

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144760.12 + phase: 0 /pseudo
(1302 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
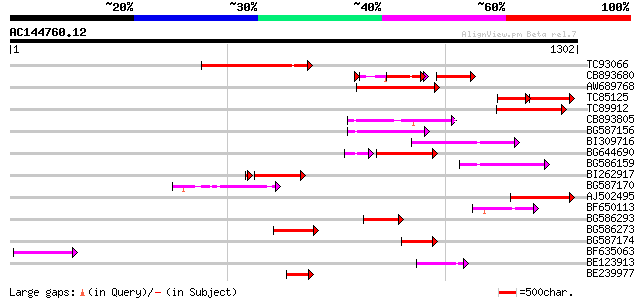
Sequences producing significant alignments: (bits) Value
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 244 2e-64
CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotia... 131 2e-56
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 186 4e-47
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 107 2e-45
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 162 1e-39
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 159 7e-39
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 155 1e-37
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 147 3e-35
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 129 5e-35
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 145 1e-34
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 142 1e-34
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 133 4e-31
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 109 6e-24
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 99 1e-20
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 87 4e-17
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 82 1e-15
BG587174 similar to PIR|A47759|A4775 retrovirus-related reverse ... 80 7e-15
BF635063 weakly similar to PIR|F84486|F84 probable retroelement ... 79 9e-15
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 74 4e-13
BE239977 weakly similar to GP|23237899|db polyprotein-like {Oryz... 63 8e-10
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 244 bits (622), Expect = 2e-64
Identities = 119/260 (45%), Positives = 172/260 (65%), Gaps = 5/260 (1%)
Frame = +1
Query: 440 EIDICEDCIL-GKQKRVSFQTSGRTPKKEKLELVHSDVWGPTTVPSIGGKHYFVTFIDDH 498
+++ C+ + G +K+VSF T+ K L+ +HSD+WGP+ V S GG+ Y +T IDD
Sbjct: 10 KLEFCKHLLFFGNRKKVSFSTATHRTKGI-LDYIHSDLWGPSKVTSYGGRRYMMTIIDDF 186
Query: 499 SRKVWVYFLKHKSEVFEAFKRWKAMVENETDLKIKKLRTDNGGEYEDTKFKKFCYEHGIR 558
RKVWVYFL++K+E F FK+W+ +VE +T +KKL TDN E+ + F +FC HGI
Sbjct: 187 PRKVWVYFLRYKNETFPTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGIA 366
Query: 559 MERTVPGTPQHNGVAERMNRTLTERARSLRVQSGL--PKKFWAEAVNTSAYLINRGPSVP 616
+T+P PQ NGVAERM RTL ERAR + +GL + W EA +T+ +L+NR P
Sbjct: 367 RHKTIPRNPQQNGVAERMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHSA 546
Query: 617 LEHKIPEEVWSGKEVKLSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRL 676
L+ K+PE++WSG V S+LR+FGC AY ++D KL P++ +CIF+ Y + GYRL
Sbjct: 547 LDFKVPEDIWSGNLVDYSNLRIFGCPAYALVND---GKLAPRAGECIFLSYASESKGYRL 717
Query: 677 W--DDENKKMVRSKDVIFNE 694
W D +++K++ S+DV FNE
Sbjct: 718 WCSDPKSQKLILSRDVTFNE 777
>CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotiana tabacum},
partial (7%)
Length = 780
Score = 131 bits (329), Expect(2) = 2e-56
Identities = 70/90 (77%), Positives = 75/90 (82%)
Frame = -3
Query: 981 MLVAGSNIDEIKNLKIQLSKEFDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYINRVLQ 1040
+LV GSNIDEIKNLK + SKE DMKDLGPAKKI+GMQI DKQKGVL LS EYI RVLQ
Sbjct: 271 LLVVGSNIDEIKNLKTRFSKEIDMKDLGPAKKIIGMQIMIDKQKGVL*LSQVEYITRVLQ 92
Query: 1041 RFNMGDAKLVSTPLASHFRLSQEQSPQTEE 1070
FNMG+A LVST LASHF LS EQSPQTE+
Sbjct: 91 IFNMGNAILVSTTLASHFCLSHEQSPQTEK 2
Score = 108 bits (269), Expect(2) = 2e-56
Identities = 60/91 (65%), Positives = 69/91 (74%), Gaps = 1/91 (1%)
Frame = -2
Query: 865 FAPVVKLNTIRSVLSIVASENLYLEQLDVKTAFLHGDLVEEIYMHQPEGFL-EEGKENMV 923
F P+VKLNTI +LSIVA ENLYLE LDVKTAFL GDLVE+IYMHQPEGF E GK MV
Sbjct: 554 FVPIVKLNTIMFLLSIVAIENLYLE*LDVKTAFLRGDLVEDIYMHQPEGFS*EVGK--MV 381
Query: 924 CMLKKSLYGLKQAPRQWYMKFESFMHKEGFQ 954
LKKS+YGLKQ PRQ ++ ++GF+
Sbjct: 380 GKLKKSMYGLKQGPRQCI*SLKALCTRKGFR 288
Score = 50.4 bits (119), Expect(2) = 4e-07
Identities = 46/166 (27%), Positives = 68/166 (40%), Gaps = 7/166 (4%)
Frame = -1
Query: 803 EEMKSLISNQTWELAKLPIGKKALHNKWVYRVKEDHDGSKRYKARLVVKGFRQKEGIDY- 861
+++KSLIS+ TWE A +D SKRYK+R+VVK +++
Sbjct: 678 KKIKSLISDHTWEQAT-------------------YDVSKRYKSRMVVKDSSKRKNFTMK 556
Query: 862 ------TEIFAPVVKLNTIRSVLSIVASENLYLEQLDVKTAFLHGDLVEEIYMHQPEGFL 915
+ + +K LS + + + ++H + G
Sbjct: 555 FCSNS*AQYYHVSIKYCCH*KSLS*IVGCEDCISSWRLS*GYIHAPT*RILIRSGENG-- 382
Query: 916 EEGKENMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNADHC 961
GK K+ K + YMKFESFMHKEGFQK N +HC
Sbjct: 381 --GKT------KEEHVWTKTRSKTMYMKFESFMHKEGFQK*NFNHC 268
Score = 23.1 bits (48), Expect(2) = 4e-07
Identities = 9/13 (69%), Positives = 12/13 (92%)
Frame = -3
Query: 791 TTDASKWELAMKE 803
TT ASKW+L+MK+
Sbjct: 712 TTCASKWKLSMKK 674
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 186 bits (473), Expect = 4e-47
Identities = 97/193 (50%), Positives = 126/193 (65%), Gaps = 1/193 (0%)
Frame = +1
Query: 796 KWELAMKEEMKSLISNQTWELAKLPIGKKALHNKWVYRVKEDHDGS-KRYKARLVVKGFR 854
+W AMK E K+LI N+TW+L LP KKA+ KWVYRVKE+ DGS ++KARLV KGF
Sbjct: 94 RWLQAMKTEYKALIDNKTWDLVPLPPHKKAIGCKWVYRVKENPDGSVNKFKARLVAKGFS 273
Query: 855 QKEGIDYTEIFAPVVKLNTIRSVLSIVASENLYLEQLDVKTAFLHGDLVEEIYMHQPEGF 914
Q G DYTE F+PV+K TIR +L+I + ++Q+D+ AFL+G L EE+YM QP+GF
Sbjct: 274 QTLGCDYTETFSPVIKPVTIRLILTIAITYKWEIQQIDINNAFLNGFLQEEVYMSQPQGF 453
Query: 915 LEEGKENMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNADHCCFFKRYKSSYIIL 974
E +++VC L KSLYGLKQAPR WY S + GF K D + I L
Sbjct: 454 -EAANKSLVCKLNKSLYGLKQAPRAWYEXLTSAQIQFGFTKSRCDPSLLIYNQNGACIYL 630
Query: 975 LLYVDDMLVAGSN 987
+YVDD+L+ GS+
Sbjct: 631 XIYVDDILITGSS 669
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 107 bits (268), Expect(2) = 2e-45
Identities = 50/102 (49%), Positives = 72/102 (70%)
Frame = +3
Query: 1195 AVTEASKELIWLQGLLTELGFMQEKSALYSDSQSAIHLAKNSAFHSRTKHIGLRYHFIRS 1254
++ +A KE IW+Q L+ ELG QE+ +Y DSQSA+H+A+N AFHSRTKHIG++YHF+R
Sbjct: 228 SLPQACKEAIWMQRLMEELGHKQEQITVYCDSQSALHIARNPAFHSRTKHIGIQYHFVRE 407
Query: 1255 LLEDEVLTLIKIQGSKNPADMLTKVVTIDKLKLCSTLVGLLE 1296
++E+ + + KI + N AD +TK + DK C + GLLE
Sbjct: 408 VVEEGSVDMQKIHTNDNLADAMTKSINTDKFIWCRSSYGLLE 533
Score = 95.1 bits (235), Expect(2) = 2e-45
Identities = 43/76 (56%), Positives = 60/76 (78%)
Frame = +1
Query: 1120 VKWILRYLRGTTEKCLYFGKGEIKVEGYVDADFAGEVDHRRSTTGYIFTVGTRSVSWMSR 1179
VK I+RY++GT+ + FG E+ V GYVD+DFAG+ D R+STTGY+FT+ +VSW+S+
Sbjct: 1 VKRIMRYIKGTSGVAVCFGGSELTVRGYVDSDFAGDHDKRKSTTGYVFTLAGGAVSWLSK 180
Query: 1180 IQKIVALSTTEVEYVA 1195
+Q +VALSTTE EY+A
Sbjct: 181 LQTVVALSTTEAEYMA 228
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 162 bits (409), Expect = 1e-39
Identities = 75/165 (45%), Positives = 113/165 (68%), Gaps = 3/165 (1%)
Frame = +1
Query: 1118 EAVKWILRYLRGTTEKCLYFGKG---EIKVEGYVDADFAGEVDHRRSTTGYIFTVGTRSV 1174
+A+KW+L+YL + + L + K E +EGYVDAD+AG VD R+S +G++FT+ ++
Sbjct: 1 QALKWVLKYLNESLKSSLKYTKAAQEEDALEGYVDADYAGNVDTRKSLSGFVFTLYGTTI 180
Query: 1175 SWMSRIQKIVALSTTEVEYVAVTEASKELIWLQGLLTELGFMQEKSALYSDSQSAIHLAK 1234
SW + Q +V LSTT+ EY+A E K+ IWL+G++ ELG QE ++ DSQSAIHLA
Sbjct: 181 SWKANQQSVVTLSTTQAEYIAFVEGVKDAIWLKGMIGELGITQEYVKIHCDSQSAIHLAN 360
Query: 1235 NSAFHSRTKHIGLRYHFIRSLLEDEVLTLIKIQGSKNPADMLTKV 1279
+ +H RTKHI +R HFIR ++E + + + K+ +NPAD+ TK+
Sbjct: 361 HQVYHERTKHIDIRLHFIRDMIESKEIVVEKMASEENPADVFTKL 495
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 159 bits (402), Expect = 7e-39
Identities = 91/267 (34%), Positives = 149/267 (55%), Gaps = 15/267 (5%)
Frame = +3
Query: 775 LLTDGGEPEDYDEACQTTDASKWELAMKEEMKSLISNQTWELAKLPIGKKALHNKWVYRV 834
+LT +P ++EA ++ KW +M EM++ N TWEL L G K + KW+++
Sbjct: 21 MLTMTSDPTTFEEAVKS---EKWRASMNNEMEATERNNTWELTDLRSGAKTIGLKWIFKT 191
Query: 835 KEDHDGS-KRYKARLVVKGFRQKEGIDYTEIFAPVVKLNTIRSVLSIVASENLYLEQLDV 893
K + +G ++YKARLV KG+ Q+ G+DYTE+FAPV + +TIR V+++ A
Sbjct: 192 KLNENGEIEKYKARLVAKGYSQQYGVDYTEVFAPVARWDTIRMVIALAAQ---------- 341
Query: 894 KTAFLHGDLVEEIYMHQPEGFLEEGKENMVCM-------------LKKSLYGLKQAPRQW 940
+ D V M + +E + + +K++LYGLKQAPR W
Sbjct: 342 ----IKRDGVCIS*M*KAHSCMEN*MRKFLLINHRVM*RRVIS*RVKRALYGLKQAPRAW 509
Query: 941 YMKFESFMHKEGFQKCNADHCCFFKRYKSSYIILL-LYVDDMLVAGSNIDEIKNLKIQLS 999
Y + E++ KEGF+KC +H F K + I+++ LYVDD++ G++ + + K +
Sbjct: 510 YSRIEAYFTKEGFEKCPYEHTLFVKLSEGGKILIISLYVDDLIFIGNDENMFEEFKKSMK 689
Query: 1000 KEFDMKDLGPAKKILGMQITRDKQKGV 1026
KEF+M DLG LG+++T++ +KG+
Sbjct: 690 KEFNMSDLGKMHYFLGVEVTQN-EKGI 767
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 155 bits (391), Expect = 1e-37
Identities = 81/189 (42%), Positives = 115/189 (59%), Gaps = 1/189 (0%)
Frame = -1
Query: 776 LTDGGEPEDYDEACQTTDASKWELAMKEEMKSLISNQTWELAKLPIGKKALHNKWVYRVK 835
L + P Y+EA + + W+ ++ E ++I N TW ++LP GKKA+ ++W++ +K
Sbjct: 576 LNENHIPRSYEEAMEDKE---WKESVGAEAGAMIKNDTWYESELPKGKKAVSSRWIFTIK 406
Query: 836 EDHDGS-KRYKARLVVKGFRQKEGIDYTEIFAPVVKLNTIRSVLSIVASENLYLEQLDVK 894
DGS +R K RLV +GF G DY E FAPV KL+TIR VLS+ + L Q+DVK
Sbjct: 405 YKADGSIERKKTRLVARGFTLTYGEDYIETFAPVAKLHTIRIVLSLAVNLGWGLWQMDVK 226
Query: 895 TAFLHGDLVEEIYMHQPEGFLEEGKENMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQ 954
AFL G+L +E+YM+ P G K V LKK++YGLKQ+PR WY K + ++ GF+
Sbjct: 225 NAFLQGELEDEVYMYPPPGLEHLVKRGNVLRLKKAIYGLKQSPRAWYNKLSTTLNGRGFR 46
Query: 955 KCNADHCCF 963
K DH F
Sbjct: 45 KSELDHTLF 19
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 147 bits (370), Expect = 3e-35
Identities = 90/250 (36%), Positives = 141/250 (56%), Gaps = 1/250 (0%)
Frame = +2
Query: 923 VCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNADHCCFFKRYKSSYIILLLYVDDML 982
VC L+KS+YGLKQA RQWY K + G+ + ++D F K SS+ LL+YVDD++
Sbjct: 20 VCELQKSIYGLKQASRQWYSKLSESLISFGYLQSSSDFSLFTKFKDSSFTTLLVYVDDIV 199
Query: 983 VAGSNIDEIKNLKIQLSKEFDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYINRVLQRF 1042
+AG++I EI+++K L F +KDLG + LG+++ R KQ G+L L+ +Y +L+
Sbjct: 200 LAGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARSKQ-GIL-LNQRKYTLELLEDS 373
Query: 1043 NMGDAKLVSTPLASHFRLSQEQSPQTEEEKELMAKIPYASAIGSLMYAMVCTRPDIGHAV 1102
K TP +L SP +E + Y IG L+Y + TRPDI AV
Sbjct: 374 GNLAVKSTLTPYDISLKLHNSDSPLYNDETQ------YRRLIGKLIY-LTTTRPDISFAV 532
Query: 1103 GVVSRFMSNPGKAHWEAVKWILRYLRGTTEKCLYF-GKGEIKVEGYVDADFAGEVDHRRS 1161
+S+F+S P + H++A +L+YL+ K L++ +K+ + D+D+A R+S
Sbjct: 533 QQLSQFVSKPQQVHYQAAIRVLQYLKTAPAKGLFYSATSNLKLSSFADSDWATCPTTRKS 712
Query: 1162 TTGYIFTVGT 1171
TGY +G+
Sbjct: 713 VTGYWVFLGS 742
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 129 bits (325), Expect(2) = 5e-35
Identities = 62/139 (44%), Positives = 92/139 (65%)
Frame = -2
Query: 843 RYKARLVVKGFRQKEGIDYTEIFAPVVKLNTIRSVLSIVASENLYLEQLDVKTAFLHGDL 902
R K++LVV+G+ QKEGIDY E F+PV ++ IR +++ A L Q+DVK+AF++GDL
Sbjct: 418 RNKSKLVVQGYNQKEGIDYDEAFSPVARMEAIRILIAFAAFMGFKLYQMDVKSAFINGDL 239
Query: 903 VEEIYMHQPEGFLEEGKENMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNADHCC 962
EE+++ QP GF + N V L K+LYGLKQAPR WY + F+ K GF++ D+
Sbjct: 238 KEEVFVKQPPGFEDAEVPNHVFRLNKTLYGLKQAPRAWYERLSKFLLKNGFKRGKIDNTL 59
Query: 963 FFKRYKSSYIILLLYVDDM 981
F + + +I+ +YVDD+
Sbjct: 58 FLLKRE*ELLIIQVYVDDI 2
Score = 38.1 bits (87), Expect(2) = 5e-35
Identities = 19/68 (27%), Positives = 32/68 (46%)
Frame = -3
Query: 768 RRYIDYMLLTDGGEPEDYDEACQTTDASKWELAMKEEMKSLISNQTWELAKLPIGKKALH 827
R + + EP++ EA + D W +M+EE+ ++ W L P GK +
Sbjct: 627 RNLVSFSAFISSIEPKNVKEALRDAD---WINSMQEELHQFERSKVWYLVPRPEGKTVIG 457
Query: 828 NKWVYRVK 835
+WV+R K
Sbjct: 456 TRWVFRNK 433
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 145 bits (365), Expect = 1e-34
Identities = 75/209 (35%), Positives = 126/209 (59%), Gaps = 2/209 (0%)
Frame = +1
Query: 1033 EYINRVLQRFNMGDAKLVSTPLASHFRLSQEQSPQTEEEKELMAKIPYASAIGSLMYAMV 1092
+Y+ +L+RF M + L P+A +L ++++ + + Y +G LMY +
Sbjct: 46 KYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATK------YKQIVGCLMY-LA 204
Query: 1093 CTRPDIGHAVGVVSRFMSNPGKAHWEAVKWILRYLRGTTEK-CLYFGKGEIKVEGYVDAD 1151
TRPD+ + + ++SRFM+ P + H AVK +LRYL GT +Y G K+E Y D+D
Sbjct: 205 ATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTINLGIMYKRNGSEKLEAYTDSD 384
Query: 1152 FAGEVDHRRSTTGYIFTVGTRSVSWMSRIQKIVALSTTEVEYVAVTEASKELIWLQGLLT 1211
+AG++D R+ST+GY+F + + +VSW S+ Q +V LSTT+ E++A + + +W++ +L
Sbjct: 385 YAGDLDDRKSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFIAAAFCACQSVWMRRVLE 564
Query: 1212 ELGFMQEKS-ALYSDSQSAIHLAKNSAFH 1239
+LG+ Q S +Y D+ S I L+KN H
Sbjct: 565 KLGYTQSGSITMYCDNNSTIKLSKNPVLH 651
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 142 bits (359), Expect(2) = 1e-34
Identities = 67/116 (57%), Positives = 81/116 (69%)
Frame = +1
Query: 563 VPGTPQHNGVAERMNRTLTERARSLRVQSGLPKKFWAEAVNTSAYLINRGPSVPLEHKIP 622
V TPQ NGVAERMNRTL ER R++ +G+ K FWAEAV T+ Y+INR PS ++ K P
Sbjct: 76 VAHTPQQNGVAERMNRTLLERTRAMLKTAGMAKSFWAEAVKTACYVINRSPSTVIDLKTP 255
Query: 623 EEVWSGKEVKLSHLRVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWD 678
E+W GK V S L VFGC YV + Q R KLDPKS+KCIF+GY ++ GY LWD
Sbjct: 256 MEMWKGKPVDYSSLHVFGCPVYVMYNSQERTKLDPKSRKCIFLGYADNVKGYXLWD 423
Score = 23.5 bits (49), Expect(2) = 1e-34
Identities = 9/17 (52%), Positives = 11/17 (63%)
Frame = +3
Query: 541 GEYEDTKFKKFCYEHGI 557
GEY D +F FC + GI
Sbjct: 9 GEYVDGEFLAFCKQEGI 59
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 133 bits (335), Expect = 4e-31
Identities = 94/257 (36%), Positives = 130/257 (50%), Gaps = 10/257 (3%)
Frame = -3
Query: 374 KISKGAMTIARGRKSGTLYKTAGA-------CHLIAVATNENPNLWHKRLGHMSEKGMKV 426
++ + + I G G LY C + ++ LWH RLGH + + +
Sbjct: 716 EVIESSQLIGEGVTKGDLYMLEKLDPVSNYKCSFTSSSSLNKDALWHARLGHPHGRALNL 537
Query: 427 MHSKGKLPSLRSIEIDICEDCILGKQ-KRVSFQTSGRTPKKEKLELVHSDVWGPTTVPSI 485
M LP + E CE CILGK K V +TS T + +L+++D+W T PS+
Sbjct: 536 M-----LPGV-VFENKNCEACILGKHCKNVFPRTS--TVYENCFDLIYTDLW---TAPSL 390
Query: 486 GGKH--YFVTFIDDHSRKVWVYFLKHKSEVFEAFKRWKAMVENETDLKIKKLRTDNGGEY 543
+ YFVTFID+ S+ W+ + K V +AFK ++A V N KIK LR+DNGGEY
Sbjct: 389 SRDNHKYFVTFIDEKSKYTWLTLIPSKDRVIDAFKNFQAYVTNHYHAKIKILRSDNGGEY 210
Query: 544 EDTKFKKFCYEHGIRMERTVPGTPQHNGVAERMNRTLTERARSLRVQSGLPKKFWAEAVN 603
FK HGI + + P TPQ NGVA+R N+ L E ARSL Q+ V+
Sbjct: 209 TSYAFKSHLDHHGILHQTSCPYTPQQNGVAKRKNKHLMEVARSLMFQAN---------VS 57
Query: 604 TSAYLINRGPSVPLEHK 620
T+ YLIN P+ L K
Sbjct: 56 TACYLINWIPTKVLRIK 6
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 109 bits (273), Expect = 6e-24
Identities = 52/148 (35%), Positives = 93/148 (62%), Gaps = 1/148 (0%)
Frame = +2
Query: 1150 ADFAGEVDHRRSTTGYIFTVGTRSVSWMSRIQKIVALSTTEVEYVAVTEASKELIWLQGL 1209
+D+AG+ + R+ST+GY F +GT ++SW S+ Q +VA ST E EY+A T + + +WL+ +
Sbjct: 2 SDWAGDTETRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRI 181
Query: 1210 LTELGFMQE-KSALYSDSQSAIHLAKNSAFHSRTKHIGLRYHFIRSLLEDEVLTLIKIQG 1268
L + Q + +Y D++SAI L+KN FH R+KHI +++H IR L+ ++ + +
Sbjct: 182 LEVMHHEQNTPTKIYCDNKSAIALSKNPVFHGRSKHIDIQFHKIRELIAEKEVVIEYCPT 361
Query: 1269 SKNPADMLTKVVTIDKLKLCSTLVGLLE 1296
+ AD+ TK + I+ ++G+++
Sbjct: 362 EEKIADIFTKPLKIESFYKLKKMLGMMK 445
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 98.6 bits (244), Expect = 1e-20
Identities = 62/163 (38%), Positives = 95/163 (58%), Gaps = 11/163 (6%)
Frame = +1
Query: 1062 QEQSPQTEEEKELMAKIPYASAIGSL-------MYAMVCTRPDIGHAVGVVSRFMSNPGK 1114
+E+ + E+E+ L P S IG + + +C RPDI ++V V+S+FM +P K
Sbjct: 1 RERKWELEQEQGLEKGEPQESRIGVVRS*IEVQVGINLC*RPDICYSVSVISKFMHDPRK 180
Query: 1115 AHWEAVKWILRYLRGTTEKCLYF---GKGEI-KVEGYVDADFAGEVDHRRSTTGYIFTVG 1170
H A ILRY+RGT E L F K E+ ++ Y D+D+ G+ RRST+GY+F
Sbjct: 181 PHLIAANRILRYVRGTMEYGLLFPYGAKSEVYELICYSDSDWCGD---RRSTSGYVFKFN 351
Query: 1171 TRSVSWMSRIQKIVALSTTEVEYVAVTEASKELIWLQGLLTEL 1213
++SW ++ Q I ALS+ E EY+A T A+ + +WL ++ EL
Sbjct: 352 DAAISWCTKKQPITALSSYEAEYIAGTFATFQALWLDSVIKEL 480
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 87.0 bits (214), Expect = 4e-17
Identities = 41/93 (44%), Positives = 65/93 (69%), Gaps = 1/93 (1%)
Frame = +2
Query: 812 QTWELAKLPIGKKALHNKWVYRVKEDHDGSK-RYKARLVVKGFRQKEGIDYTEIFAPVVK 870
QT +L K P G K + +W+Y++K + DG+ +YKARLV KG+ +++GID+ E+FAPVV+
Sbjct: 53 QTLKLVKKPTGVKPIGLRWIYKIKRNEDGTLIKYKARLVAKGYVKQQGIDFDEVFAPVVR 232
Query: 871 LNTIRSVLSIVASENLYLEQLDVKTAFLHGDLV 903
+ TI +L++ A+ + +DVK AFL+G V
Sbjct: 233 IETI*LLLALAATNGC*IHHIDVKIAFLNGHFV 331
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 82.4 bits (202), Expect = 1e-15
Identities = 41/105 (39%), Positives = 69/105 (65%), Gaps = 1/105 (0%)
Frame = -2
Query: 605 SAYLINRGPSVPLEHKIPEEVWSGKEVKLSHLRVFGCVAYVHISDQGRNKLDPKSKKCIF 664
+ YLINR P+ L+ + P EV + ++ L+++RVFGC+ YV + + RNKL+ +S+K +F
Sbjct: 704 ACYLINRIPTRVLKDQAPFEVLNQRKPSLTYMRVFGCLCYVLVPGELRNKLEARSRKAMF 525
Query: 665 IGYGEDEFGYRLWDDENKKMVRSKDVIF-NERVMYKDKHNTTTND 708
IGY + GY+ +D E ++++ S+DV F ER Y++K+ D
Sbjct: 524 IGYSTTQKGYKCYDPEARRVLVSRDVKFIEERGYYEEKNQEDLRD 390
>BG587174 similar to PIR|A47759|A4775 retrovirus-related reverse
transcriptase homolog - rape retrotransposon copia-like
(fragment), partial (84%)
Length = 249
Score = 79.7 bits (195), Expect = 7e-15
Identities = 44/83 (53%), Positives = 54/83 (65%), Gaps = 1/83 (1%)
Frame = -1
Query: 901 DLVEEIYMHQPEGFLEEGKENMVCMLKKSLYGLKQAPRQWYMKFESFMHKEGFQKCNADH 960
+L E+IYM QPEGFL GKE+ VC L+KSLYGLKQ+PRQWY +F+S+
Sbjct: 249 ELEEKIYMTQPEGFLFPGKEDHVCKLRKSLYGLKQSPRQWYKRFDSYRSSWATTGVLMTV 70
Query: 961 CCFFKRYK-SSYIILLLYVDDML 982
R + S YI L+LYVDDML
Sbjct: 69 VST*TR*RMSRYIYLVLYVDDML 1
>BF635063 weakly similar to PIR|F84486|F84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(4%)
Length = 677
Score = 79.3 bits (194), Expect = 9e-15
Identities = 46/151 (30%), Positives = 84/151 (55%), Gaps = 3/151 (1%)
Frame = -2
Query: 8 IEKFDGA-DFGFWKMQIEDYLYQKKLHQPLTEKK--PDSMKDDEWSLLDRQALGVVRLSL 64
IEKF G DFG WK+++E L Q+K + L + P +M E + + +A + L L
Sbjct: 550 IEKFTGDNDFGLWKVKMEAVLIQQKCEKALKGEVSLPVTMSRAEKTEMVDKARSAIVLCL 371
Query: 65 SRNVAFNIAKEKTTAGLMKALSSMYEKPPSSNKVHLMRRLFTLRMAEGMSVAQHINELNI 124
V +AKE+T A + L S+Y +++ L ++L++ RM E ++ + + E N
Sbjct: 370 GDKVLREVAKERTAASMWAKL*SLYMTKSLAHRQFLKQQLYSFRMVESKAIMEQLTEFNK 191
Query: 125 VTTQLSSVGIEFDDEVRALILLSSLPDSWSA 155
+ L ++ ++ +DE +A++LL +LP S+ +
Sbjct: 190 ILDDLENIEVQLEDEEKAILLLCALPKSFES 98
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 73.9 bits (180), Expect = 4e-13
Identities = 42/120 (35%), Positives = 67/120 (55%), Gaps = 1/120 (0%)
Frame = +1
Query: 935 QAPRQWYMKFESFMHKEGFQKCNADHCCFFKRYKS-SYIILLLYVDDMLVAGSNIDEIKN 993
Q+PR W+ +F + K G+ +C DH F K + IL++YVDD+ + G + IK
Sbjct: 1 QSPRDWFDRFT*VVKKFGYIQCQTDHAMFIKHSSTVKKAILIVYVDDIFLTGDHGK*IKR 180
Query: 994 LKIQLSKEFDMKDLGPAKKILGMQITRDKQKGVLQLS*AEYINRVLQRFNMGDAKLVSTP 1053
LK L++EF++KDLG K LGM++ R K+ +S +Y+ +L+ M K + P
Sbjct: 181 LKNLLAEEFEIKDLGNLKYFLGMEVARWKKGS--SISQRKYVLDLLKETRMIGCKTIRDP 354
>BE239977 weakly similar to GP|23237899|db polyprotein-like {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 514
Score = 62.8 bits (151), Expect = 8e-10
Identities = 26/62 (41%), Positives = 40/62 (63%)
Frame = -2
Query: 637 RVFGCVAYVHISDQGRNKLDPKSKKCIFIGYGEDEFGYRLWDDENKKMVRSKDVIFNERV 696
R+FGC ++VHI GR+K D ++ KC+FI Y + GYR + ++K S+DV F+E+
Sbjct: 456 RIFGCTSFVHIHSDGRSKFDHRALKCVFIRYSSTQKGYRCYHPPSRKYFVSRDVTFHEQE 277
Query: 697 MY 698
Y
Sbjct: 276 SY 271
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.133 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,747,146
Number of Sequences: 36976
Number of extensions: 558152
Number of successful extensions: 3061
Number of sequences better than 10.0: 83
Number of HSP's better than 10.0 without gapping: 2955
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3024
length of query: 1302
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1194
effective length of database: 5,021,319
effective search space: 5995454886
effective search space used: 5995454886
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC144760.12


