

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
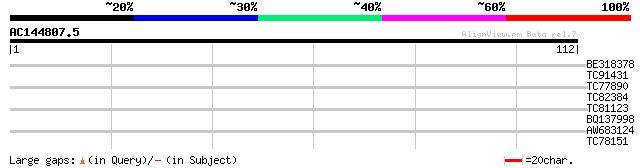
Query= AC144807.5 + phase: 0
(112 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BE318378 similar to GP|6681351|dbj| ETAG-A3 {Lycopersicon escule... 26 2.0
TC91431 similar to GP|10177753|dbj|BAB11066. gene_id:MDN11.10~un... 26 2.6
TC77890 similar to GP|20466476|gb|AAM20555.1 alpha-mannosidase {... 26 2.6
TC82384 similar to PIR|T51591|T51591 dimethyladenosine transfera... 25 3.4
TC81123 similar to PIR|E96789|E96789 protein T23E18.10 [imported... 25 4.5
BQ137998 GP|6165483|emb putative nuclear transport receptor {Sch... 25 5.9
AW683124 homologue to PIR|E96600|E96 protein F14J16.20 [imported... 25 5.9
TC78151 similar to GP|22655006|gb|AAM98094.1 At1g76400/F15M4_10 ... 25 5.9
>BE318378 similar to GP|6681351|dbj| ETAG-A3 {Lycopersicon esculentum},
partial (21%)
Length = 654
Score = 26.2 bits (56), Expect = 2.0
Identities = 11/28 (39%), Positives = 15/28 (53%)
Frame = -2
Query: 40 QKIIEALEAEKDSLTKTNKSIRNTLWDW 67
QK+I A +S TK+ K + LW W
Sbjct: 638 QKVINAKIVHSNSPTKSKKPVFTNLWMW 555
>TC91431 similar to GP|10177753|dbj|BAB11066. gene_id:MDN11.10~unknown
protein {Arabidopsis thaliana}, partial (86%)
Length = 1047
Score = 25.8 bits (55), Expect = 2.6
Identities = 15/49 (30%), Positives = 23/49 (46%)
Frame = +1
Query: 41 KIIEALEAEKDSLTKTNKSIRNTLWDWKQQLYAEAASDSAPVPLARLYE 89
+ I+A+ A K+ + +LW L+A ASD+ PL R E
Sbjct: 853 RTIDAINAAKEQ--------KRSLWRVSSTLFAHIASDAVLKPLGRFAE 975
>TC77890 similar to GP|20466476|gb|AAM20555.1 alpha-mannosidase {Arabidopsis
thaliana}, partial (55%)
Length = 2422
Score = 25.8 bits (55), Expect = 2.6
Identities = 14/50 (28%), Positives = 23/50 (46%)
Frame = +3
Query: 9 DIAKRASENNTVINVGLGAVFAILGARSYNQQKIIEALEAEKDSLTKTNK 58
DIA R+ + VGLG + + + + I K+SL +T+K
Sbjct: 849 DIAHRSGNESDAFEVGLGNLKLVYSRKEGKLTQYINRKRKVKESLEQTHK 998
>TC82384 similar to PIR|T51591|T51591 dimethyladenosine transferase (EC
2.1.1.-) PFC1 [validated] - Arabidopsis thaliana,
partial (78%)
Length = 1138
Score = 25.4 bits (54), Expect = 3.4
Identities = 12/35 (34%), Positives = 16/35 (45%)
Frame = +2
Query: 38 NQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLY 72
N Q + A K +T I +LW WKQQ +
Sbjct: 68 NLQLPPQTATAAKSYRRQTESKILRSLWPWKQQSF 172
>TC81123 similar to PIR|E96789|E96789 protein T23E18.10 [imported] -
Arabidopsis thaliana, partial (68%)
Length = 1242
Score = 25.0 bits (53), Expect = 4.5
Identities = 10/35 (28%), Positives = 21/35 (59%)
Frame = -2
Query: 38 NQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLY 72
N++++I+A+E + + ++ R LW WK +Y
Sbjct: 254 NKERLIQAIENTNNQVIISD---RFNLWSWKLSIY 159
>BQ137998 GP|6165483|emb putative nuclear transport receptor
{Schizosaccharomyces pombe}, partial (11%)
Length = 656
Score = 24.6 bits (52), Expect = 5.9
Identities = 13/37 (35%), Positives = 21/37 (56%), Gaps = 5/37 (13%)
Frame = -2
Query: 5 HKIVDIAKRASENNTVINVGLGAV-----FAILGARS 36
H ++ IAK A + ++V +GLG F ++ ARS
Sbjct: 286 HSVICIAKLAKQTHSVNRIGLGPA*KTVRFLMMEARS 176
>AW683124 homologue to PIR|E96600|E96 protein F14J16.20 [imported] -
Arabidopsis thaliana, partial (11%)
Length = 399
Score = 24.6 bits (52), Expect = 5.9
Identities = 14/52 (26%), Positives = 24/52 (45%)
Frame = +3
Query: 35 RSYNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLYAEAASDSAPVPLAR 86
R + +Q ++ AL + ++ TLW WK + AE S++ L R
Sbjct: 66 RLFKEQGLLLALLLGLSAFFSLAETSITTLWPWKVRELAEKESENGVFRLLR 221
>TC78151 similar to GP|22655006|gb|AAM98094.1 At1g76400/F15M4_10
{Arabidopsis thaliana}, partial (88%)
Length = 2171
Score = 24.6 bits (52), Expect = 5.9
Identities = 9/17 (52%), Positives = 11/17 (63%)
Frame = -2
Query: 55 KTNKSIRNTLWDWKQQL 71
K N S+RN LW W Q +
Sbjct: 526 KLNVSLRNFLWKWLQYM 476
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.131 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,881,999
Number of Sequences: 36976
Number of extensions: 26212
Number of successful extensions: 125
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 125
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 125
length of query: 112
length of database: 9,014,727
effective HSP length: 88
effective length of query: 24
effective length of database: 5,760,839
effective search space: 138260136
effective search space used: 138260136
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 50 (23.9 bits)
Medicago: description of AC144807.5


