

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144766.6 - phase: 0
(381 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
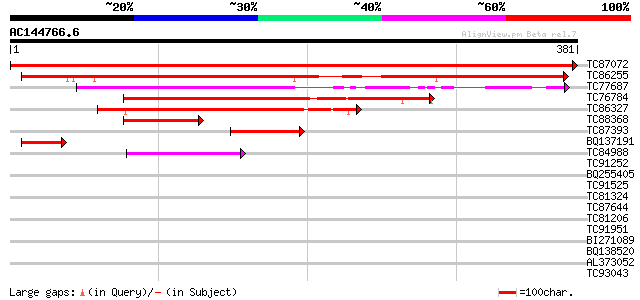
Score E
Sequences producing significant alignments: (bits) Value
TC87072 similar to PIR|D84581|D84581 probable CCCH-type zinc fin... 805 0.0
TC86255 similar to PIR|D84581|D84581 probable CCCH-type zinc fin... 453 e-128
TC77687 similar to PIR|D84581|D84581 probable CCCH-type zinc fin... 247 6e-66
TC76784 similar to PIR|G84825|G84825 probable CCCH-type zinc fin... 200 9e-52
TC86327 similar to GP|14335106|gb|AAK59832.1 At2g41900/T6D20.20 ... 192 2e-49
TC88368 similar to GP|15810487|gb|AAL07131.1 unknown protein {Ar... 84 1e-16
TC87393 similar to PIR|G84825|G84825 probable CCCH-type zinc fin... 54 1e-07
BQ137191 44 1e-04
TC84988 similar to GP|14018368|emb|CAC38358. zinc finger protein... 41 7e-04
TC91252 similar to GP|9988428|dbj|BAB12694.1 putative zinc finge... 40 0.001
BQ255405 similar to GP|9988428|dbj| putative zinc finger transcr... 39 0.003
TC91525 similar to GP|14335106|gb|AAK59832.1 At2g41900/T6D20.20 ... 38 0.007
TC81324 similar to GP|21553871|gb|AAM62964.1 unknown {Arabidopsi... 37 0.017
TC87644 similar to GP|10177307|dbj|BAB10568. contains similarity... 35 0.037
TC81206 similar to GP|8777433|dbj|BAA97023.1 gene_id:MHM17.4~unk... 35 0.048
TC91951 similar to GP|5441893|dbj|BAA82391.1 ESTs C99174(E10437)... 34 0.082
BI271089 similar to GP|9294325|dbj| gene_id:K24M9.13~unknown pro... 34 0.082
BQ138520 similar to SP|O59953|RL5_ 60S ribosomal protein L5 (CPR... 33 0.14
AL373052 similar to GP|21593014|gb| putative aldolase {Arabidops... 32 0.31
TC93043 weakly similar to PIR|C84918|C84918 probable ATP-depende... 32 0.41
>TC87072 similar to PIR|D84581|D84581 probable CCCH-type zinc finger protein
[imported] - Arabidopsis thaliana, partial (47%)
Length = 1599
Score = 805 bits (2080), Expect = 0.0
Identities = 381/381 (100%), Positives = 381/381 (100%)
Frame = +3
Query: 1 MMLGEPPHRTNPTVHVPPWPTLNNPTAEIFSPLTSNDDYSQFYMQEALSAFQHYVNENND 60
MMLGEPPHRTNPTVHVPPWPTLNNPTAEIFSPLTSNDDYSQFYMQEALSAFQHYVNENND
Sbjct: 27 MMLGEPPHRTNPTVHVPPWPTLNNPTAEIFSPLTSNDDYSQFYMQEALSAFQHYVNENND 206
Query: 61 SDSDSEIFPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKY 120
SDSDSEIFPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKY
Sbjct: 207 SDSDSEIFPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKY 386
Query: 121 NYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTE 180
NYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTE
Sbjct: 387 NYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTE 566
Query: 181 QLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRS 240
QLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRS
Sbjct: 567 QLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRS 746
Query: 241 LGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSPRGPVIRPGFCSLPSTP 300
LGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSPRGPVIRPGFCSLPSTP
Sbjct: 747 LGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSPRGPVIRPGFCSLPSTP 926
Query: 301 TQVPSRGRVNHFDLWDQSCEEEPVMERVESGRDIRVKMFEKLSKENSFNGSGMGSGSGLG 360
TQVPSRGRVNHFDLWDQSCEEEPVMERVESGRDIRVKMFEKLSKENSFNGSGMGSGSGLG
Sbjct: 927 TQVPSRGRVNHFDLWDQSCEEEPVMERVESGRDIRVKMFEKLSKENSFNGSGMGSGSGLG 1106
Query: 361 EVVEDPDVGWVSELVSPFLGD 381
EVVEDPDVGWVSELVSPFLGD
Sbjct: 1107EVVEDPDVGWVSELVSPFLGD 1169
>TC86255 similar to PIR|D84581|D84581 probable CCCH-type zinc finger protein
[imported] - Arabidopsis thaliana, partial (47%)
Length = 1508
Score = 453 bits (1166), Expect = e-128
Identities = 240/400 (60%), Positives = 273/400 (68%), Gaps = 33/400 (8%)
Frame = +3
Query: 9 RTNPTVHVPPWPTLNNPTAEIFSPLTSND----------------------DYSQ--FYM 44
R NPTVHVPPWP ++PTAEI S D DYS +Y+
Sbjct: 48 RANPTVHVPPWPNHDDPTAEILSSCFGYDNNNIAGDFTGAGDYTAGDFSTGDYSSGDYYL 227
Query: 45 QEALSAFQHYV--NENNDSDSDSEIFPTHES-VDSYSNDHFRMFEFKIRRCARGRSHDWT 101
+EAL+A Q Y+ NE NDSD +S +S VD+YS DHFRM+EFKIRRCARGRSHDWT
Sbjct: 228 REALAALQRYLPSNEFNDSDLESPATAAADSTVDAYSCDHFRMYEFKIRRCARGRSHDWT 407
Query: 102 ECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQP 161
ECP++HPGEKARRRDPRK++YSGT+CPDFRKG+CKKGD+CE AHGVFECWLHP+RYRTQP
Sbjct: 408 ECPYAHPGEKARRRDPRKFHYSGTACPDFRKGNCKKGDACEHAHGVFECWLHPARYRTQP 587
Query: 162 CKDGTSCRRPVCFFAHTTEQLRAPTQQSP---RSVPSVDSYDGSPLRLAFESSCVKTLQF 218
CKDGTSCRR VCFFAHT EQLR QQSP RSV S DSYDGSPLR
Sbjct: 588 CKDGTSCRRRVCFFAHTPEQLRLNVQQSPSSSRSVNSPDSYDGSPLRQMV---------- 737
Query: 219 MSSPGSVSPPVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGS 278
+++ P ESPPMSPM +EMVASLRNLQLG MK++P + NV +GS
Sbjct: 738 -----TITTPPESPPMSPM------------ASEMVASLRNLQLGRMKTMPINRNVTIGS 866
Query: 279 PRFGSPR---GPVIRPGFCSLPSTPTQVPSRGRVNHFDLWDQSCEEEPVMERVESGRDIR 335
P FGS G +R GF SLP+TPT+ P GRV FDLWDQSCEEEP+MERVESGRDIR
Sbjct: 867 PVFGSGSPVFGSPLRSGFLSLPNTPTKKPGLGRVGGFDLWDQSCEEEPIMERVESGRDIR 1046
Query: 336 VKMFEKLSKENSFNGSGMGSGSGLGEVVEDPDVGWVSELV 375
KMFEKLSKENS SG GL PDVGWV +L+
Sbjct: 1047AKMFEKLSKENSLENENGNSGLGLESGQPGPDVGWVCDLL 1166
>TC77687 similar to PIR|D84581|D84581 probable CCCH-type zinc finger protein
[imported] - Arabidopsis thaliana, partial (35%)
Length = 1096
Score = 247 bits (630), Expect = 6e-66
Identities = 147/332 (44%), Positives = 184/332 (55%), Gaps = 1/332 (0%)
Frame = +3
Query: 46 EALSAFQHYVNENNDSDSDSEIFPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPF 105
E ++ H+++ + D + + +S D FRMFEFK+R+C RGRSHDWT+CP+
Sbjct: 90 EEVTFSNHFISNDIDIHTSCMTEDFDLPIHVFSTDQFRMFEFKVRKCQRGRSHDWTDCPY 269
Query: 106 SHPGEKARRRDPRKYNYSGTSCPDFRK-GSCKKGDSCEFAHGVFECWLHPSRYRTQPCKD 164
SHPGEKARRRDP+KYNYSG CP+FRK G+C KGDSC FAHGVFECWLHPSRYRTQ C D
Sbjct: 270 SHPGEKARRRDPQKYNYSGNPCPEFRKLGNCTKGDSCHFAHGVFECWLHPSRYRTQLCND 449
Query: 165 GTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGS 224
GT CRR VCFFAHT +QLR SP S F+SSP S
Sbjct: 450 GTLCRRRVCFFAHTIDQLRVSNNASPES-------------------------FVSSPTS 554
Query: 225 VSPPVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSP 284
V ++S P R V +V E+V +R++++ + +MGS FGSP
Sbjct: 555 V---LDSSP-----RKSRYGVPPVNVRELVGFMRSVRVDEWSPVS-----KMGSV-FGSP 692
Query: 285 RGPVIRPGFCSLPSTPTQVPSRGRVNHFDLWDQSCEEEPVMERVESGRDIRVKMFEKLSK 344
R R GF SLPS E MERVESGRD+R K++EK +
Sbjct: 693 RP---RGGFLSLPSN--------------------YEGVGMERVESGRDLRAKIYEKFGR 803
Query: 345 ENSFNGSGMGSGSGLGEVVEDPDVGWVSELVS 376
NS +G V PD+GWV+ELV+
Sbjct: 804 LNSNDGV---------VSVPVPDIGWVAELVN 872
>TC76784 similar to PIR|G84825|G84825 probable CCCH-type zinc finger protein
[imported] - Arabidopsis thaliana, partial (56%)
Length = 2631
Score = 200 bits (508), Expect = 9e-52
Identities = 103/213 (48%), Positives = 134/213 (62%), Gaps = 4/213 (1%)
Frame = +3
Query: 77 YSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGSCK 136
Y +D FRM+ FK++ C+R SHDWTECPF HPGE ARRRDPRKY YS CP+FRKGSC+
Sbjct: 852 YGSDEFRMYSFKVKPCSRAYSHDWTECPFVHPGENARRRDPRKYPYSCVPCPEFRKGSCQ 1031
Query: 137 KGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSV 196
KGDSCE+AHGVFE WLHP++YRT+ CKD T C R VCFFAH E+LR + ++PS
Sbjct: 1032 KGDSCEYAHGVFESWLHPAQYRTRLCKDETGCNRKVCFFAHRPEELRPVYASTGSAMPSP 1211
Query: 197 DSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRSLGRSVGSSSVNEMVAS 256
SY S + + + L SS +S V +PPMSP+ S G+ N++ +
Sbjct: 1212 KSYSAS----GMDMTSMSPLSLSSSSLPMS-TVSTPPMSPLAGSSSPKSGNMWQNKLNLT 1376
Query: 257 LRNLQL--GTMKSLPSSWNVQMGSPRFG--SPR 285
+LQL +KS S+ ++ + G SPR
Sbjct: 1377 PPSLQLPGSRLKSALSARDLDLEMELLGLDSPR 1475
>TC86327 similar to GP|14335106|gb|AAK59832.1 At2g41900/T6D20.20 {Arabidopsis
thaliana}, partial (50%)
Length = 3018
Score = 192 bits (488), Expect = 2e-49
Identities = 92/186 (49%), Positives = 120/186 (64%), Gaps = 9/186 (4%)
Frame = +2
Query: 60 DSDSDSEIFPTHESVDS-----YSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARR 114
+S S+ + +P S+ Y+ D FRM+ FK+R C+R SHDWTECPF HPGE ARR
Sbjct: 962 NSTSEKKEYPVDPSLPDIKNSMYATDEFRMYSFKVRPCSRAYSHDWTECPFVHPGENARR 1141
Query: 115 RDPRKYNYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCF 174
RDPRK++YS CPDFRKG+C++ D CE+AHGVFECWLHP++YRT+ CKDG C R VCF
Sbjct: 1142 RDPRKFHYSCVPCPDFRKGACRRSDMCEYAHGVFECWLHPAQYRTRLCKDGMGCNRRVCF 1321
Query: 175 FAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVS----PPVE 230
FAH+ E+LR + +VPS S + + + +L F SP S+S P
Sbjct: 1322 FAHSPEELRPLYVSTGSAVPSPRS--AASTANVMDMAAAMSL-FPGSPSSISLMSQSPFA 1492
Query: 231 SPPMSP 236
PP+SP
Sbjct: 1493 QPPLSP 1510
>TC88368 similar to GP|15810487|gb|AAL07131.1 unknown protein {Arabidopsis
thaliana}, partial (25%)
Length = 665
Score = 83.6 bits (205), Expect = 1e-16
Identities = 33/54 (61%), Positives = 41/54 (75%)
Frame = +3
Query: 77 YSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDF 130
+ D FRM+ FK++ C+RG +HDWTECPF HPGE ARRRDPRK+ YS P+F
Sbjct: 504 FVTDEFRMYSFKVKTCSRGYTHDWTECPFVHPGENARRRDPRKFPYSCVPXPEF 665
>TC87393 similar to PIR|G84825|G84825 probable CCCH-type zinc finger protein
[imported] - Arabidopsis thaliana, partial (16%)
Length = 975
Score = 53.5 bits (127), Expect = 1e-07
Identities = 25/50 (50%), Positives = 32/50 (64%)
Frame = +1
Query: 149 ECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSVDS 198
E LHPS+YRT+ CKD C R VCFFAH E+LR + ++PS +S
Sbjct: 1 ESLLHPSQYRTRLCKDEIRCTRKVCFFAHKHEELRPLYASTGSAMPSQES 150
>BQ137191
Length = 970
Score = 43.5 bits (101), Expect = 1e-04
Identities = 19/30 (63%), Positives = 22/30 (73%)
Frame = +3
Query: 9 RTNPTVHVPPWPTLNNPTAEIFSPLTSNDD 38
R NPTVHVPPWP ++PTAEI S S D+
Sbjct: 54 RANPTVHVPPWPNHDDPTAEILSS*FSYDN 143
>TC84988 similar to GP|14018368|emb|CAC38358. zinc finger protein {Pisum
sativum}, partial (28%)
Length = 656
Score = 41.2 bits (95), Expect = 7e-04
Identities = 24/81 (29%), Positives = 33/81 (40%), Gaps = 1/81 (1%)
Frame = +1
Query: 79 NDHFRMFEFKIRRCARGRSH-DWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGSCKK 137
NDHF FK R A CP S + R + C FR G+C+
Sbjct: 358 NDHFEPHSFKRARIADNNPPPSGLMCPPSRMIQPPPNRGTSSIFFKTRICTKFRFGTCRN 537
Query: 138 GDSCEFAHGVFECWLHPSRYR 158
G++C +AHG E P ++
Sbjct: 538 GENCNYAHGADEIRQPPRNWQ 600
>TC91252 similar to GP|9988428|dbj|BAB12694.1 putative zinc finger
transcription factor {Oryza sativa (japonica
cultivar-group)}, partial (9%)
Length = 655
Score = 40.4 bits (93), Expect = 0.001
Identities = 30/99 (30%), Positives = 46/99 (46%), Gaps = 4/99 (4%)
Frame = +1
Query: 169 RRPVCFFAHTTEQLRA----PTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGS 224
+R +CFFAHT QLR P SP + P ++ +S T +S P
Sbjct: 25 KRKICFFAHTPRQLRVLLLPPLSPSP-TPPQKNNKKCCSFCHCCSNSSSPTSTLLSGPHF 201
Query: 225 VSPPVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLG 263
S SPP+SP++ S + V +NE++ S+ + G
Sbjct: 202 SSS--NSPPLSPLSNSNSKDV--HVLNELIRSMESFNFG 306
>BQ255405 similar to GP|9988428|dbj| putative zinc finger transcription
factor {Oryza sativa (japonica cultivar-group)}, partial
(5%)
Length = 269
Score = 39.3 bits (90), Expect = 0.003
Identities = 20/51 (39%), Positives = 32/51 (62%), Gaps = 3/51 (5%)
Frame = +1
Query: 46 EALSAFQHYVNENNDSDSDSEIFPTHES---VDSYSNDHFRMFEFKIRRCA 93
EAL Q + E++ ++S+ E T E+ VD+Y+ DH RM+E+ +RCA
Sbjct: 112 EALEELQRSI*EDDFNESELECHATEEAD*TVDAYACDHDRMYEYTTKRCA 264
>TC91525 similar to GP|14335106|gb|AAK59832.1 At2g41900/T6D20.20
{Arabidopsis thaliana}, partial (18%)
Length = 672
Score = 37.7 bits (86), Expect = 0.007
Identities = 28/78 (35%), Positives = 37/78 (46%)
Frame = +2
Query: 173 CFFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESP 232
CFFAHT E+LR + +VPS S S + A S + SS +SP +P
Sbjct: 2 CFFAHTPEELRPLYVSTGSAVPSPRSSTSSAMDFAAAMSMLPGSP--SSMSVMSPSPFTP 175
Query: 233 PMSPMTRSLGRSVGSSSV 250
PMSP G + +SV
Sbjct: 176 PMSPS----GNGISHNSV 217
>TC81324 similar to GP|21553871|gb|AAM62964.1 unknown {Arabidopsis
thaliana}, partial (75%)
Length = 1156
Score = 36.6 bits (83), Expect = 0.017
Identities = 19/48 (39%), Positives = 22/48 (45%), Gaps = 2/48 (4%)
Frame = +1
Query: 104 PFSHPGEKARRRDPRKY--NYSGTSCPDFRKGSCKKGDSCEFAHGVFE 149
P S+ G P + N C +F KGSC GD C FAHG E
Sbjct: 841 PKSNQGPPGHHGAPGHHGNNLKTKLCENFAKGSCTFGDRCHFAHGAVE 984
Score = 27.7 bits (60), Expect = 7.7
Identities = 10/15 (66%), Positives = 11/15 (72%)
Frame = +1
Query: 135 CKKGDSCEFAHGVFE 149
CK GD C FAHG +E
Sbjct: 436 CKFGDKCHFAHGEWE 480
>TC87644 similar to GP|10177307|dbj|BAB10568. contains similarity to zinc
finger protein~gene_id:MDC12.23 {Arabidopsis thaliana},
partial (36%)
Length = 1541
Score = 35.4 bits (80), Expect = 0.037
Identities = 25/81 (30%), Positives = 33/81 (39%), Gaps = 21/81 (25%)
Frame = +1
Query: 110 EKARRRDPRKYNYSGTSCPDF-RKGSCKKGDSCEFAH--------------------GVF 148
EK R R+ + N T C + R G CK G +C++ H G
Sbjct: 373 EKVREREEPEENAGQTECKYYQRSGGCKFGKACKYNHSRGFTAPISELNFLGLPIRLGER 552
Query: 149 ECWLHPSRYRTQPCKDGTSCR 169
EC P RT CK G++CR
Sbjct: 553 EC---PYYMRTGSCKFGSNCR 606
>TC81206 similar to GP|8777433|dbj|BAA97023.1 gene_id:MHM17.4~unknown
protein {Arabidopsis thaliana}, partial (5%)
Length = 842
Score = 35.0 bits (79), Expect = 0.048
Identities = 23/76 (30%), Positives = 28/76 (36%), Gaps = 18/76 (23%)
Frame = +3
Query: 90 RRCARGRSHDWTECPFSHPGEKARRRDP------RKYNYSGTSCPD------------FR 131
R GR H EC FSH + P + G CP
Sbjct: 120 RHYMNGRCHKGDECNFSHDAIPLTKSKPCVHYAHHRSCMKGNDCPYDHDLFNYPCSNFVS 299
Query: 132 KGSCKKGDSCEFAHGV 147
KGSC +GD+C F+H V
Sbjct: 300 KGSCYRGDACLFSHQV 347
Score = 30.4 bits (67), Expect = 1.2
Identities = 18/65 (27%), Positives = 27/65 (40%), Gaps = 5/65 (7%)
Frame = +3
Query: 109 GEKARRRDPRKYNYSGTSCPDFRKGSCKKGDSCEFAHGVF-----ECWLHPSRYRTQPCK 163
G K + DP + C + G C KGD C F+H + +H + +R+ C
Sbjct: 66 GVKRPKLDPEQ-KPKPKQCRHYMNGRCHKGDECNFSHDAIPLTKSKPCVHYAHHRS--CM 236
Query: 164 DGTSC 168
G C
Sbjct: 237 KGNDC 251
>TC91951 similar to GP|5441893|dbj|BAA82391.1 ESTs C99174(E10437)
D22295(C10709) correspond to a region of the predicted
gene.~Similar to, partial (23%)
Length = 526
Score = 34.3 bits (77), Expect = 0.082
Identities = 13/23 (56%), Positives = 15/23 (64%)
Frame = +3
Query: 127 CPDFRKGSCKKGDSCEFAHGVFE 149
C +F KGSC G+ C FAHG E
Sbjct: 216 CENFAKGSCTFGERCHFAHGAAE 284
>BI271089 similar to GP|9294325|dbj| gene_id:K24M9.13~unknown protein
{Arabidopsis thaliana}, partial (2%)
Length = 481
Score = 34.3 bits (77), Expect = 0.082
Identities = 41/150 (27%), Positives = 53/150 (35%), Gaps = 14/150 (9%)
Frame = +3
Query: 70 THESVD-----SYSNDHFR---MFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYN 121
THES D SND + M EF R GR H + F H G R
Sbjct: 3 THESRDYSLKSGRSNDEYGKVFMDEFTREREV-GRRH--RDNSFEHGGRHVPNRT----- 158
Query: 122 YSGTSCPDFRKGSCKKGDSCEFAHGVFEC-----WLHPSRYRTQPCKDGTSC-RRPVCFF 175
C F G+C+ G C F+H C L R+ P +D RR +
Sbjct: 159 -IDAPCKFFASGNCRNGKHCRFSHDTQACRSPIRRLRDDRWTRNPSRDHQMLDRRKLSDS 335
Query: 176 AHTTEQLRAPTQQSPRSVPSVDSYDGSPLR 205
++LR S + D + SP R
Sbjct: 336 ISPNKRLRDDRWGSDSDMADPDRVEDSPKR 425
>BQ138520 similar to SP|O59953|RL5_ 60S ribosomal protein L5 (CPR4).
{Neurospora crassa}, partial (63%)
Length = 584
Score = 33.5 bits (75), Expect = 0.14
Identities = 40/129 (31%), Positives = 51/129 (39%), Gaps = 4/129 (3%)
Frame = +2
Query: 179 TEQLRAPTQQSPRSVPSVDSYDGSPLRL-AFESSCVKTLQFMSSPGSVSP---PVESPPM 234
T +R T S S S SPLRL A SS ++TL SS SP P PPM
Sbjct: 101 TSTMRPSTASSSASPTRTSSLRLSPLRLVATRSSWLRTLT--SSSPMASPTV*PTGLPPM 274
Query: 235 SPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSPRGPVIRPGFC 294
+ S S SSS + LR + T P V+ +P S + P
Sbjct: 275 PLVFCSPAVSSRSSSSTRLSPVLRRPMVSTPSPRPPRLTVRSAAPSRSSLMSVLPAPPPA 454
Query: 295 SLPSTPTQV 303
+ S P +V
Sbjct: 455 PVSSVP*RV 481
>AL373052 similar to GP|21593014|gb| putative aldolase {Arabidopsis
thaliana}, partial (19%)
Length = 459
Score = 32.3 bits (72), Expect = 0.31
Identities = 34/151 (22%), Positives = 61/151 (39%), Gaps = 22/151 (14%)
Frame = +3
Query: 182 LRAPTQQSPRSVPSVDSYDGSPLRLAFESS------------C-VKTLQFMSSPGSVSPP 228
L P+ ++ PS+ S + +PL L +S C ++ ++ S+P +PP
Sbjct: 18 LTPPSLSHHQNQPSLPSLNPTPLPLPISTSNSFPSLPPLTWQCPLERIRHFSNPSPTTPP 197
Query: 229 VESPPM---------SPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSP 279
+SPP +P++ + S SSS S ++ SL S+WN +
Sbjct: 198 SQSPPQPPPHPTATSNPVSVLMKPSTASSSSPSPPLSPKSPPSPATTSLSSTWNTVTAA- 374
Query: 280 RFGSPRGPVIRPGFCSLPSTPTQVPSRGRVN 310
P + P +P Q+P + V+
Sbjct: 375 ------SPTLSP---VSTHSPPQIPQQSSVS 440
>TC93043 weakly similar to PIR|C84918|C84918 probable ATP-dependent RNA
helicase A [imported] - Arabidopsis thaliana, partial
(14%)
Length = 735
Score = 32.0 bits (71), Expect = 0.41
Identities = 17/56 (30%), Positives = 29/56 (51%), Gaps = 4/56 (7%)
Frame = +1
Query: 130 FRKGSCKKGDSCEFAHGVF----ECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQ 181
F +GSC +G+SC F+H + +C + Q C++G S C F+H ++
Sbjct: 7 FMRGSCSRGNSCSFSHTLQAKRPQCKFF---FSLQGCRNGGS-----CLFSHDVDR 150
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.133 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,137,343
Number of Sequences: 36976
Number of extensions: 247366
Number of successful extensions: 1898
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 1669
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1879
length of query: 381
length of database: 9,014,727
effective HSP length: 98
effective length of query: 283
effective length of database: 5,391,079
effective search space: 1525675357
effective search space used: 1525675357
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144766.6


