

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144807.8 - phase: 0
(361 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
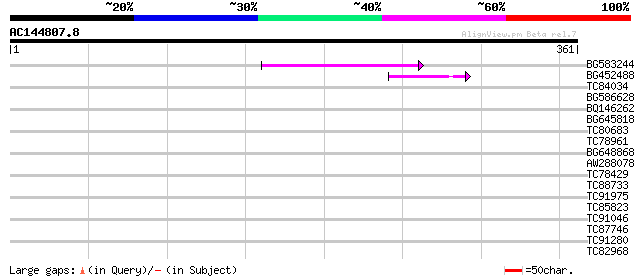
Sequences producing significant alignments: (bits) Value
BG583244 58 5e-09
BG452488 45 4e-05
TC84034 29 0.69
BG586628 similar to GP|6513940|gb| sulfite oxidase (SOX) {Arabid... 31 0.85
BQ146262 30 1.1
BG645818 PIR|S58801|S58 ubiquinol--cytochrome-c reductase (EC 1.... 30 1.5
TC80683 homologue to GP|8777424|dbj|BAA97014.1 gb|AAF56406.1~gen... 28 4.2
TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 28 4.2
BG648868 28 4.2
AW288078 28 7.2
TC78429 similar to GP|16604607|gb|AAL24096.1 putative AtBgamma p... 28 7.2
TC88733 weakly similar to GP|14334730|gb|AAK59543.1 unknown prot... 28 7.2
TC91975 27 9.4
TC85823 weakly similar to GP|13622914|gb|AAK34593.1 hypothetical... 27 9.4
TC91046 similar to GP|9279622|dbj|BAB01080.1 gene_id:MOJ10.6~unk... 27 9.4
TC87746 similar to GP|15021761|gb|AAK77908.1 AAA-metalloprotease... 27 9.4
TC91280 similar to GP|15982807|gb|AAL09751.1 At2g15290/F27O10.6 ... 27 9.4
TC82968 weakly similar to PIR|T04972|T04972 hypothetical protein... 27 9.4
>BG583244
Length = 633
Score = 58.2 bits (139), Expect = 5e-09
Identities = 33/104 (31%), Positives = 49/104 (46%), Gaps = 1/104 (0%)
Frame = -1
Query: 161 DARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIG-TVLASELYEYPDKKLIIKIKVN 219
D ++ IWVQ LP +G +GS I V+ E Y + +KI+V
Sbjct: 312 DHMAIPMNTIDIWVQDHQLPFGFMDTTIGALVGSHIC*MVIFDEENNYGPWRKYVKIRVE 133
Query: 220 LAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFCFACGLIGHTE 263
+ + P+K + + W+ F+YE L +FCF CG IGH E
Sbjct: 132 IEIEQPLKQDLILEGDMCKNIWLVFKYEKLDKFCFVCGSIGHRE 1
>BG452488
Length = 643
Score = 45.1 bits (105), Expect = 4e-05
Identities = 20/52 (38%), Positives = 31/52 (59%)
Frame = +1
Query: 242 IDFRYENLPQFCFACGLIGHTEANCKLHDTRTNAESRNKNVLGPWLRYNHFG 293
++FRYE FC+ CGLIGHT+ +C + E + + GP+LR ++ G
Sbjct: 37 VNFRYEKPGNFCYECGLIGHTDGSCPKRFEKGFVEGQQQ--WGPYLRSDYVG 186
>TC84034
Length = 485
Score = 28.9 bits (63), Expect(2) = 0.69
Identities = 15/46 (32%), Positives = 22/46 (47%)
Frame = -2
Query: 156 WRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLA 201
W R ++ HA IWV+L P + K+ +I +GT LA
Sbjct: 349 WSRGFKPQAQVRTHAQIWVRLMHFPQEYWRKRTLFEIAYGLGTPLA 212
Score = 20.8 bits (42), Expect(2) = 0.69
Identities = 7/12 (58%), Positives = 8/12 (66%)
Frame = -1
Query: 242 IDFRYENLPQFC 253
ID +YE P FC
Sbjct: 92 IDIQYEKQPPFC 57
>BG586628 similar to GP|6513940|gb| sulfite oxidase (SOX) {Arabidopsis
thaliana}, partial (35%)
Length = 548
Score = 30.8 bits (68), Expect = 0.85
Identities = 17/46 (36%), Positives = 22/46 (46%), Gaps = 2/46 (4%)
Frame = -2
Query: 302 RKFSSNPTKCKNYGHFMPPIPPKMLEEMAKM--SMLNHIKDHPQNH 345
R SNP C H PPIP + E + +M S N K+ +NH
Sbjct: 283 RNIPSNPFHCPRLAHSCPPIPCTLKEFICRMK*SSENLCKNTKKNH 146
>BQ146262
Length = 687
Score = 30.4 bits (67), Expect = 1.1
Identities = 12/36 (33%), Positives = 23/36 (63%), Gaps = 1/36 (2%)
Frame = -3
Query: 246 YENLPQFCFACGLIGHTEANC-KLHDTRTNAESRNK 280
YE P FC+ C L+GH+ C ++++ + +A +N+
Sbjct: 478 YERKPSFCYNCKLLGHSIQLCNRINNIQHHAAPKNQ 371
>BG645818 PIR|S58801|S58 ubiquinol--cytochrome-c reductase (EC 1.10.2.2)
cytochrome b - Leptothorax acervorum mitochondrion,
partial (10%)
Length = 701
Score = 30.0 bits (66), Expect = 1.5
Identities = 14/39 (35%), Positives = 19/39 (47%)
Frame = +2
Query: 242 IDFRYENLPQFCFACGLIGHTEANCKLHDTRTNAESRNK 280
ID +YE P F C ++GH NC+ + N E K
Sbjct: 155 IDIQYEK*PPF*VNCKVLGHNLQNCRKLNGSNNTEGSVK 271
>TC80683 homologue to GP|8777424|dbj|BAA97014.1
gb|AAF56406.1~gene_id:K9P8.7~strong similarity to unknown
protein {Arabidopsis thaliana}, partial (16%)
Length = 1360
Score = 28.5 bits (62), Expect = 4.2
Identities = 12/39 (30%), Positives = 18/39 (45%), Gaps = 6/39 (15%)
Frame = +1
Query: 250 PQFCFACGLIGHTEANCK------LHDTRTNAESRNKNV 282
P+ C+ C +GH +CK LH T+ N N+
Sbjct: 1069 PKICYKCKKVGHLSRDCKEQPNDLLHSHATSEAEENPNM 1185
>TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (71%)
Length = 974
Score = 28.5 bits (62), Expect = 4.2
Identities = 13/41 (31%), Positives = 20/41 (48%), Gaps = 1/41 (2%)
Frame = +2
Query: 240 HWI-DFRYENLPQFCFACGLIGHTEANCKLHDTRTNAESRN 279
HW D + + C+ CG GH E NCK + + +R+
Sbjct: 383 HWARDCKAGDWKNKCYRCGERGHIEKNCKNSPKKLSRHARS 505
>BG648868
Length = 801
Score = 28.5 bits (62), Expect = 4.2
Identities = 10/24 (41%), Positives = 16/24 (66%)
Frame = -2
Query: 245 RYENLPQFCFACGLIGHTEANCKL 268
++E + F F CGL+GHT+ + L
Sbjct: 797 KHEKIGDFYFLCGLLGHTDKSFSL 726
>AW288078
Length = 494
Score = 27.7 bits (60), Expect = 7.2
Identities = 14/34 (41%), Positives = 20/34 (58%), Gaps = 1/34 (2%)
Frame = +3
Query: 2 WPISPN-NPYPTQNSTQINNKLMKNYLCSLPLMK 34
W SP+ YP ST IN+K+ + +C L L+K
Sbjct: 84 WKKSPS*RRYPLA*STCINSKIFTSLICQLNLLK 185
>TC78429 similar to GP|16604607|gb|AAL24096.1 putative AtBgamma protein
{Arabidopsis thaliana}, partial (62%)
Length = 1596
Score = 27.7 bits (60), Expect = 7.2
Identities = 13/39 (33%), Positives = 24/39 (61%)
Frame = +2
Query: 70 YDDSDIQERLEECQNSILGKIVSEKAIHRNSIQNALSNI 108
+D D +ER +C ++L +I + +HR I+ ++SNI
Sbjct: 1040 FDSEDPRER--DCLKTVLHRIYGKFMVHRPFIRKSVSNI 1150
>TC88733 weakly similar to GP|14334730|gb|AAK59543.1 unknown protein
{Arabidopsis thaliana}, partial (27%)
Length = 986
Score = 27.7 bits (60), Expect = 7.2
Identities = 16/31 (51%), Positives = 19/31 (60%), Gaps = 1/31 (3%)
Frame = +2
Query: 329 MAKMSML-NHIKDHPQNHNTNEEHTKTTKKV 358
+ KMSML IKDH Q N ++ TKT K V
Sbjct: 758 LRKMSMLLKKIKDHVQIENLVKDDTKTAKGV 850
>TC91975
Length = 724
Score = 27.3 bits (59), Expect = 9.4
Identities = 14/43 (32%), Positives = 19/43 (43%)
Frame = +3
Query: 249 LPQFCFACGLIGHTEANCKLHDTRTNAESRNKNVLGPWLRYNH 291
+P FCF L + C +H T + RN L+YNH
Sbjct: 339 IPFFCFRLNLF*SIVSTCGIHSTHKSINGRNT----IQLQYNH 455
>TC85823 weakly similar to GP|13622914|gb|AAK34593.1 hypothetical protein
{Streptococcus pyogenes M1 GAS}, partial (18%)
Length = 813
Score = 27.3 bits (59), Expect = 9.4
Identities = 14/43 (32%), Positives = 19/43 (43%), Gaps = 3/43 (6%)
Frame = -2
Query: 3 PISPNNPYPTQ---NSTQINNKLMKNYLCSLPLMKMHHQNAIT 42
PI NP PT N ++ + + Y C P + HH A T
Sbjct: 518 PIHRENPKPTMASNNEAEVYTSIEQAYPCLTPELAKHH*KATT 390
>TC91046 similar to GP|9279622|dbj|BAB01080.1 gene_id:MOJ10.6~unknown
protein {Arabidopsis thaliana}, partial (6%)
Length = 978
Score = 27.3 bits (59), Expect = 9.4
Identities = 10/26 (38%), Positives = 17/26 (64%)
Frame = +1
Query: 328 EMAKMSMLNHIKDHPQNHNTNEEHTK 353
E A+ S+L+ +KD Q H + +EH +
Sbjct: 49 EAARASLLSQLKDALQEHESKQEHVR 126
>TC87746 similar to GP|15021761|gb|AAK77908.1 AAA-metalloprotease FtsH
{Pisum sativum}, partial (49%)
Length = 1274
Score = 27.3 bits (59), Expect = 9.4
Identities = 16/40 (40%), Positives = 18/40 (45%), Gaps = 8/40 (20%)
Frame = +2
Query: 288 RYNHFGRRIMDYKD--------RKFSSNPTKCKNYGHFMP 319
R N FG + D+K R FSS K KNY F P
Sbjct: 245 RNNGFGSNLYDFKSIAANRMLHRMFSSESPKKKNYEKFYP 364
>TC91280 similar to GP|15982807|gb|AAL09751.1 At2g15290/F27O10.6
{Arabidopsis thaliana}, partial (61%)
Length = 818
Score = 27.3 bits (59), Expect = 9.4
Identities = 10/20 (50%), Positives = 15/20 (75%)
Frame = +2
Query: 27 LCSLPLMKMHHQNAITSQDP 46
+C++ L+KM+H NA QDP
Sbjct: 629 ICAMLLIKMNHDNASAIQDP 688
>TC82968 weakly similar to PIR|T04972|T04972 hypothetical protein T16L1.40 -
Arabidopsis thaliana, partial (16%)
Length = 507
Score = 27.3 bits (59), Expect = 9.4
Identities = 22/64 (34%), Positives = 28/64 (43%), Gaps = 11/64 (17%)
Frame = -2
Query: 98 RNSIQNALSNIW-------CNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGN----PWFF 146
R+S ++ L + W CN F H KLFHFF DE T I + PW +
Sbjct: 404 RHSNRSTLHSCWNFQSARPCNRYCFLNTH---KLFHFFSDEITSTWDIKVSHHCTTPW*W 234
Query: 147 RNSW 150
N W
Sbjct: 233 WNVW 222
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.135 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,583,175
Number of Sequences: 36976
Number of extensions: 216171
Number of successful extensions: 1172
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 1161
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1172
length of query: 361
length of database: 9,014,727
effective HSP length: 97
effective length of query: 264
effective length of database: 5,428,055
effective search space: 1433006520
effective search space used: 1433006520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144807.8


