

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144893.11 + phase: 0 /pseudo/partial
(652 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
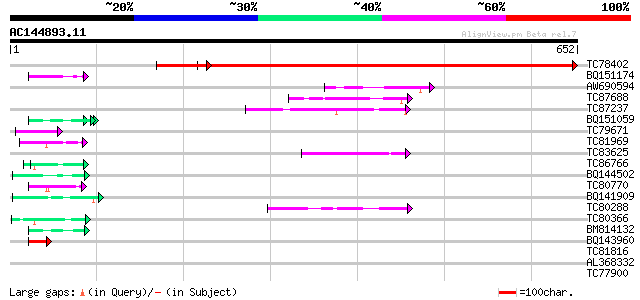
Sequences producing significant alignments: (bits) Value
TC78402 weakly similar to PIR|B84853|B84853 hypothetical protein... 858 0.0
BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xy... 51 1e-06
AW690594 similar to GP|23498163|emb hypothetical protein {Plasmo... 46 5e-05
TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~un... 45 1e-04
TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-inf... 45 1e-04
BQ151059 similar to GP|7291525|gb|A CG9888-PA {Drosophila melano... 45 1e-04
TC79671 similar to GP|21537305|gb|AAM61646.1 unknown {Arabidopsi... 44 1e-04
TC81969 similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich... 44 2e-04
TC83625 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana taba... 44 2e-04
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 44 3e-04
BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 44 3e-04
TC80770 similar to GP|4680336|gb|AAD27627.1| hypothetical protei... 44 3e-04
BQ141909 weakly similar to GP|14253159|emb VMP3 protein {Volvox ... 44 3e-04
TC80288 weakly similar to PIR|T06029|T06029 hypothetical protein... 43 4e-04
TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like... 43 4e-04
BM814132 weakly similar to OMNI|MT3615.1 PE_PGRS family protein ... 42 6e-04
BQ143960 weakly similar to SP|P21997|SSGP_ Sulfated surface glyc... 42 7e-04
TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana taba... 41 0.001
AL368332 weakly similar to GP|21322711|e pherophorin-dz1 protein... 41 0.001
TC77900 similar to GP|9759182|dbj|BAB09797.1 ATP-dependent Clp p... 41 0.002
>TC78402 weakly similar to PIR|B84853|B84853 hypothetical protein At2g42370
[imported] - Arabidopsis thaliana, partial (14%)
Length = 1982
Score = 858 bits (2216), Expect = 0.0
Identities = 423/436 (97%), Positives = 425/436 (97%)
Frame = +2
Query: 217 VGELSAEDIAFLDELASNWMLLHDDTHIMTKDVMQQLGLIKEGNLEKMDWAGLMWSMLEK 276
+G L + L SNWMLLHDDTHIMTKDVMQQLGLIKEGNLEKMDWAGLMWSMLEK
Sbjct: 146 LGNLVQRILLSLMSWQSNWMLLHDDTHIMTKDVMQQLGLIKEGNLEKMDWAGLMWSMLEK 325
Query: 277 ELKATHLEECYYASHLQHLIKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEE 336
ELKATHLEECYYASHLQHLIKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEE
Sbjct: 326 ELKATHLEECYYASHLQHLIKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEE 505
Query: 337 EDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMD 396
EDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMD
Sbjct: 506 EDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMD 685
Query: 397 FEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDD 456
FEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDD
Sbjct: 686 FEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDD 865
Query: 457 VEEDEHDVGFHFSTKHHLEGMPSGTGSSIQAMEAVQMPFGSGIHLHDSSVGDFLSARDDP 516
VEEDEHDVGFHFSTKHHLEGMPSGTGSSIQAMEAVQMPFGSGIHLHDSSVGDFLSARDDP
Sbjct: 866 VEEDEHDVGFHFSTKHHLEGMPSGTGSSIQAMEAVQMPFGSGIHLHDSSVGDFLSARDDP 1045
Query: 517 QMIHGSSLFGNGHKRDIGLDNHNSHHTLNGSNKRLRSDSPWSSQPIDFEGCMEQMQHFME 576
QMIHGSSLFGNGHKRDIGLDNHNSHHTLNGSNKRLRSDSPWSSQPIDFEGCMEQMQHFME
Sbjct: 1046QMIHGSSLFGNGHKRDIGLDNHNSHHTLNGSNKRLRSDSPWSSQPIDFEGCMEQMQHFME 1225
Query: 577 KARMMYASKDQAAEESAANQQVLLNELQRRDEMVAHLQKARIDQTQKTQIEVYRLEKELY 636
KARMMYASKDQAAEESAANQQVLLNELQRRDEMVAHLQKARIDQTQKTQIEVYRLEKELY
Sbjct: 1226KARMMYASKDQAAEESAANQQVLLNELQRRDEMVAHLQKARIDQTQKTQIEVYRLEKELY 1405
Query: 637 MMQSLVEGYRKALKET 652
MMQSLVEGYRKALKET
Sbjct: 1406MMQSLVEGYRKALKET 1453
Score = 124 bits (312), Expect = 9e-29
Identities = 64/64 (100%), Positives = 64/64 (100%)
Frame = +1
Query: 169 VAQVIASYNSTQRCSFVNDVKIMVNRAELGRAFKLPKKNSAAGGSVVDVGELSAEDIAFL 228
VAQVIASYNSTQRCSFVNDVKIMVNRAELGRAFKLPKKNSAAGGSVVDVGELSAEDIAFL
Sbjct: 1 VAQVIASYNSTQRCSFVNDVKIMVNRAELGRAFKLPKKNSAAGGSVVDVGELSAEDIAFL 180
Query: 229 DELA 232
DELA
Sbjct: 181 DELA 192
Score = 30.0 bits (66), Expect = 2.9
Identities = 28/97 (28%), Positives = 38/97 (38%), Gaps = 10/97 (10%)
Frame = -1
Query: 4 PIADTPNETLNPIT---TEQCPEQPPPPPP--PPPNPQ----PDPHFSETLENPQIQTI- 53
P + + ET +P T Q P P PP PP Q PD L P+ ++
Sbjct: 746 PHSSSDQETTSPFLLS*TAQNPSFVPLPPALFPPVKSQLYLAPDSIQYCVLPTPEPDSLQ 567
Query: 54 PDPDSTNPNHDQEDTLMDEDPTHPQIEDDPEPEPPSP 90
P P S + +H + P P + P P PSP
Sbjct: 566 PHPFSHHHSHPPHPSRPPHLPLPPHLSPHPLPPHPSP 456
>BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xylostella
granulovirus}, partial (42%)
Length = 909
Score = 51.2 bits (121), Expect = 1e-06
Identities = 28/69 (40%), Positives = 32/69 (45%)
Frame = +1
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P + PPPPPPP PQ P T + P+ T P P T P DPT PQ
Sbjct: 127 PPRGQPPPPPPPPPQAPPTKPPTRQTPKNNTPPPPPHTPP-----------DPTPPQ--- 264
Query: 82 DPEPEPPSP 90
P P+PP P
Sbjct: 265 -PPPQPPKP 288
Score = 40.8 bits (94), Expect = 0.002
Identities = 32/122 (26%), Positives = 43/122 (35%), Gaps = 11/122 (9%)
Frame = +3
Query: 9 PNETLNPI--------TTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTN 60
PN+T N TT P +P PP PPP Q P + DP +T
Sbjct: 177 PNQTTNTTNPKKQHTPTTPTHPPRPHTPPTPPPTTQ---------TTPPREAPTDPGTTR 329
Query: 61 ---PNHDQEDTLMDEDPTHPQIEDDPEPEPPSPAATTVRRGHKRKKFGGKRTAAQKRKSH 117
P D T P + P PP P R GH+ + KRT + +
Sbjct: 330 GTPPTADDAQTSPAHHPGEGGGKGGAPPPPPRPPPRHGRGGHQNQTREAKRTTTEPQGKR 509
Query: 118 EK 119
++
Sbjct: 510 DR 515
Score = 37.7 bits (86), Expect = 0.014
Identities = 31/117 (26%), Positives = 41/117 (34%), Gaps = 22/117 (18%)
Frame = +3
Query: 25 PPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTH-------- 76
PPPPP PPP H ++T E + T P P D P H
Sbjct: 408 PPPPPRPPPRHGRGGHQNQTREAKRTTTEPQGKRDRPRTDPRK--KKHPPPHQKEGGNTQ 581
Query: 77 --PQIEDDPE------------PEPPSPAATTVRRGHKRKKFGGKRTAAQKRKSHEK 119
P+ ED P P PP P T RG + + G R +K + ++
Sbjct: 582 KEPRGEDPPPHTPAGGGGGPGGPRPPPPKKTA--RGGEERPPAGARARREKTRGDQQ 746
Score = 36.6 bits (83), Expect = 0.031
Identities = 28/102 (27%), Positives = 34/102 (32%), Gaps = 3/102 (2%)
Frame = +1
Query: 4 PIADTPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNH 63
P TP P P+ P PP PPP P PH + P+ P T P
Sbjct: 190 PTRQTPKNNTPPPPPHTPPD--PTPPQPPPQPPKPPHHEKRPRTPEPPGGRPPPPTTPKQ 363
Query: 64 DQEDTLMDEDPTHPQIEDDPEPEPPSPAA---TTVRRGHKRK 102
+ T E P P PP T R G +R+
Sbjct: 364 ARPTT--QERGGARGARPRPPPAPPQGTGGEDTKTRPGKQRE 483
Score = 36.6 bits (83), Expect = 0.031
Identities = 25/79 (31%), Positives = 31/79 (38%)
Frame = +3
Query: 18 TEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHP 77
T++ P + PPP P P P P P QT ++TNP T PTHP
Sbjct: 96 TKKSPRKKEKPPPRPATPPPAP----APPGPPNQT---TNTTNPKKQHTPT----TPTHP 242
Query: 78 QIEDDPEPEPPSPAATTVR 96
P PP+ T R
Sbjct: 243 PRPHTPPTPPPTTQTTPPR 299
Score = 36.2 bits (82), Expect = 0.041
Identities = 25/81 (30%), Positives = 33/81 (39%)
Frame = +3
Query: 14 NPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDED 73
+P E+ P +P PPP P P P P+ + NP+ Q P T P H
Sbjct: 105 SPRKKEKPPPRPATPPPAPAPPGP-PNQTTNTTNPKKQHTP----TTPTHPPRPHTPPTP 269
Query: 74 PTHPQIEDDPEPEPPSPAATT 94
P P + P E P+ TT
Sbjct: 270 P--PTTQTTPPREAPTDPGTT 326
Score = 35.0 bits (79), Expect = 0.091
Identities = 28/116 (24%), Positives = 41/116 (35%), Gaps = 5/116 (4%)
Frame = +2
Query: 4 PIADTPNETLNPITTEQCPEQPPPPP-----PPPPNPQPDPHFSETLENPQIQTIPDPDS 58
P +TPN ++ P++ PP PP P P P + + PQ +
Sbjct: 56 PKPETPNRKKPQKNKKEPPKKRKTPPEASHPPPRPRPPRPPQPNHQHDKPQ-------KT 214
Query: 59 TNPNHDQEDTLMDEDPTHPQIEDDPEPEPPSPAATTVRRGHKRKKFGGKRTAAQKR 114
T+P+H T P P P P P T R H + G ++R
Sbjct: 215 THPHHPH---------TPPPTPHPPNPPPNHPNHPTTRSAHGPRNHPGDAPHRRRR 355
Score = 29.3 bits (64), Expect = 5.0
Identities = 28/107 (26%), Positives = 35/107 (32%), Gaps = 18/107 (16%)
Frame = +2
Query: 20 QCPEQPPPPPPPPP--NPQPDPHFSE------------TLENPQIQTIPDPDSTNPNHDQ 65
+ P PPPPP P+PDP E T P+ +T P H +
Sbjct: 404 RAPAPPPPPPKAREGRTPKPDPGSKENHNRTPGKEGPATHGPPEKKTPPPTPKRRGKHTK 583
Query: 66 EDTLMDEDPTHPQI----EDDPEPEPPSPAATTVRRGHKRKKFGGKR 108
P HP P P PP + R R + G KR
Sbjct: 584 RAPGGGPPPPHPGRGGGGPGGPPPPPPKKNSERGGRAAPRGRPGTKR 724
Score = 29.3 bits (64), Expect = 5.0
Identities = 15/38 (39%), Positives = 21/38 (54%)
Frame = +3
Query: 84 EPEPPSPAATTVRRGHKRKKFGGKRTAAQKRKSHEKFE 121
+P PSP AT R+GH +K R+ + +KS K E
Sbjct: 9 DPPTPSPQATPQRKGHPSQKPPTGRSHKKTKKSPRKKE 122
>AW690594 similar to GP|23498163|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (10%)
Length = 633
Score = 45.8 bits (107), Expect = 5e-05
Identities = 39/132 (29%), Positives = 59/132 (44%), Gaps = 6/132 (4%)
Frame = +3
Query: 363 QELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLR 422
+++EEHN E G+ VE+E+ E ++ + E ++EE E
Sbjct: 36 RKVEEHNEEEDEGE-------VEEEEDEHDEEEEDEHDEEEEEE---------------- 146
Query: 423 PCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVGFHFST------KHHLEG 476
+ + E +DE EEEE E+EEEEEDD EE E D ST H L
Sbjct: 147 -------EEEEEEDEHDDEEEEEEEEEEEEEEDDDEEGEEDEIDRISTLPDPLLLHILSF 305
Query: 477 MPSGTGSSIQAM 488
+P+ T + ++
Sbjct: 306 LPTRTSVATMSL 341
Score = 37.4 bits (85), Expect = 0.018
Identities = 27/85 (31%), Positives = 44/85 (51%)
Frame = +3
Query: 299 QHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVE 358
+H E +E V E E E +E ++E+E E ++EEEEE+ D+ + + + E
Sbjct: 48 EHNEEEDEGEVEEEEDEHDE---EEEDEHDEEEEEEEEEEEEDEHDDEEEEEEEEEEEEE 218
Query: 359 ENQVQELEEHNIELSLGQDKVETLP 383
E+ +E EE I D++ TLP
Sbjct: 219 EDDDEEGEEDEI------DRISTLP 275
Score = 34.3 bits (77), Expect = 0.15
Identities = 20/51 (39%), Positives = 31/51 (60%)
Frame = +3
Query: 297 KSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVD 347
+ +H E EE E E E +E ++EEEE E ++EEE++D +DE+D
Sbjct: 114 EDEHDEEEEEE---EEEEEEDEHDDEEEEEEEEEEEEEEDDDEEGEEDEID 257
>TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~unknown
protein {Arabidopsis thaliana}, partial (46%)
Length = 2077
Score = 44.7 bits (104), Expect = 1e-04
Identities = 42/147 (28%), Positives = 67/147 (45%), Gaps = 4/147 (2%)
Frame = +2
Query: 321 VKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIELSLGQDKVE 380
V+ EEEE + DEE ++D D+ +G K G E + + EE + E G + +
Sbjct: 365 VELEEEESDDFDEELDDDD----DDFEGVEKKKAKGGSEGE-DDFEEEDDEDEEGSEDED 529
Query: 381 TLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKED 440
EKE+ +G + D E TE + NY KG + ++ ED
Sbjct: 530 DEEDEKEKVKGGGIEDKFLKIDELTEYLEKEEDNY-------------EKGEERDEADED 670
Query: 441 EGEEEEHEQ----EEEEEDDVEEDEHD 463
E++E E+ E ++EDD ++DE D
Sbjct: 671 SEEDDELEKAGEFEMDDEDDDDDDEED 751
Score = 40.4 bits (93), Expect = 0.002
Identities = 40/169 (23%), Positives = 75/169 (43%), Gaps = 9/169 (5%)
Frame = +2
Query: 296 IKSQHKELFEETLVVEVEGEGEEGVVKDE-------EEEGEAKDEEEEEDGVVVKDEVDG 348
++ + + F+E L + + + EGV K + E++ E +D+E+EE DE D
Sbjct: 371 LEEEESDDFDEEL--DDDDDDFEGVEKKKAKGGSEGEDDFEEEDDEDEEGSEDEDDEEDE 544
Query: 349 SGDVKMGGVEEN--QVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETE 406
VK GG+E+ ++ EL E+ L +D E +GE+ + ++ +E+ E
Sbjct: 545 KEKVKGGGIEDKFLKIDELTEY---LEKEEDNYE---------KGEERDEADEDSEEDDE 688
Query: 407 MWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEED 455
+ G E +DE ++++ E++EE ED
Sbjct: 689 LEKAG-----------------------EFEMDDEDDDDDDEEDEEAED 766
Score = 32.0 bits (71), Expect = 0.77
Identities = 26/107 (24%), Positives = 50/107 (46%), Gaps = 6/107 (5%)
Frame = +2
Query: 363 QELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLR 422
+E+ + + L +G+ + VE E+ E + DF++ ++ +
Sbjct: 302 EEIAQFKVPLDVGKKLEKKKRVELEEEESD---DFDEELDDDDD---------------- 424
Query: 423 PCHNRDRKGIDCEQVK-----EDEGEEEEHEQEE-EEEDDVEEDEHD 463
D +G++ ++ K ED+ EEE+ E EE E++D EEDE +
Sbjct: 425 -----DFEGVEKKKAKGGSEGEDDFEEEDDEDEEGSEDEDDEEDEKE 550
>TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-infected
erythrocyte surface antigen {Plasmodium falciparum},
partial (2%)
Length = 2007
Score = 44.7 bits (104), Expect = 1e-04
Identities = 50/198 (25%), Positives = 85/198 (42%), Gaps = 8/198 (4%)
Frame = +1
Query: 272 SMLEKELKATHLEECYYASHLQHLIKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAK 331
S LE E E SHL++ + E EE +E E KDEE+ +
Sbjct: 10 SHLENEESKETGESSEEKSHLENEESKETGESSEEKSHLENEEN------KDEEKSKQEN 171
Query: 332 DEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIELSL---GQDKVETLPVEKEQ 388
+E ++ + + ++E + + EEN+ +E + EL G+ + + EK +
Sbjct: 172 EEIKDGEKIQQENEENKDEEKSQQENEENKDEEKSQQENELKKNEGGEKETGEITEEKSK 351
Query: 389 GEGEQMMDFEQSKKEETEMWFLGQ--KNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEE 446
E E+ + KE E G K VGE H +D K E + GE+EE
Sbjct: 352 QENEETSETNSKDKENEESNQNGSDAKEQVGEN-----HEQDSKQGTEETNGTEGGEKEE 516
Query: 447 HEQEEEE---EDDVEEDE 461
H++ +E+ ++ V++ E
Sbjct: 517 HDKIKEDTSSDNQVQDGE 570
Score = 31.2 bits (69), Expect = 1.3
Identities = 45/251 (17%), Positives = 98/251 (38%), Gaps = 11/251 (4%)
Frame = +1
Query: 385 EKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKED---- 440
EK E E+ + +S +E++ + K GE S H + + D E+ K++
Sbjct: 4 EKSHLENEESKETGESSEEKSHLENEESKE-TGESSEEKSHLENEENKDEEKSKQENEEI 180
Query: 441 -EGEEEEHEQEEEEEDDVEEDEHDVGFHFSTKHHLEGMPSGTGSSIQAMEAVQMPFGSGI 499
+GE+ + E EE ++++ + E++ + G + E + S
Sbjct: 181 KDGEKIQQENEENKDEEKSQQENEENKDEEKSQQENELKKNEGGEKETGEITEEK--SKQ 354
Query: 500 HLHDSSVGDFLSARDDPQMIHGSSL---FGNGHKRDIGLDNHNSHHTLNGSNK---RLRS 553
++S + ++ +GS G H++D ++ T G + +++
Sbjct: 355 ENEETSETNSKDKENEESNQNGSDAKEQVGENHEQDSKQGTEETNGTEGGEKEEHDKIKE 534
Query: 554 DSPWSSQPIDFEGCMEQMQHFMEKARMMYASKDQAAEESAANQQVLLNELQRRDEMVAHL 613
D+ +Q D E E + A D ++ES+ + ++ + ++E
Sbjct: 535 DTSSDNQVQDGEKNNEAREENYSGDNASSAVVDNKSQESSNKTEEQFDK-KEKNEFELES 711
Query: 614 QKARIDQTQKT 624
QK + T+ T
Sbjct: 712 QKNSNETTEST 744
>BQ151059 similar to GP|7291525|gb|A CG9888-PA {Drosophila melanogaster},
partial (26%)
Length = 308
Score = 44.7 bits (104), Expect = 1e-04
Identities = 27/81 (33%), Positives = 27/81 (33%)
Frame = -1
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P PPPPPPPPP P P P P P P P P
Sbjct: 221 PPPPPPPPPPPPPPPPPPP-------------PPPPPPPPPPPPPPPPPPPPPPPPPPPP 81
Query: 82 DPEPEPPSPAATTVRRGHKRK 102
P P PP P RG RK
Sbjct: 80 PPPPPPPPPPPPPPPRGFPRK 18
Score = 43.9 bits (102), Expect = 2e-04
Identities = 24/69 (34%), Positives = 24/69 (34%)
Frame = -2
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P PPPPPPPPP P P P P P P P P P
Sbjct: 301 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 122
Query: 82 DPEPEPPSP 90
P P PP P
Sbjct: 121 PPPPPPPPP 95
Score = 43.9 bits (102), Expect = 2e-04
Identities = 24/69 (34%), Positives = 24/69 (34%)
Frame = -1
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P PPPPPPPPP P P P P P P P P P
Sbjct: 305 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 126
Query: 82 DPEPEPPSP 90
P P PP P
Sbjct: 125 PPPPPPPPP 99
Score = 43.9 bits (102), Expect = 2e-04
Identities = 24/69 (34%), Positives = 24/69 (34%)
Frame = -1
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P PPPPPPPPP P P P P P P P P P
Sbjct: 293 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 114
Query: 82 DPEPEPPSP 90
P P PP P
Sbjct: 113 PPPPPPPPP 87
Score = 43.9 bits (102), Expect = 2e-04
Identities = 24/69 (34%), Positives = 24/69 (34%)
Frame = -3
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P PPPPPPPPP P P P P P P P P P
Sbjct: 285 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 106
Query: 82 DPEPEPPSP 90
P P PP P
Sbjct: 105 PPPPPPPPP 79
Score = 43.9 bits (102), Expect = 2e-04
Identities = 24/69 (34%), Positives = 24/69 (34%)
Frame = -2
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P PPPPPPPPP P P P P P P P P P
Sbjct: 283 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 104
Query: 82 DPEPEPPSP 90
P P PP P
Sbjct: 103 PPPPPPPPP 77
Score = 43.9 bits (102), Expect = 2e-04
Identities = 24/69 (34%), Positives = 24/69 (34%)
Frame = -3
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P PPPPPPPPP P P P P P P P P P
Sbjct: 306 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 127
Query: 82 DPEPEPPSP 90
P P PP P
Sbjct: 126 PPPPPPPPP 100
Score = 43.9 bits (102), Expect = 2e-04
Identities = 24/69 (34%), Positives = 24/69 (34%)
Frame = -1
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P PPPPPPPPP P P P P P P P P P
Sbjct: 299 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 120
Query: 82 DPEPEPPSP 90
P P PP P
Sbjct: 119 PPPPPPPPP 93
Score = 42.4 bits (98), Expect = 6e-04
Identities = 26/77 (33%), Positives = 27/77 (34%)
Frame = -3
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P PPPPPPPPP P P P P P P P P P
Sbjct: 198 PPPPPPPPPPPPPPPPPP------PPPPPPPPPPPPPPPP---------PPPPPPPPPPP 64
Query: 82 DPEPEPPSPAATTVRRG 98
P P PP P + RG
Sbjct: 63 PPPPPPPPPPGFSPERG 13
>TC79671 similar to GP|21537305|gb|AAM61646.1 unknown {Arabidopsis
thaliana}, partial (36%)
Length = 947
Score = 44.3 bits (103), Expect = 1e-04
Identities = 20/54 (37%), Positives = 26/54 (48%)
Frame = +1
Query: 7 DTPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTN 60
+ P +N I PE PPPPP P P+P P F+E L N + + D N
Sbjct: 28 EEPVPDMNEIKALPAPENYTPPPPPEPEPEPKPQFTEDLVNLREDAVTADDQGN 189
>TC81969 similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (11%)
Length = 556
Score = 43.9 bits (102), Expect = 2e-04
Identities = 27/81 (33%), Positives = 36/81 (44%), Gaps = 3/81 (3%)
Frame = -2
Query: 12 TLNPITTEQCPEQPPPPPPPPPNPQPDPHF---SETLENPQIQTIPDPDSTNPNHDQEDT 68
+LNP ++ P PPPPPP +P P P+ LE P + P+P + + Q
Sbjct: 372 SLNPSSSSPPPPSPPPPPPSFSHPSPSPNLPFSPPPLEPPSSGSSPNPSCSKVS--QSTL 199
Query: 69 LMDEDPTHPQIEDDPEPEPPS 89
L P P P P PPS
Sbjct: 198 LCIPPPQSPSC--PPSPRPPS 142
Score = 28.9 bits (63), Expect = 6.5
Identities = 18/68 (26%), Positives = 35/68 (51%), Gaps = 3/68 (4%)
Frame = +3
Query: 390 EGEQMMDFEQSKKEETEMWFLGQ---KNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEE 446
E E+ +D ++ + E +G+ K G+ + + K + E+ K+ + +EEE
Sbjct: 147 EVEEKVDTMETGEGECIAM*IGKL*NKKDSGKNQKKEVQEEEEKTANLEKAKDGKTKEEE 326
Query: 447 HEQEEEEE 454
E++EEEE
Sbjct: 327 EEEKEEEE 350
>TC83625 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana tabacum},
partial (9%)
Length = 908
Score = 43.9 bits (102), Expect = 2e-04
Identities = 31/126 (24%), Positives = 63/126 (49%)
Frame = +2
Query: 336 EEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMM 395
EE + +K + +GDVK VE + E++ + + K + + + EG+
Sbjct: 461 EESNIPIKIDDKSAGDVKEDKVEIEKDLEIKSVEKDDEKKEKKDKEKKDKTDVDEGKDKK 640
Query: 396 DFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEED 455
D E+ KKE+ E G++ E + +++K E K+ +GEE++ ++++E++
Sbjct: 641 DKEKKKKEKKEENVKGEEEDGDEKKDKEKKKKEKKEKGKED-KDKDGEEKKSKKDKEKKK 817
Query: 456 DVEEDE 461
D ED+
Sbjct: 818 DKNEDD 835
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 43.5 bits (101), Expect = 3e-04
Identities = 27/76 (35%), Positives = 29/76 (37%), Gaps = 2/76 (2%)
Frame = +2
Query: 17 TTEQCPEQPPPPPP--PPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDP 74
T C PPPPPP PPP P P P FS P + P P P H P
Sbjct: 5 TPPYCVRSPPPPPPNSPPPPPPPAPVFSPPPPVPYYYSSPPPP---PAH---------SP 148
Query: 75 THPQIEDDPEPEPPSP 90
P P P P+P
Sbjct: 149PPPPXSPPPPPHSPTP 196
Score = 42.7 bits (99), Expect = 4e-04
Identities = 24/69 (34%), Positives = 28/69 (39%), Gaps = 3/69 (4%)
Frame = +2
Query: 25 PPP---PPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
PPP PPPPP +P P PH P + P P P H + P P
Sbjct: 128 PPPAHSPPPPPXSPPPPPHSPTPPVYPYLSPPPPP----PVHSPPPPVYSPPPPSPPPCV 295
Query: 82 DPEPEPPSP 90
+P P PP P
Sbjct: 296 EPPPPPPPP 322
Score = 38.9 bits (89), Expect = 0.006
Identities = 29/92 (31%), Positives = 33/92 (35%), Gaps = 10/92 (10%)
Frame = +2
Query: 9 PNETLNPITTEQCPEQPPP---PPPP---PPNPQPDPHFSETLENPQIQTIPDPDSTNPN 62
P+ P+ P PPP PPPP PP P P P P P P S+
Sbjct: 179 PHSPTPPVYPYLSPPPPPPVHSPPPPVYSPPPPSPPPCVEPPPPPPPPCVEPPPPSSPAP 358
Query: 63 HDQEDTLMDEDPTHPQIEDDPEPEP----PSP 90
H + P HP P P P PSP
Sbjct: 359 H--------QTPYHPPPSPSPPPSPVYAYPSP 430
Score = 38.5 bits (88), Expect = 0.008
Identities = 31/109 (28%), Positives = 38/109 (34%), Gaps = 7/109 (6%)
Frame = +2
Query: 4 PIADTPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFS----ETLENPQIQTIPDPDST 59
P+ P +P P PPPPPPPP +P P S +T +P P P
Sbjct: 233 PVHSPPPPVYSPPPPSPPPCVEPPPPPPPPCVEPPPPSSPAPHQTPYHPPPSPSPPPSPV 412
Query: 60 NPNHDQEDTLMDEDPTHPQIEDDPEPEPP---SPAATTVRRGHKRKKFG 105
+ P P + P P PP SP V G FG
Sbjct: 413 YAYPSPPPPVYTSPPPSP-VYAYPSPPPPVYSSPPPPPVYEGPIPPVFG 556
Score = 34.3 bits (77), Expect = 0.15
Identities = 23/75 (30%), Positives = 30/75 (39%), Gaps = 3/75 (4%)
Frame = +2
Query: 22 PEQPP--PPPPPP-PNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQ 78
P PP PPPPP P P P+ S P + + P P + P + P P
Sbjct: 146 PPPPPXSPPPPPHSPTPPVYPYLSPP-PPPPVHSPPPPVYSPPPPSPPPCVEPPPPPPPP 322
Query: 79 IEDDPEPEPPSPAAT 93
+ P P P+P T
Sbjct: 323 CVEPPPPSSPAPHQT 367
>BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (39%)
Length = 1358
Score = 43.5 bits (101), Expect = 3e-04
Identities = 27/88 (30%), Positives = 30/88 (33%)
Frame = -2
Query: 4 PIADTPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNH 63
P A P P P +PPPPPPPPP P P P P + P
Sbjct: 223 PRASPPPPPPPPGPAASAPPRPPPPPPPPPQRPPPP--------------PPPHTHQP-- 92
Query: 64 DQEDTLMDEDPTHPQIEDDPEPEPPSPA 91
E PT P P P P P+
Sbjct: 91 --------EPPTSPPRRQPPAPRAPPPS 32
Score = 40.4 bits (93), Expect = 0.002
Identities = 28/93 (30%), Positives = 33/93 (35%), Gaps = 3/93 (3%)
Frame = -1
Query: 1 MTNPIADTPNETLNP---ITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPD 57
+ I P L P + T P +PPPP PPPP+P P SE P P P
Sbjct: 638 LAKKIGGQPRPALVPLVRVATYPRPPRPPPPVPPPPSPPRAPPQSEAAPPPP----PPPR 471
Query: 58 STNPNHDQEDTLMDEDPTHPQIEDDPEPEPPSP 90
P P P DP P +P
Sbjct: 470 GARP----------RAPLRPPSAPDPAGHPRAP 402
Score = 36.6 bits (83), Expect = 0.031
Identities = 26/86 (30%), Positives = 31/86 (35%), Gaps = 10/86 (11%)
Frame = -2
Query: 14 NPITTEQCPEQPPPPPP----PPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTL 69
+P+ P P PPPP PPP P P P S P+ P P P
Sbjct: 331 SPVPCPALPHPPGPPPPLDRAPPPPPAPLPLASRL--RPRASPPPPPPPPGPAASAPPRP 158
Query: 70 MDEDPTHPQIEDDP------EPEPPS 89
P PQ P +PEPP+
Sbjct: 157 PPPPPPPPQRPPPPPPPHTHQPEPPT 80
Score = 35.8 bits (81), Expect = 0.053
Identities = 27/106 (25%), Positives = 36/106 (33%), Gaps = 12/106 (11%)
Frame = -1
Query: 4 PIADTPNETLNPITTEQCPEQPPPP------PPPPPNPQPDPHFSETLENPQIQTIPDPD 57
P+ P+ P +E P PPPP P P PDP + +P P
Sbjct: 548 PVPPPPSPPRAPPQSEAAPPPPPPPRGARPRAPLRPPSAPDPAGHPRAPRQGVAVVPGPR 369
Query: 58 STNP------NHDQEDTLMDEDPTHPQIEDDPEPEPPSPAATTVRR 97
S N + PT P P P PP+P T + +
Sbjct: 368 SGNSPRAAHRGPEPRPLPRPPPPTRPPAAPGPGP-PPAPGPTPLSK 234
Score = 35.4 bits (80), Expect = 0.069
Identities = 20/69 (28%), Positives = 26/69 (36%)
Frame = -3
Query: 25 PPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIEDDPE 84
PPPPPPPPP+ P + + P + P+ PT P P
Sbjct: 696 PPPPPPPPPSSPPGMGTRPSCQENWWAAPPGARPSRPSR------YIPPPTSPPPASPPA 535
Query: 85 PEPPSPAAT 93
P PP +T
Sbjct: 534 PVPPQSPST 508
Score = 33.1 bits (74), Expect = 0.34
Identities = 31/117 (26%), Positives = 41/117 (34%), Gaps = 20/117 (17%)
Frame = -3
Query: 3 NPIADTPNETLNPITTEQCPEQPPP------PPPPPPNPQP------DPHFSETLENPQI 50
+P A P ++ + I P PPP PPP P P+P P ++
Sbjct: 546 SPPAPVPPQSPSTIRGRAPPPAPPPRRPPARPPPAPQRPRPRRPPARPPSGRRRGARAKV 367
Query: 51 QTIPD---PDSTNPNHDQEDTLMDEDPTHPQIE-----DDPEPEPPSPAATTVRRGH 99
+ +P P S P+ PTHP P P P SP RGH
Sbjct: 366 RKLPKGGTPGSRAPSP------APPSPTHPAPRRPWTGPPPRPRPHSP*QAACGRGH 214
Score = 32.7 bits (73), Expect = 0.45
Identities = 25/90 (27%), Positives = 36/90 (39%), Gaps = 3/90 (3%)
Frame = -1
Query: 4 PIADTPNET--LNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIP-DPDSTN 60
P A P+ +P T P PP PPPPPP + P + P+ +P +T
Sbjct: 746 PCARAPHSIPRAHPAATPLLP--PPRPPPPPPEWERVPLAKKIGGQPRPALVPLVRVATY 573
Query: 61 PNHDQEDTLMDEDPTHPQIEDDPEPEPPSP 90
P + + P+ P+ E PP P
Sbjct: 572 PRPPRPPPPVPPPPSPPRAPPQSEAAPPPP 483
Score = 30.8 bits (68), Expect = 1.7
Identities = 25/94 (26%), Positives = 33/94 (34%), Gaps = 10/94 (10%)
Frame = -1
Query: 9 PNETLNPITTEQCPE-----QPPPPPPPP----PNPQPDPHFSETLENPQIQTIPDPDST 59
P +P + PE +PPPP PP P P P P + + P + IP +
Sbjct: 374 PRSGNSPRAAHRGPEPRPLPRPPPPTRPPAAPGPGPPPAPGPTPLSKPPAAEGIPPATAP 195
Query: 60 NPNHDQ-EDTLMDEDPTHPQIEDDPEPEPPSPAA 92
P + P P P P PP A
Sbjct: 194 PPRARRLRPPSAPAPPPTPPAAAPPAPTPPHAPA 93
Score = 30.0 bits (66), Expect = 2.9
Identities = 23/96 (23%), Positives = 33/96 (33%)
Frame = -3
Query: 2 TNPIADTPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNP 61
T+P +P + P + + PPP PPP P P + P+ P
Sbjct: 564 TSPPPASPPAPVPPQSPSTIRGRAPPPAPPPRRPPARPPPAPQRPRPRRPPARPPSGRRR 385
Query: 62 NHDQEDTLMDEDPTHPQIEDDPEPEPPSPAATTVRR 97
+ + + T P P PPSP RR
Sbjct: 384 GARAKVRKLPKGGTPG--SRAPSPAPPSPTHPAPRR 283
>TC80770 similar to GP|4680336|gb|AAD27627.1| hypothetical protein {Oryza
sativa subsp. indica} [Oryza sativa (indica
cultivar-group)], partial (7%)
Length = 507
Score = 43.5 bits (101), Expect = 3e-04
Identities = 27/81 (33%), Positives = 33/81 (40%), Gaps = 14/81 (17%)
Frame = -2
Query: 22 PEQPPPPPPPPPNPQPDPH---------FSE-----TLENPQIQTIPDPDSTNPNHDQED 67
P + PP PP P+PQP PH FS L +P Q P P P
Sbjct: 281 PSKISPPQPPHPSPQPSPHPLLFFMLS*FSHPPLPTILPHPPPQPAPQPPPHPPPQPPLF 102
Query: 68 TLMDEDPTHPQIEDDPEPEPP 88
T+ + P HP P P+PP
Sbjct: 101 TIFPQPPPHP--PPQPPPQPP 45
Score = 33.1 bits (74), Expect = 0.34
Identities = 17/37 (45%), Positives = 18/37 (47%), Gaps = 5/37 (13%)
Frame = -2
Query: 9 PNETLNPITTEQCPEQPPPPPPPPPN-----PQPDPH 40
P T+ P Q QPPP PPP P PQP PH
Sbjct: 185 PLPTILPHPPPQPAPQPPPHPPPQPPLFTIFPQPPPH 75
Score = 30.4 bits (67), Expect = 2.2
Identities = 15/36 (41%), Positives = 16/36 (43%), Gaps = 4/36 (11%)
Frame = -2
Query: 9 PNETLNPITTEQCPEQPPPPPPPPP----NPQPDPH 40
P L I + P PP PPP PP P P PH
Sbjct: 119 PQPPLFTIFPQPPPHPPPQPPPQPPLLIIFPHPPPH 12
Score = 28.5 bits (62), Expect = 8.5
Identities = 15/40 (37%), Positives = 18/40 (44%), Gaps = 7/40 (17%)
Frame = -2
Query: 3 NPIADTPNETLNPITTEQCPEQPPPPPP-------PPPNP 35
+P P T+ P P QPPP PP PPP+P
Sbjct: 128 HPPPQPPLFTIFPQPPPHPPPQPPPQPPLLIIFPHPPPHP 9
>BQ141909 weakly similar to GP|14253159|emb VMP3 protein {Volvox carteri f.
nagariensis}, partial (13%)
Length = 1223
Score = 43.5 bits (101), Expect = 3e-04
Identities = 33/111 (29%), Positives = 45/111 (39%), Gaps = 6/111 (5%)
Frame = -2
Query: 4 PIADTPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNH 63
P+ +P+ TL P P PPPPPPP P P P P S P +++ P P
Sbjct: 352 PLRSSPSLTLPPSFCSSPPFLPPPPPPPSPPPPPPPPSSPL--RPPLRSPPPLPPLPP-- 185
Query: 64 DQEDTLMDEDPTHPQIEDDPEPEPPSPAATTV------RRGHKRKKFGGKR 108
L P+ P P ++ V +R K+KK GG+R
Sbjct: 184 -----LPPFTPSSHLTTSSPPPSFLRCISSVVFIFPLRKRDGKKKKIGGRR 47
Score = 30.4 bits (67), Expect = 2.2
Identities = 14/41 (34%), Positives = 21/41 (51%)
Frame = -1
Query: 1 MTNPIADTPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHF 41
+++P +P + P ++ PPP PPPPP P P F
Sbjct: 296 LSSPSPSSPLPSPPPPSSLFPSPTPPPLPPPPPPPPPASPF 174
>TC80288 weakly similar to PIR|T06029|T06029 hypothetical protein T28I19.100
- Arabidopsis thaliana, partial (14%)
Length = 1460
Score = 42.7 bits (99), Expect = 4e-04
Identities = 41/169 (24%), Positives = 75/169 (44%), Gaps = 2/169 (1%)
Frame = +3
Query: 297 KSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKD-EEEEEDGVVVKDEVDGSGDVKMG 355
+ +E +E +V ++ + E + E EEG + EEE ED V +
Sbjct: 393 EGHEEEEEDEHIVYNMQNKREHDEQQQEGEEGNKHETEEESEDNVHER------------ 536
Query: 356 GVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNY 415
E Q +E +H E+ Q++ E+ E E G+ +D +K E +
Sbjct: 537 --REEQDEEENKHGAEV---QEENESKSEEVEDEGGDVEIDENDHEKSEAD--------- 674
Query: 416 VGEPSLRPCHNRDRKGIDCEQVKEDEGEEE-EHEQEEEEEDDVEEDEHD 463
++R+ + +D E+ KE+EG++E E+E +E+EE + H+
Sbjct: 675 ---------NDREDEVVDEEKDKEEEGDDETENEDKEDEEKGGLVENHE 794
Score = 35.8 bits (81), Expect = 0.053
Identities = 36/141 (25%), Positives = 60/141 (42%), Gaps = 8/141 (5%)
Frame = +3
Query: 331 KDEEEEEDGVVVKDEVD-----GSGDVKMGGVEENQVQELEEHNIELSLGQDKVETLPVE 385
K+ ++ + + ++ E D G D+ G VE ++ + EE +D+ ++
Sbjct: 282 KEFDKNDTKLPIRTETDQILKLGRKDLHPGKVEADKNEGHEEEE------EDEHIVYNMQ 443
Query: 386 KEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEE 445
++ EQ + E+ K ETE E S H R + + E E +EE
Sbjct: 444 NKREHDEQQQEGEEGNKHETE-----------EESEDNVHERREEQDEEENKHGAEVQEE 590
Query: 446 EHEQEEEEED---DVEEDEHD 463
+ EE ED DVE DE+D
Sbjct: 591 NESKSEEVEDEGGDVEIDEND 653
>TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like protein
{Arabidopsis thaliana}, partial (16%)
Length = 814
Score = 42.7 bits (99), Expect = 4e-04
Identities = 30/98 (30%), Positives = 39/98 (39%), Gaps = 8/98 (8%)
Frame = +3
Query: 3 NPIADTPNETLNPITTEQCPEQPPP-------PPPPPPNPQPDPHFSETLENPQIQ-TIP 54
NP TP+ P +T P PPP PP P N P P + + P T P
Sbjct: 54 NPPPVTPSAP--PPSTPATPSSPPPSTTPSAPPPSTPSNSPPSPPTTPAISPPSGGGTTP 227
Query: 55 DPDSTNPNHDQEDTLMDEDPTHPQIEDDPEPEPPSPAA 92
P S P D+ P+ P + P P PPS ++
Sbjct: 228 SPPSRTPPSS------DDSPSPPSSKTPPPPSPPSSSS 323
Score = 33.5 bits (75), Expect = 0.26
Identities = 24/80 (30%), Positives = 28/80 (35%)
Frame = +3
Query: 15 PITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDP 74
P T P P PPP P P T +P P ST P+ T + P
Sbjct: 15 PATPSSPPPSTPANPPPVTPSAPPPSTPATPSSP-------PPSTTPSAPPPSTPSNSPP 173
Query: 75 THPQIEDDPEPEPPSPAATT 94
+ P P PPS TT
Sbjct: 174SPP---TTPAISPPSGGGTT 224
>BM814132 weakly similar to OMNI|MT3615.1 PE_PGRS family protein
{Mycobacterium tuberculosis CDC1551}, partial (7%)
Length = 164
Score = 42.4 bits (98), Expect = 6e-04
Identities = 25/70 (35%), Positives = 26/70 (36%)
Frame = -3
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P PPPPPPPPP P P P P + P P P P P
Sbjct: 162 PPPPPPPPPPPPPPPPPP--------PPPRPPPPPPPPPP-----------PPPPPPPPP 40
Query: 82 DPEPEPPSPA 91
P P PP PA
Sbjct: 39 PPPPPPPPPA 10
Score = 40.8 bits (94), Expect = 0.002
Identities = 24/69 (34%), Positives = 24/69 (34%)
Frame = -1
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P PPPPPPPPP P P P P P P P P P
Sbjct: 164 PPPPPPPPPPPPPPPPPP--------PPPPAPPPPPPPPP---------PRPPPPPPPPP 36
Query: 82 DPEPEPPSP 90
P P PP P
Sbjct: 35 PPPPPPPPP 9
Score = 38.1 bits (87), Expect = 0.011
Identities = 13/18 (72%), Positives = 14/18 (77%)
Frame = -2
Query: 22 PEQPPPPPPPPPNPQPDP 39
P PPPPPPPPP P+P P
Sbjct: 55 PPPPPPPPPPPPPPRPPP 2
Score = 37.4 bits (85), Expect = 0.018
Identities = 13/18 (72%), Positives = 13/18 (72%)
Frame = -1
Query: 22 PEQPPPPPPPPPNPQPDP 39
P PPPPPPPPP P P P
Sbjct: 59 PPPPPPPPPPPPPPPPPP 6
Score = 35.4 bits (80), Expect = 0.069
Identities = 16/34 (47%), Positives = 16/34 (47%)
Frame = -1
Query: 4 PIADTPNETLNPITTEQCPEQPPPPPPPPPNPQP 37
P A P P P PPPPPPPPP P P
Sbjct: 104 PPAPPPPPPPPPPRPPPPPPPPPPPPPPPPPPPP 3
>BQ143960 weakly similar to SP|P21997|SSGP_ Sulfated surface glycoprotein
185 (SSG 185). {Volvox carteri}, partial (13%)
Length = 1023
Score = 42.0 bits (97), Expect = 7e-04
Identities = 16/27 (59%), Positives = 17/27 (62%)
Frame = +3
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENP 48
P PPPPPPPPP P PHFS +P
Sbjct: 30 PPSPPPPPPPPPPPSFPPHFSPIFFSP 110
Score = 30.0 bits (66), Expect = 2.9
Identities = 13/25 (52%), Positives = 14/25 (56%), Gaps = 5/25 (20%)
Frame = +1
Query: 22 PEQPPPPP-----PPPPNPQPDPHF 41
P +PP PP PPP P P PHF
Sbjct: 4 PPRPPAPPFPLPLLPPPPPLPPPHF 78
>TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana tabacum},
partial (13%)
Length = 663
Score = 41.2 bits (95), Expect = 0.001
Identities = 43/168 (25%), Positives = 87/168 (51%), Gaps = 14/168 (8%)
Frame = +3
Query: 310 VEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHN 369
VE+E + E V+ ++E+ E K ++E++D K +VD D K +E + +E +E N
Sbjct: 132 VEIEKDLEIKSVEKDDEKKE-KKDKEKKD----KTDVDEGKDKK---DKEKKKKEKKEEN 287
Query: 370 I-----ELSLGQDKVETLPVEKEQGEGEQMMDFEQ--SKKEETEMWFLGQKNYVGEPSLR 422
+ + +DK + +KE+G+ ++ D E+ SKK++ + + + GE +
Sbjct: 288 VKGEEEDGDEKKDKEKKKKEKKEKGKEDKDKDGEEKKSKKDKEKKKDKNEDDDEGEDGSK 467
Query: 423 PCHNRDRKGIDCEQVKEDEG-------EEEEHEQEEEEEDDVEEDEHD 463
N+D+K E+ +++EG + EE +E +E+ +ED+ +
Sbjct: 468 KKKNKDKKEKKKEEDEKEEGKVSVRDIDIEETAKEGKEKKKKKEDKEE 611
>AL368332 weakly similar to GP|21322711|e pherophorin-dz1 protein {Volvox
carteri f. nagariensis}, partial (10%)
Length = 478
Score = 41.2 bits (95), Expect = 0.001
Identities = 28/91 (30%), Positives = 34/91 (36%), Gaps = 7/91 (7%)
Frame = +1
Query: 8 TPNETLNPITTEQCPEQPP------PPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNP 61
+P NP PP PPPPPPP P P S P P P +P
Sbjct: 133 SPFHPFNPPPPHSIKSPPPAPSHSSPPPPPPPRKXPPPPPSHPFSPPPPHNHPPP---SP 303
Query: 62 NHDQEDTLMDEDPTHPQIEDDPEPE-PPSPA 91
+H P+ P + P P PPSP+
Sbjct: 304 HHPIA-------PSPPHVRPSPPPPLPPSPS 375
Score = 31.2 bits (69), Expect = 1.3
Identities = 18/63 (28%), Positives = 21/63 (32%)
Frame = +1
Query: 26 PPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIEDDPEP 85
PPPP P NP P P P P +P + + P P P P
Sbjct: 67 PPPPSHPLNPPPSPFHPIN---------PPPSPFHPFNPPPPHSIKSPPPAPSHSSPPPP 219
Query: 86 EPP 88
PP
Sbjct: 220 PPP 228
Score = 29.6 bits (65), Expect = 3.8
Identities = 12/29 (41%), Positives = 15/29 (51%)
Frame = +1
Query: 9 PNETLNPITTEQCPEQPPPPPPPPPNPQP 37
P +PI +P PPPP PP+P P
Sbjct: 292 PPSPHHPIAPSPPHVRPSPPPPLPPSPSP 378
>TC77900 similar to GP|9759182|dbj|BAB09797.1 ATP-dependent Clp protease
regulatory subunit CLPX {Arabidopsis thaliana}, partial
(61%)
Length = 2144
Score = 40.8 bits (94), Expect = 0.002
Identities = 27/69 (39%), Positives = 31/69 (44%), Gaps = 4/69 (5%)
Frame = -2
Query: 4 PIADTPNETLNPITTEQCPEQP----PPPPPPPPNPQPDPHFSETLENPQIQTIPDPDST 59
PI D P + + PE P PPPPPPPP P ET P TIP PD T
Sbjct: 664 PILDPPQHQ-----SFEFPESPELAAPPPPPPPPPP-------ETS*FPGRTTIPPPDQT 521
Query: 60 NPNHDQEDT 68
+ E+T
Sbjct: 520 SGGSTNEET 494
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.131 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,886,222
Number of Sequences: 36976
Number of extensions: 376599
Number of successful extensions: 17549
Number of sequences better than 10.0: 686
Number of HSP's better than 10.0 without gapping: 4709
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 9722
length of query: 652
length of database: 9,014,727
effective HSP length: 102
effective length of query: 550
effective length of database: 5,243,175
effective search space: 2883746250
effective search space used: 2883746250
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144893.11


