

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144765.5 - phase: 0 /pseudo
(443 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
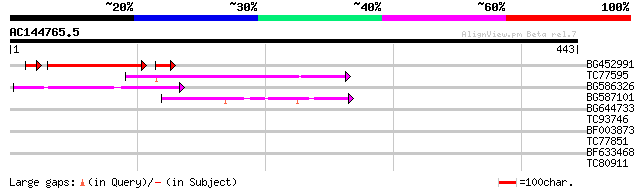
Score E
Sequences producing significant alignments: (bits) Value
BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, pa... 109 2e-27
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 109 2e-24
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 64 9e-11
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 46 3e-05
BG644733 weakly similar to GP|15289942|db putative polyprotein {... 40 0.002
TC93746 weakly similar to GP|22830935|dbj|BAC15800. hypothetical... 35 0.076
BF003873 similar to GP|14715222|em putative polyprotein {Cicer a... 32 0.38
TC77851 similar to GP|15982840|gb|AAL09767.1 AT5g03040/F15A17_70... 30 1.9
BF633468 similar to GP|20259516|gb unknown protein {Arabidopsis ... 28 7.1
TC80911 similar to GP|3668118|emb|CAA11819.1 hypothetical protei... 28 9.3
>BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, partial (33%)
Length = 560
Score = 109 bits (273), Expect(3) = 2e-27
Identities = 54/78 (69%), Positives = 67/78 (85%)
Frame = +3
Query: 30 VKEFELLEQFRDLSLVCELSSQSVQLGMLKINSDFLGSIREAQQVDVKFVDLMVVSNQAE 89
++ + LEQFRDLSLVCE+S QSV+LGMLKIN++FL SI+EAQ+VDVK VDLM +NQ E
Sbjct: 51 LESWSCLEQFRDLSLVCEVSPQSVKLGMLKINNEFLDSIKEAQKVDVKLVDLMFGNNQTE 230
Query: 90 ESDFKVDEQGVLRFRGRI 107
+ DFKVD+QGVL+FR RI
Sbjct: 231 DGDFKVDDQGVLQFRDRI 284
Score = 25.4 bits (54), Expect(3) = 2e-27
Identities = 9/15 (60%), Positives = 14/15 (93%)
Frame = +2
Query: 115 LKKLILEEGHKSNLS 129
+KK+ILEE H+SN++
Sbjct: 284 MKKMILEESHRSNVN 328
Score = 25.4 bits (54), Expect(3) = 2e-27
Identities = 11/13 (84%), Positives = 12/13 (91%)
Frame = +1
Query: 13 VVADALSRKTLHM 25
VVAD LSRKTLH+
Sbjct: 1 VVADVLSRKTLHV 39
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial
(14%)
Length = 1708
Score = 109 bits (272), Expect = 2e-24
Identities = 64/184 (34%), Positives = 96/184 (51%), Gaps = 8/184 (4%)
Frame = +2
Query: 91 SDFKVDEQGVLRFRGRICIPDNE-------ELKKLILEEGHKSNLSIHLGATKMYQDLKK 143
S+ ++D L FRGRI +P ++ EL+ +++E H S + H G + + +
Sbjct: 74 SECQLDSLKRLTFRGRIWVPGSDDEESPLNELRTKLVQESHDSTAAGHPGRNGTLEIVSR 253
Query: 144 LFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLTPLDVPEWKWDSISMDFVSSLPNT- 202
F+W G + V RFV C C + Q G L PL VP +SMDF++SLP T
Sbjct: 254 KFFWPGQSQTVRRFVRNCDVCGGIHIWRQAKRGFLKPLPVPNRLHSDLSMDFITSLPPTR 433
Query: 203 SRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYIQNIVKLHGVPSSIVSDRDPRFTS 262
RG +WV+VDRL+KS ++ + A+ ++ + HG+P SIVSDR +
Sbjct: 434 GRGSQYLWVIVDRLSKSVTLEEMD-TMEAEACAQRFLSCHYRFHGMPQSIVSDRGSNWVG 610
Query: 263 RFWR 266
RFWR
Sbjct: 611 RFWR 622
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 64.3 bits (155), Expect = 9e-11
Identities = 42/134 (31%), Positives = 72/134 (53%), Gaps = 1/134 (0%)
Frame = +2
Query: 4 LNYHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKIN-S 62
+ Y+P KA +VADALSR+ + +SA +E + L+ + L+ + LG+ +N +
Sbjct: 341 ITYYPGKANLVADALSRRRVDVSA--EREADDLDGMVRALRLNVLTKATESLGLEAVNQA 514
Query: 63 DFLGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLILEE 122
D IR AQ D + Q + ++++ + G + GRI +P++ LK+ I+ E
Sbjct: 515 DLFTRIRLAQGQDENLQKVA----QNDRTEYQTAKDGTILVNGRISVPNDRSLKEEIMSE 682
Query: 123 GHKSNLSIHLGATK 136
HKS S+H GA +
Sbjct: 683 AHKSRFSVHPGAPR 724
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 45.8 bits (107), Expect = 3e-05
Identities = 40/156 (25%), Positives = 69/156 (43%), Gaps = 6/156 (3%)
Frame = +2
Query: 119 ILEEGHKSNLSIHLGATKMYQDLKKL-FWWSGLKKDVARFVYACLTCQKS---KVEHQRP 174
IL H SN + H +K +++ FWW + KD F+ C CQ+ ++ P
Sbjct: 104 ILFHCHGSNYAGHFAVSKTVSKIQQAGFWWPTMFKDAHSFISKCDPCQRQGNIS*RNEMP 283
Query: 175 AGLLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFI--PINISYPVA 232
+ ++V +D +DF+ P +S + I V VD ++K I P N + V
Sbjct: 284 QNFILEVEV----FDVWGIDFMGPFP-SSYNNKYILVAVDYVSKWVEAIASPTNDATVVV 448
Query: 233 QLAEIYIQNIVKLHGVPSSIVSDRDPRFTSRFWRSL 268
++ + I GVP ++SD F ++ + L
Sbjct: 449 KM---FKSVIFPRFGVPRVVISDGGSHFINKVFEKL 547
>BG644733 weakly similar to GP|15289942|db putative polyprotein {Oryza sativa
(japonica cultivar-group)}, partial (1%)
Length = 174
Score = 40.0 bits (92), Expect = 0.002
Identities = 21/43 (48%), Positives = 26/43 (59%)
Frame = -3
Query: 204 RGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYIQNIVKLH 246
+ ++SI VVVDRLTKS FIP SY A I + IV +H
Sbjct: 130 KSYESI*VVVDRLTKSTLFIPFKTSYSAK*YARILLDEIVCIH 2
>TC93746 weakly similar to GP|22830935|dbj|BAC15800. hypothetical
protein~similar to gag-pol polyprotein {Oryza sativa
(japonica cultivar-group)}, partial (4%)
Length = 1019
Score = 34.7 bits (78), Expect = 0.076
Identities = 24/84 (28%), Positives = 39/84 (45%), Gaps = 13/84 (15%)
Frame = +2
Query: 153 DVARFVYACLTCQKSKVEHQRPAGLLTPLD-------------VPEWKWDSISMDFVSSL 199
D + +A L + K E +P G L ++ VP+ W+ +++DF L
Sbjct: 350 DAIKIKFAPLPLNEGKEEF-KPLGFLADMEPFKDKTKLCLLRPVPKPPWEDVTIDFSLGL 526
Query: 200 PNTSRGHDSIWVVVDRLTKSAHFI 223
T + DS VV D+ ++ AHFI
Sbjct: 527 L*TQQLKDSKMVVGDKFSRMAHFI 598
>BF003873 similar to GP|14715222|em putative polyprotein {Cicer arietinum},
partial (82%)
Length = 559
Score = 32.3 bits (72), Expect = 0.38
Identities = 18/29 (62%), Positives = 21/29 (72%)
Frame = +3
Query: 415 DVH*SRES*LQSSSVLITFQKELELWRIQ 443
DV *S+ S*L S V I +QKELE WRI+
Sbjct: 9 DVL*SQRS*L*DSLVRIRYQKELERWRIE 95
>TC77851 similar to GP|15982840|gb|AAL09767.1 AT5g03040/F15A17_70
{Arabidopsis thaliana}, partial (42%)
Length = 1921
Score = 30.0 bits (66), Expect = 1.9
Identities = 13/34 (38%), Positives = 21/34 (61%)
Frame = -1
Query: 30 VKEFELLEQFRDLSLVCELSSQSVQLGMLKINSD 63
V E E+L+QF D++L C + S ++ +NSD
Sbjct: 361 VDEKEVLDQFLDMNLKCSIFSSQTISSLIHLNSD 260
>BF633468 similar to GP|20259516|gb unknown protein {Arabidopsis thaliana},
partial (44%)
Length = 589
Score = 28.1 bits (61), Expect = 7.1
Identities = 16/35 (45%), Positives = 20/35 (56%)
Frame = +2
Query: 367 LQRKSR*LERR*KRRRVDKRAIMTSVGKTLSFRKG 401
L+R+ R ERR KRR V K I + K + FR G
Sbjct: 14 LEREKRTKERRLKRRTVKKERIPLNSDKKVRFRYG 118
>TC80911 similar to GP|3668118|emb|CAA11819.1 hypothetical protein {Brassica
napus}, partial (22%)
Length = 860
Score = 27.7 bits (60), Expect = 9.3
Identities = 10/29 (34%), Positives = 19/29 (65%)
Frame = +3
Query: 243 VKLHGVPSSIVSDRDPRFTSRFWRSLQMR 271
+++HG+P +S R+ + SR WR ++ R
Sbjct: 759 LQVHGLPVGFISLRNTQLLSRRWRGVRFR 845
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.341 0.148 0.472
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,278,792
Number of Sequences: 36976
Number of extensions: 175305
Number of successful extensions: 1325
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1311
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1321
length of query: 443
length of database: 9,014,727
effective HSP length: 99
effective length of query: 344
effective length of database: 5,354,103
effective search space: 1841811432
effective search space used: 1841811432
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144765.5


