

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144759.9 - phase: 0
(749 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
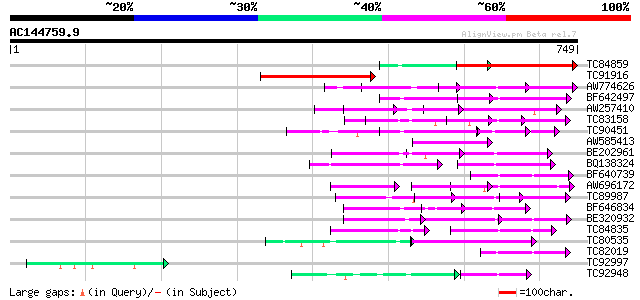
Score E
Sequences producing significant alignments: (bits) Value
TC84859 weakly similar to GP|9758774|dbj|BAB09072.1 selenium-bin... 320 2e-87
TC91916 similar to GP|14530645|emb|CAB61040. contains similarity... 304 9e-83
AW774626 similar to PIR|T47909|T47 hypothetical protein T20K12.7... 87 3e-17
BF642497 weakly similar to PIR|H96713|H96 hypothetical protein T... 84 2e-16
AW257410 weakly similar to GP|10177533|db selenium-binding prote... 82 1e-15
TC83158 weakly similar to PIR|H86191|H86191 hypothetical protein... 80 4e-15
TC90451 similar to PIR|H84508|H84508 hypothetical protein At2g13... 79 5e-15
AW585413 weakly similar to PIR|B85153|B851 hypothetical protein ... 74 3e-13
BE202961 weakly similar to GP|6016735|gb|A hypothetical protein ... 71 2e-12
BQ138324 weakly similar to GP|12322902|gb| hypothetical protein;... 68 1e-11
BF640739 weakly similar to GP|7363286|dbj Similar to Arabidopsis... 65 1e-10
AW696172 weakly similar to PIR|H96620|H96 hypothetical protein F... 63 4e-10
TC89987 weakly similar to PIR|D86443|D86443 probable PPR-repeat ... 61 2e-09
BF646834 weakly similar to PIR|A84784|A847 hypothetical protein ... 59 5e-09
BE320932 weakly similar to PIR|T04548|T045 hypothetical protein ... 58 1e-08
TC84835 weakly similar to PIR|T02328|T02328 hypothetical protein... 56 6e-08
TC80535 weakly similar to GP|14335136|gb|AAK59848.1 At2g15690/F9... 55 1e-07
TC82019 weakly similar to GP|11994382|dbj|BAB02341. gb|AAB82628.... 54 3e-07
TC92997 similar to GP|21743256|dbj|BAC03254. contains ESTs AU030... 50 2e-06
TC92948 weakly similar to GP|15623829|dbj|BAB67888. hypothetical... 50 4e-06
>TC84859 weakly similar to GP|9758774|dbj|BAB09072.1 selenium-binding
protein-like {Arabidopsis thaliana}, partial (6%)
Length = 532
Score = 320 bits (819), Expect = 2e-87
Identities = 159/159 (100%), Positives = 159/159 (100%)
Frame = +1
Query: 591 NTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQV 650
NTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQV
Sbjct: 1 NTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQV 180
Query: 651 HADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGL 710
HADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGL
Sbjct: 181 HADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGL 360
Query: 711 YIEAVKFVYQMKAAGVQPHEPLLEKLRIACGSSNFSSMN 749
YIEAVKFVYQMKAAGVQPHEPLLEKLRIACGSSNFSSMN
Sbjct: 361 YIEAVKFVYQMKAAGVQPHEPLLEKLRIACGSSNFSSMN 477
Score = 47.4 bits (111), Expect = 2e-05
Identities = 41/152 (26%), Positives = 61/152 (39%), Gaps = 2/152 (1%)
Frame = +1
Query: 489 SWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNV-PLGMQ 547
+W VS + A+ F M +G + +S +LKAC N G Q
Sbjct: 10 TWTAKIVSSCRERHFSEALGDFKKM----GRVGVKKDSFTFSSVLKACGRMQNRGSCGEQ 177
Query: 548 VHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRV-SRHNTLTWTAKIVSSCRER 606
VH +KLG + SLI YGR L DA +VF + N + A ++ +
Sbjct: 178 VHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNG 357
Query: 607 HFSEALGDFKKMGRVGVKKDSFTFSSVLKACG 638
+ EA+ +M GV+ + ACG
Sbjct: 358 LYIEAVKFVYQMKAAGVQPHEPLLEKLRIACG 453
>TC91916 similar to GP|14530645|emb|CAB61040. contains similarity to Pfam
domain: PF00096 (Zinc finger C2H2 type) Score=103.9
E-value=9.8e-28, partial (1%)
Length = 515
Score = 304 bits (778), Expect = 9e-83
Identities = 149/152 (98%), Positives = 150/152 (98%)
Frame = +1
Query: 332 MMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPSSQPIT 391
M IVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNL+LIHPSSQPIT
Sbjct: 58 MEAIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLALIHPSSQPIT 237
Query: 392 PPKKSKRRRKCDTTSHILPLMDALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRG 451
PPKKSKRRRKCDTTSHILPLMDALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRG
Sbjct: 238 PPKKSKRRRKCDTTSHILPLMDALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRG 417
Query: 452 IELPLTLLNRILIMFVSCGLLENARRVFDVMS 483
IELPLTLLNRILIMFVSCGLLENARRVFDVMS
Sbjct: 418 IELPLTLLNRILIMFVSCGLLENARRVFDVMS 513
>AW774626 similar to PIR|T47909|T47 hypothetical protein T20K12.70 -
Arabidopsis thaliana, partial (12%)
Length = 703
Score = 86.7 bits (213), Expect = 3e-17
Identities = 55/187 (29%), Positives = 100/187 (53%), Gaps = 4/187 (2%)
Frame = +3
Query: 567 LIRFYGRFKCLEDANMVFNRVS--RHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVK 624
L+ Y + KC+ +A +F + R N + WTA + + +A+ F+ M GV+
Sbjct: 3 LVDMYAKCKCVSEAEFLFKGLEFDRKNHVLWTAMVTGYAQNGDGYKAVEFFRYMHAQGVE 182
Query: 625 KDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAEL 684
+ +TF ++L AC + R GEQVH +K G S+ YVQ +L+ MY + G L++A+
Sbjct: 183 CNQYTFPTILTACSSVLAR-CFGEQVHGFIVKSGFGSNVYVQSALVDMYAKCGDLKNAKN 359
Query: 685 VFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIAC--GS 742
+ E T + +V S N++++G++++GL EA++ M ++ + + C GS
Sbjct: 360 MLE-TMEDDDVVSWNSLMVGFVRHGLEEEALRLFKNMHGRNMKIDDYTFPSVLNCCVVGS 536
Query: 743 SNFSSMN 749
N S++
Sbjct: 537 INPKSVH 557
Score = 84.7 bits (208), Expect = 1e-16
Identities = 63/227 (27%), Positives = 107/227 (46%), Gaps = 4/227 (1%)
Frame = +3
Query: 465 MFVSCGLLENARRVFDVMSV--RDFHSWATLFVSYYENGEYENAIDVFVSMLCQ-LDVMG 521
M+ C + A +F + ++ W + Y +NG+ A++ F M Q ++
Sbjct: 12 MYAKCKCVSEAEFLFKGLEFDRKNHVLWTAMVTGYAQNGDGYKAVEFFRYMHAQGVECNQ 191
Query: 522 FSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDAN 581
++FP +L AC+ + G QVHG ++K G +V + S+L+ Y + L++A
Sbjct: 192 YTFPT-----ILTACSSVLARCFGEQVHGFIVKSGFGSNVYVQSALVDMYAKCGDLKNAK 356
Query: 582 MVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQ 641
+ + + ++W + +V R EAL FK M +K D +TF SVL C
Sbjct: 357 NMLETMEDDDVVSWNSLMVGFVRHGLEEEALRLFKNMHGRNMKIDDYTFPSVLNCC---- 524
Query: 642 NRGSCG-EQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFE 687
GS + VH IK G ++ V +L+ MY ++G + A VFE
Sbjct: 525 VVGSINPKSVHGLIIKTGFENYKLVSNALVDMYAKTGDMDCAYTVFE 665
Score = 64.7 bits (156), Expect = 1e-10
Identities = 42/182 (23%), Positives = 90/182 (49%), Gaps = 1/182 (0%)
Frame = +3
Query: 417 FPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENAR 476
FP + +S++ C ++H I+ G + + + ++ M+ CG L+NA+
Sbjct: 198 FPTILTACSSVLARCF-------GEQVHGFIVKSGFGSNVYVQSALVDMYAKCGDLKNAK 356
Query: 477 RVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQ-LDVMGFSFPPWIWSCLLKA 535
+ + M D SW +L V + +G E A+ +F +M + + + ++FP + +C
Sbjct: 357 NMLETMEDDDVVSWNSLMVGFVRHGLEEEALRLFKNMHGRNMKIDDYTFPS-VLNC---- 521
Query: 536 CACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTW 595
C + VHG ++K G ++ L+S++L+ Y + ++ A VF ++ + ++W
Sbjct: 522 --CVVGSINPKSVHGLIIKTGFENYKLVSNALVDMYAKTGDMDCAYTVFEKMLEKDVISW 695
Query: 596 TA 597
T+
Sbjct: 696 TS 701
>BF642497 weakly similar to PIR|H96713|H96 hypothetical protein T6L1.11
[imported] - Arabidopsis thaliana, partial (20%)
Length = 461
Score = 84.0 bits (206), Expect = 2e-16
Identities = 41/152 (26%), Positives = 79/152 (51%)
Frame = +3
Query: 489 SWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQV 548
SW ++ + +NG +AID+F M + + + +L AC M + G QV
Sbjct: 9 SWTSMITGFTQNGLDRDAIDIFREMKLE----NLQMDQYTFGSVLTACGGVMALQEGKQV 176
Query: 549 HGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHF 608
H +++ D++ ++S+L+ Y + K ++ A VF +++ N ++WTA +V + +
Sbjct: 177 HAYIIRTDYKDNIFVASALVDMYCKCKNIKSAEAVFKKMTCKNVVSWTAMLVGYGQNGYS 356
Query: 609 SEALGDFKKMGRVGVKKDSFTFSSVLKACGRM 640
EA+ F M + G++ D FT SV+ +C +
Sbjct: 357 EEAVKTFSDMQKYGIEPDDFTLGSVISSCANL 452
Score = 76.6 bits (187), Expect = 3e-14
Identities = 49/151 (32%), Positives = 81/151 (53%)
Frame = +3
Query: 592 TLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVH 651
+++WT+ I + +A+ F++M ++ D +TF SVL ACG + G+QVH
Sbjct: 3 SISWTSMITGFTQNGLDRDAIDIFREMKLENLQMDQYTFGSVLTACGGVMALQE-GKQVH 179
Query: 652 ADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLY 711
A I+ + +V +L+ MY + ++ AE VF+ +NV S AML+GY QNG
Sbjct: 180 AYIIRTDYKDNIFVASALVDMYCKCKNIKSAEAVFKKMTC-KNVVSWTAMLVGYGQNGYS 356
Query: 712 IEAVKFVYQMKAAGVQPHEPLLEKLRIACGS 742
EAVK M+ G++P + L + +C +
Sbjct: 357 EEAVKTFSDMQKYGIEPDDFTLGSVISSCAN 449
Score = 40.8 bits (94), Expect = 0.002
Identities = 24/97 (24%), Positives = 47/97 (47%), Gaps = 2/97 (2%)
Frame = +3
Query: 419 ITIDIYT--SLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENAR 476
+ +D YT S++ C + ++H II + + + + ++ M+ C +++A
Sbjct: 96 LQMDQYTFGSVLTACGGVMALQEGKQVHAYIIRTDYKDNIFVASALVDMYCKCKNIKSAE 275
Query: 477 RVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSM 513
VF M+ ++ SW + V Y +NG E A+ F M
Sbjct: 276 AVFKKMTCKNVVSWTAMLVGYGQNGYSEEAVKTFSDM 386
>AW257410 weakly similar to GP|10177533|db selenium-binding protein-like
{Arabidopsis thaliana}, partial (21%)
Length = 724
Score = 81.6 bits (200), Expect = 1e-15
Identities = 57/214 (26%), Positives = 97/214 (44%), Gaps = 32/214 (14%)
Frame = +1
Query: 547 QVHGCLLKLGACDHVLISSSLIRFYG--RFKCLEDANMVFNRVSRHNTLTWTAKIVSSCR 604
Q++ LK G H L S L+ Y F L A MVF+R+S NT+ W I +
Sbjct: 31 QIYVQFLKKGTIRHKLTVSRLLTTYASMEFSNLTYARMVFDRISSPNTVMWNTMIRAYSN 210
Query: 605 ERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSY 664
EAL + +M + +++TF +LKAC + Q+H IK G S+ Y
Sbjct: 211 SNDPEEALLLYHQMLHHSIPHNAYTFPFLLKACSALSALAET-HQIHVQIIKRGFGSEVY 387
Query: 665 VQCSLIAMYGRSGLLRDAELVFEMTRN------------------------------ERN 694
SL+ +Y SG ++ A ++F++ + E+N
Sbjct: 388 ATNSLLRVYAISGSIKSAHVLFDLLPSRDIVSWNTMIDGYIKCGNVEMAYKIFQAMPEKN 567
Query: 695 VDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQP 728
V S +M++G+++ G++ +A+ + QM A ++P
Sbjct: 568 VISWTSMIVGFVRTGMHKKALSLLQQMLVARIKP 669
Score = 52.8 bits (125), Expect = 5e-07
Identities = 39/161 (24%), Positives = 78/161 (48%), Gaps = 3/161 (1%)
Frame = +1
Query: 442 ELHTQIITRGIELPLTLLNRILIMFVSCGL--LENARRVFDVMSVRDFHSWATLFVSYYE 499
+++ Q + +G ++R+L + S L AR VFD +S + W T+ +Y
Sbjct: 31 QIYVQFLKKGTIRHKLTVSRLLTTYASMEFSNLTYARMVFDRISSPNTVMWNTMIRAYSN 210
Query: 500 NGEYENAIDVFVSMLCQ-LDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGAC 558
+ + E A+ ++ ML + ++FP LLKAC+ + Q+H ++K G
Sbjct: 211 SNDPEEALLLYHQMLHHSIPHNAYTFP-----FLLKACSALSALAETHQIHVQIIKRGFG 375
Query: 559 DHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKI 599
V ++SL+R Y ++ A+++F+ + + ++W I
Sbjct: 376 SEVYATNSLLRVYAISGSIKSAHVLFDLLPSRDIVSWNTMI 498
Score = 52.8 bits (125), Expect = 5e-07
Identities = 33/113 (29%), Positives = 54/113 (47%), Gaps = 2/113 (1%)
Frame = +1
Query: 403 DTTSHILPLMDALHFPITIDIYTS--LVKECTLSTDPETAIELHTQIITRGIELPLTLLN 460
D +L LH I + YT L+K C+ + ++H QII RG + N
Sbjct: 217 DPEEALLLYHQMLHHSIPHNAYTFPFLLKACSALSALAETHQIHVQIIKRGFGSEVYATN 396
Query: 461 RILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSM 513
+L ++ G +++A +FD++ RD SW T+ Y + G E A +F +M
Sbjct: 397 SLLRVYAISGSIKSAHVLFDLLPSRDIVSWNTMIDGYIKCGNVEMAYKIFQAM 555
>TC83158 weakly similar to PIR|H86191|H86191 hypothetical protein [imported]
- Arabidopsis thaliana, partial (42%)
Length = 1194
Score = 79.7 bits (195), Expect = 4e-15
Identities = 61/215 (28%), Positives = 110/215 (50%), Gaps = 2/215 (0%)
Frame = +2
Query: 470 GLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIW 529
G +++A ++FD + V++ SW + + + YE A++ F M QL + F I
Sbjct: 2 GDVDDALKLFDKLPVKNVVSWTVVIGGFVKKECYEEALECFREM--QLAGVVPDFVTVI- 172
Query: 530 SCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSR 589
++ ACA + LG+ VH ++K D+V + +SLI Y R C+E A VF+ +S+
Sbjct: 173 -AIISACANLGALGLGLWVHRLVMKKEFRDNVKVLNSLIDMYARCGCIELARQVFDGMSQ 349
Query: 590 HNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQ 649
N ++W + IV +AL F+ M + G++ + +++S L AC G +
Sbjct: 350 RNLVSWNSIIVGFAVNGLADKALSFFRSMKKEGLEPNGVSYTSALTACSHAGLIDE-GLK 526
Query: 650 VHADAIKLGLDSD--SYVQCSLIAMYGRSGLLRDA 682
+ AD + +S + C L+ +Y R+G L++A
Sbjct: 527 IFADIKRDHRNSPRIEHYGC-LVDLYSRAGRLKEA 628
Score = 75.9 bits (185), Expect = 5e-14
Identities = 50/164 (30%), Positives = 84/164 (50%)
Frame = +2
Query: 577 LEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKA 636
++DA +F+++ N ++WT I ++ + EAL F++M GV D T +++ A
Sbjct: 8 VDDALKLFDKLPVKNVVSWTVVIGGFVKKECYEEALECFREMQLAGVVPDFVTVIAIISA 187
Query: 637 CGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVD 696
C + G G VH +K + V SLI MY R G + A VF+ ++RN+
Sbjct: 188 CANLGALG-LGLWVHRLVMKKEFRDNVKVLNSLIDMYARCGCIELARQVFD-GMSQRNLV 361
Query: 697 SLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIAC 740
S N++++G+ NGL +A+ F MK G++P+ AC
Sbjct: 362 SWNSIIVGFAVNGLADKALSFFRSMKKEGLEPNGVSYTSALTAC 493
Score = 51.6 bits (122), Expect = 1e-06
Identities = 47/206 (22%), Positives = 94/206 (44%), Gaps = 9/206 (4%)
Frame = +2
Query: 443 LHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGE 502
+H ++ + + +LN ++ M+ CG +E AR+VFD MS R+ SW ++ V + NG
Sbjct: 224 VHRLVMKKEFRDNVKVLNSLIDMYARCGCIELARQVFDGMSQRNLVSWNSIIVGFAVNGL 403
Query: 503 YENAIDVFVSMLCQ-LDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDH- 560
+ A+ F SM + L+ G S+ + L AC+ + G+++ + + DH
Sbjct: 404 ADKALSFFRSMKKEGLEPNGVSY-----TSALTACSHAGLIDEGLKIFADIKR----DHR 556
Query: 561 ----VLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRER---HFSEALG 613
+ L+ Y R L++A V ++ ++++CR + +E +
Sbjct: 557 NSPRIEHYGCLVDLYSRAGRLKEAWDVIKKMPMMPNEVVLGSLLAACRTQGDVELAEKVM 736
Query: 614 DFKKMGRVGVKKDSFTFSSVLKACGR 639
++ G + FS++ A G+
Sbjct: 737 KYQVELYPGGDSNYVLFSNIYAAVGK 814
>TC90451 similar to PIR|H84508|H84508 hypothetical protein At2g13600
[imported] - Arabidopsis thaliana, partial (12%)
Length = 1362
Score = 79.3 bits (194), Expect = 5e-15
Identities = 69/240 (28%), Positives = 108/240 (44%), Gaps = 2/240 (0%)
Frame = +2
Query: 489 SWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQV 548
+W + SY +NA+ ++VSML G + +LKA + + + LG QV
Sbjct: 386 NWNNIIRSYTRLESPQNALRIYVSMLRA----GVLPDRYTLPIVLKAVSQSFAIQLGQQV 553
Query: 549 HGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHF 608
H +KLG + S I Y + + A+ VF+ +W A I +
Sbjct: 554 HSYGIKLGLQSNEYCESGFINLYCKAGDFDSAHKVFDENHEPKLGSWNALISGLSQGGLA 733
Query: 609 SEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYV--Q 666
+A+ F M R G + D T SV+ ACG + + Q+H + + + +
Sbjct: 734 MDAIVVFVDMKRHGFEPDGITMVSVMSACGSIGDL-YLALQLHKYVFQAKTNEWTVILMS 910
Query: 667 CSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGV 726
SLI MYG+ G + A VF T +RNV S +M++GY +G EA+ + MK GV
Sbjct: 911 NSLIDMYGKCGRMDLAYEVF-ATMEDRNVSSWTSMIVGYAMHGHAKEALGCFHCMKGVGV 1087
Score = 64.7 bits (156), Expect(2) = 4e-12
Identities = 71/264 (26%), Positives = 108/264 (40%), Gaps = 6/264 (2%)
Frame = +2
Query: 366 LLRLPLRRNPKPKNLSLIHPSSQPITPPKKSKRRRKCDTTSHILPLMDALHFPITIDIYT 425
LL L NP N + I S + P+ + R + +LP D PI +
Sbjct: 347 LLTRFLESNPASFNWNNIIRSYTRLESPQNALRIYVSMLRAGVLP--DRYTLPIVLK--- 511
Query: 426 SLVKECTLSTDPETAIELHTQIITRGIELPLT----LLNRILIMFVSCGLLENARRVFDV 481
+ AI+L Q+ + GI+L L + + ++ G ++A +VFD
Sbjct: 512 --------AVSQSFAIQLGQQVHSYGIKLGLQSNEYCESGFINLYCKAGDFDSAHKVFDE 667
Query: 482 MSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMN 541
SW L + G +AI VFV M GF ++ AC +
Sbjct: 668 NHEPKLGSWNALISGLSQGGLAMDAIVVFVDMKRH----GFEPDGITMVSVMSACGSIGD 835
Query: 542 VPLGMQVHGCLL--KLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKI 599
+ L +Q+H + K +L+S+SLI YG+ ++ A VF + N +WT+ I
Sbjct: 836 LYLALQLHKYVFQAKTNEWTVILMSNSLIDMYGKCGRMDLAYEVFATMEDRNVSSWTSMI 1015
Query: 600 VSSCRERHFSEALGDFKKMGRVGV 623
V H EALG F M VGV
Sbjct: 1016VGYAMHGHAKEALGCFHCMKGVGV 1087
Score = 25.0 bits (53), Expect(2) = 4e-12
Identities = 22/72 (30%), Positives = 34/72 (46%), Gaps = 3/72 (4%)
Frame = +1
Query: 619 GRVGVKKDSFTFSSVLKACGRMQNRGSCGE-QVHADAIK--LGLDSDSYVQCSLIAMYGR 675
G GVK + TF VL AC + G+ E + + D +K G+ ++ + GR
Sbjct: 1075 GSRGVKPNYVTFIGVLSAC---VHGGTVQEGRFYFDMMKNIYGITPQLQHYGCMVDLLGR 1245
Query: 676 SGLLRDAELVFE 687
+GL DA + E
Sbjct: 1246 AGLFDDARRMVE 1281
>AW585413 weakly similar to PIR|B85153|B851 hypothetical protein AT4g14050
[imported] - Arabidopsis thaliana, partial (5%)
Length = 577
Score = 73.6 bits (179), Expect = 3e-13
Identities = 34/105 (32%), Positives = 63/105 (59%)
Frame = +1
Query: 533 LKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNT 592
L +CA T+N LG+Q+H +++ G D++ + S+L+ FY + + DAN +F + +H+
Sbjct: 208 LSSCAKTLNWHLGIQIHAYMIRSGYEDNLFLCSALVDFYAKCFAIVDANKIFRAMKQHDQ 387
Query: 593 LTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKAC 637
++WT+ I + +AL FK+M ++ + FT +SV+ AC
Sbjct: 388 VSWTSLIAGFSANKQGRDALLLFKEMLGTQIRPNCFTLTSVINAC 522
>BE202961 weakly similar to GP|6016735|gb|A hypothetical protein {Arabidopsis
thaliana}, partial (14%)
Length = 613
Score = 70.9 bits (172), Expect = 2e-12
Identities = 47/179 (26%), Positives = 88/179 (48%), Gaps = 3/179 (1%)
Frame = +2
Query: 426 SLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVR 485
S+++ C ++ E +++H + G E+ T+ ++ M++ C E A +F+ M +
Sbjct: 62 SVLRACACISNLEEGMKIHELAVNYGFEMETTVSTALMDMYMKCFSPEKAVDIFNRMPKK 241
Query: 486 DFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLG 545
D +WA LF Y +NG ++ VF +ML + P I L+K + +
Sbjct: 242 DVIAWAVLFSGYADNGMVHESMWVFRNML-----SSGTRPDAI--ALVKILTTVSELGIL 400
Query: 546 MQV---HGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVS 601
Q H ++K G ++ I +SLI Y + +EDAN VF ++ + +TW++ I +
Sbjct: 401 QQAVCFHAFVIKNGFENNQFIGASLIEVYAKCSSIEDANKVFKGMTYKDVVTWSSIIAA 577
Score = 67.0 bits (162), Expect = 3e-11
Identities = 48/195 (24%), Positives = 91/195 (46%), Gaps = 3/195 (1%)
Frame = +2
Query: 525 PPWIWSC-LLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMV 583
P W+ +L+ACAC N+ GM++H + G +S++L+ Y + E A +
Sbjct: 41 PNWVTVVSVLRACACISNLEEGMKIHELAVNYGFEMETTVSTALMDMYMKCFSPEKAVDI 220
Query: 584 FNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNR 643
FNR+ + + + W E++ F+ M G + D+ +L +
Sbjct: 221 FNRMPKKDVIAWAVLFSGYADNGMVHESMWVFRNMLSSGTRPDAIALVKILTTVSEL--- 391
Query: 644 GSCGEQV--HADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAM 701
G + V HA IK G +++ ++ SLI +Y + + DA VF+ ++V + +++
Sbjct: 392 GILQQAVCFHAFVIKNGFENNQFIGASLIEVYAKCSSIEDANKVFK-GMTYKDVVTWSSI 568
Query: 702 LMGYIQNGLYIEAVK 716
+ Y +G EA+K
Sbjct: 569 IAAYGFHGQGEEALK 613
>BQ138324 weakly similar to GP|12322902|gb| hypothetical protein; 50785-52656
{Arabidopsis thaliana}, partial (7%)
Length = 619
Score = 67.8 bits (164), Expect = 1e-11
Identities = 42/130 (32%), Positives = 66/130 (50%)
Frame = +1
Query: 592 TLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVH 651
T W + ++R F EAL ++ M R ++FTF +LK+C + + G Q+H
Sbjct: 67 TTAWNCYLRELSKQRKFMEALTVYRHMLRSSFFPNTFTFPVLLKSCALL-SLPFTGSQLH 243
Query: 652 ADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLY 711
+ +K G D Y SLI MY ++ L A VF+ + + S NAM+ GY N +
Sbjct: 244 SHILKTGSQPDPYTHSSLINMYSKTSLPCLARKVFDESPVNLTI-SYNAMISGYTNNMMI 420
Query: 712 IEAVKFVYQM 721
+EA+K +M
Sbjct: 421 VEAIKLFRRM 450
Score = 50.1 bits (118), Expect = 3e-06
Identities = 43/176 (24%), Positives = 74/176 (41%)
Frame = +1
Query: 396 SKRRRKCDTTSHILPLMDALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELP 455
SK+R+ + + ++ + FP T + L+K C L + P T +LH+ I+ G +
Sbjct: 100 SKQRKFMEALTVYRHMLRSSFFPNTFT-FPVLLKSCALLSLPFTGSQLHSHILKTGSQPD 276
Query: 456 LTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLC 515
+ ++ M+ L AR+VFD V S+ + Y N AI +F MLC
Sbjct: 277 PYTHSSLINMYSKTSLPCLARKVFDESPVNLTISYNAMISGYTNNMMIVEAIKLFRRMLC 456
Query: 516 QLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFY 571
+ F L+ + LG +HGC K G + + + + + Y
Sbjct: 457 E---NRFFVNSVTMLGLVSGILVPEKLRLGFCLHGCCFKFGFENDLXVGNXFLTMY 615
Score = 39.7 bits (91), Expect = 0.004
Identities = 41/156 (26%), Positives = 66/156 (42%), Gaps = 4/156 (2%)
Frame = +1
Query: 522 FSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDAN 581
F+FP LLK+CA G Q+H +LK G+ SSLI Y + A
Sbjct: 175 FTFP-----VLLKSCALLSLPFTGSQLHSHILKTGSQPDPYTHSSLINMYSKTSLPCLAR 339
Query: 582 MVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSV----LKAC 637
VF+ + T+++ A I EA+ F++M + ++ F +SV L +
Sbjct: 340 KVFDESPVNLTISYNAMISGYTNNMMIVEAIKLFRRM----LCENRFFVNSVTMLGLVSG 507
Query: 638 GRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMY 673
+ + G +H K G ++D V + MY
Sbjct: 508 ILVPEKLRLGFCLHGCCFKFGFENDLXVGNXFLTMY 615
>BF640739 weakly similar to GP|7363286|dbj Similar to Arabidopsis thaliana
DNA chromosome 4 BAC clone F4I10; vegetative storage
protein., partial (14%)
Length = 444
Score = 65.1 bits (157), Expect = 1e-10
Identities = 45/138 (32%), Positives = 74/138 (53%), Gaps = 1/138 (0%)
Frame = +1
Query: 609 SEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCS 668
+EAL F M G+K DSFTF SV+ A + + +H AI+ +D++ +V +
Sbjct: 4 NEALNLFCTMQSQGIKPDSFTFVSVITALADLSVTRQ-AKWIHGLAIRTNMDTNVFVATA 180
Query: 669 LIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMK-AAGVQ 727
L+ MY + G + A +F+M + ER+V + NAM+ GY +GL A+ M+ A ++
Sbjct: 181 LVDMYAKCGAIETARELFDMMQ-ERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEASLK 357
Query: 728 PHEPLLEKLRIACGSSNF 745
P++ + AC S F
Sbjct: 358 PNDITFLSVISACSHSGF 411
Score = 40.4 bits (93), Expect = 0.003
Identities = 19/71 (26%), Positives = 36/71 (49%)
Frame = +1
Query: 443 LHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGE 502
+H I ++ + + ++ M+ CG +E AR +FD+M R +W + Y +G
Sbjct: 124 IHGLAIRTNMDTNVFVATALVDMYAKCGAIETARELFDMMQERHVITWNAMIDGYGTHGL 303
Query: 503 YENAIDVFVSM 513
+ A+D+F M
Sbjct: 304 GKAALDLFDDM 336
>AW696172 weakly similar to PIR|H96620|H96 hypothetical protein F23H11.3
[imported] - Arabidopsis thaliana, partial (19%)
Length = 649
Score = 63.2 bits (152), Expect = 4e-10
Identities = 52/169 (30%), Positives = 78/169 (45%), Gaps = 5/169 (2%)
Frame = +2
Query: 583 VFNRVSRHNTLTWTAKIVSSCRER-HFSEALGDFKKMGRVGVKK---DSFTFSSVLKACG 638
+ + N+ TW I S + H +A+ +K + + D T+ VLKAC
Sbjct: 134 ILRTIHTPNSFTWNILIQSYSKSTLHKQKAILLYKAIITEQENELFPDKHTYPFVLKACA 313
Query: 639 RMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSL 698
+ + G+QVHA +KLG + D+Y+ SLI Y G L A VF+ RNV S
Sbjct: 314 YLFSLFE-GKQVHAHVLKLGFELDTYICNSLIHFYASCGYLETARKVFDRMCEWRNVVSW 490
Query: 699 NAMLMGYIQNGLY-IEAVKFVYQMKAAGVQPHEPLLEKLRIACGSSNFS 746
N M+ Y + G Y I + F MK +P ++ + ACG F+
Sbjct: 491 NVMIDSYAKVGDYDIVLIMFCEMMKV--YEPDCYTMQSVIRACGGFRFA 631
Score = 58.9 bits (141), Expect = 7e-09
Identities = 40/108 (37%), Positives = 57/108 (52%), Gaps = 1/108 (0%)
Frame = +2
Query: 532 LLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSR-H 590
+LKACA ++ G QVH +LKLG I +SLI FY LE A VF+R+
Sbjct: 296 VLKACAYLFSLFEGKQVHAHVLKLGFELDTYICNSLIHFYASCGYLETARKVFDRMCEWR 475
Query: 591 NTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACG 638
N ++W I S + + L F +M +V + D +T SV++ACG
Sbjct: 476 NVVSWNVMIDSYAKVGDYDIVLIMFCEMMKV-YEPDCYTMQSVIRACG 616
Score = 53.1 bits (126), Expect = 4e-07
Identities = 28/92 (30%), Positives = 47/92 (50%), Gaps = 1/92 (1%)
Frame = +2
Query: 424 YTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVM- 482
Y ++K C ++H ++ G EL + N ++ + SCG LE AR+VFD M
Sbjct: 287 YPFVLKACAYLFSLFEGKQVHAHVLKLGFELDTYICNSLIHFYASCGYLETARKVFDRMC 466
Query: 483 SVRDFHSWATLFVSYYENGEYENAIDVFVSML 514
R+ SW + SY + G+Y+ + +F M+
Sbjct: 467 EWRNVVSWNVMIDSYAKVGDYDIVLIMFCEMM 562
>TC89987 weakly similar to PIR|D86443|D86443 probable PPR-repeat protein
[imported] - Arabidopsis thaliana, partial (62%)
Length = 1559
Score = 60.8 bits (146), Expect = 2e-09
Identities = 44/146 (30%), Positives = 74/146 (50%), Gaps = 1/146 (0%)
Frame = +1
Query: 535 ACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLT 594
AC+ V G+QVHG + K+G V++ +SLI YG+ +++A VFN + + +
Sbjct: 130 ACSLLGVVDEGIQVHGHVFKMGLEGDVIVQNSLINMYGKCGEIKNACDVFNGMDEKSVAS 309
Query: 595 WTAKIVSSCRERHFSEALGDFKKMGRVG-VKKDSFTFSSVLKACGRMQNRGSCGEQVHAD 653
W+A I + ++E L KM G + + T +VL AC + G+ +H
Sbjct: 310 WSAIIGAHACVEMWNECLMLLGKMSSEGRCRVEESTLVNVLSACTHL-GSPDLGKCIHGI 486
Query: 654 AIKLGLDSDSYVQCSLIAMYGRSGLL 679
++ + + V+ SLI MY +SG L
Sbjct: 487 LLRNISELNVVVKTSLIDMYVKSGCL 564
Score = 49.3 bits (116), Expect = 5e-06
Identities = 39/164 (23%), Positives = 76/164 (45%), Gaps = 4/164 (2%)
Frame = +1
Query: 431 CTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSW 490
C+L + I++H + G+E + + N ++ M+ CG ++NA VF+ M + SW
Sbjct: 133 CSLLGVVDEGIQVHGHVFKMGLEGDVIVQNSLINMYGKCGEIKNACDVFNGMDEKSVASW 312
Query: 491 ATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSC----LLKACACTMNVPLGM 546
+ + ++ ++++ L L M + +L AC + LG
Sbjct: 313 SAIIGAH-------ACVEMWNECLMLLGKMSSEGRCRVEESTLVNVLSACTHLGSPDLGK 471
Query: 547 QVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRH 590
+HG LL+ + +V++ +SLI Y + CLE F R+ ++
Sbjct: 472 CIHGILLRNISELNVVVKTSLIDMYVKSGCLEKR---FTRIQKY 594
Score = 44.3 bits (103), Expect = 2e-04
Identities = 31/95 (32%), Positives = 52/95 (54%), Gaps = 1/95 (1%)
Frame = +1
Query: 647 GEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYI 706
G QVH K+GL+ D VQ SLI MYG+ G +++A VF +E++V S +A++ +
Sbjct: 160 GIQVHGHVFKMGLEGDVIVQNSLINMYGKCGEIKNACDVFN-GMDEKSVASWSAIIGAHA 336
Query: 707 QNGLYIEAVKFVYQMKAAG-VQPHEPLLEKLRIAC 740
++ E + + +M + G + E L + AC
Sbjct: 337 CVEMWNECLMLLGKMSSEGRCRVEESTLVNVLSAC 441
Score = 33.1 bits (74), Expect = 0.40
Identities = 37/159 (23%), Positives = 59/159 (36%), Gaps = 19/159 (11%)
Frame = +2
Query: 556 GACDHVLISSSLI------RFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFS 609
G+C L++S L+ R L VF +S N ++T I
Sbjct: 482 GSC*GTLVNSMLL*RLR*LICMSRVDVLRKGLRVFKNMSEKNRYSYTVMISGLAIHGRGK 661
Query: 610 EALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCS- 668
EAL F +M G+ D + V AC HA ++ GL +Q
Sbjct: 662 EALKVFSEMIEEGLAPDDVVYVGVFSACS------------HAGLVEEGLQCFKSMQFEH 805
Query: 669 -----------LIAMYGRSGLLRDA-ELVFEMTRNERNV 695
++ + GR G+L++A EL+ M+ +V
Sbjct: 806 KIEPTVQHYGCMVDLLGRFGMLKEAYELIKSMSIKPNDV 922
>BF646834 weakly similar to PIR|A84784|A847 hypothetical protein At2g36730
[imported] - Arabidopsis thaliana, partial (10%)
Length = 602
Score = 59.3 bits (142), Expect = 5e-09
Identities = 50/147 (34%), Positives = 72/147 (48%), Gaps = 3/147 (2%)
Frame = +2
Query: 544 LGMQVHGCLLKLGACDHVLISSSLIRFYGR--FKCLEDAN-MVFNRVSRHNTLTWTAKIV 600
L Q+H L L H+L S L+ F+ FK L A +VF+ + + ++W I
Sbjct: 122 LQAQIH--LNSLHNDTHIL--SQLVYFFSLSPFKNLSHARKLVFHFSNNPSPISWNILIR 289
Query: 601 SSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLD 660
E++ FKKM GVK + T+ + K+C M G+QVHAD +K GLD
Sbjct: 290 GYASSDSPIESIWVFKKMRENGVKPNKLTYPFIFKSCA-MALVLCEGKQVHADLVKFGLD 466
Query: 661 SDSYVQCSLIAMYGRSGLLRDAELVFE 687
SD YV ++I YG + A VF+
Sbjct: 467 SDVYVCNNMINFYGXCKKIVYARKVFD 547
Score = 43.9 bits (102), Expect = 2e-04
Identities = 40/165 (24%), Positives = 70/165 (42%), Gaps = 3/165 (1%)
Frame = +2
Query: 442 ELHTQIITRGIELPLTLLNRILIMFVSCGL--LENARR-VFDVMSVRDFHSWATLFVSYY 498
+L QI + +L++++ F L +AR+ VF + SW L Y
Sbjct: 119 QLQAQIHLNSLHNDTHILSQLVYFFSLSPFKNLSHARKLVFHFSNNPSPISWNILIRGYA 298
Query: 499 ENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGAC 558
+ +I VF M G + + K+CA + + G QVH L+K G
Sbjct: 299 SSDSPIESIWVFKKMREN----GVKPNKLTYPFIFKSCAMALVLCEGKQVHADLVKFGLD 466
Query: 559 DHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSC 603
V + +++I FYG K + A VF+ + ++W + + + C
Sbjct: 467 SDVYVCNNMINFYGXCKKIVYARKVFDEMCVXTIVSWNSVMTAWC 601
Score = 35.4 bits (80), Expect = 0.081
Identities = 16/74 (21%), Positives = 36/74 (48%)
Frame = +2
Query: 424 YTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMS 483
Y + K C ++ ++H ++ G++ + + N ++ + C + AR+VFD M
Sbjct: 377 YPFIFKSCAMALVLCEGKQVHADLVKFGLDSDVYVCNNMINFYGXCKKIVYARKVFDEMC 556
Query: 484 VRDFHSWATLFVSY 497
V SW ++ ++
Sbjct: 557 VXTIVSWNSVMTAW 598
>BE320932 weakly similar to PIR|T04548|T045 hypothetical protein F28J12.180 -
Arabidopsis thaliana, partial (14%)
Length = 354
Score = 58.2 bits (139), Expect = 1e-08
Identities = 37/108 (34%), Positives = 53/108 (48%)
Frame = +3
Query: 545 GMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSSCR 604
G Q+HG ++K V I +SLI Y + + + VF+R+ NT TWT+ I R
Sbjct: 33 GTQLHGAIVKKICKSDVFIGTSLIDMYAKCGEIVSSKKVFDRMKVRNTATWTSIISGYAR 212
Query: 605 ERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHA 652
EAL F+ M R V + T V+ ACG ++ G +VHA
Sbjct: 213 NGFGEEALNFFRLMKRKKVYVNKSTLVCVMTACGTIK-ASLIGREVHA 353
Score = 52.8 bits (125), Expect = 5e-07
Identities = 31/97 (31%), Positives = 51/97 (51%)
Frame = +3
Query: 646 CGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGY 705
CG Q+H +K SD ++ SLI MY + G + ++ VF+ + RN + +++ GY
Sbjct: 30 CGTQLHGAIVKKICKSDVFIGTSLIDMYAKCGEIVSSKKVFDRMK-VRNTATWTSIISGY 206
Query: 706 IQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIACGS 742
+NG EA+ F MK V ++ L + ACG+
Sbjct: 207 ARNGFGEEALNFFRLMKRKKVYVNKSTLVCVMTACGT 317
Score = 45.8 bits (107), Expect = 6e-05
Identities = 27/108 (25%), Positives = 53/108 (49%)
Frame = +3
Query: 442 ELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENG 501
+LH I+ + + + + ++ M+ CG + ++++VFD M VR+ +W ++ Y NG
Sbjct: 39 QLHGAIVKKICKSDVFIGTSLIDMYAKCGEIVSSKKVFDRMKVRNTATWTSIISGYARNG 218
Query: 502 EYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVH 549
E A++ F M + + S C++ AC +G +VH
Sbjct: 219 FGEEALNFFRLMKRKKVYVNKS----TLVCVMTACGTIKASLIGREVH 350
>TC84835 weakly similar to PIR|T02328|T02328 hypothetical protein At2g34400
[imported] - Arabidopsis thaliana, partial (5%)
Length = 647
Score = 55.8 bits (133), Expect = 6e-08
Identities = 39/140 (27%), Positives = 67/140 (47%)
Frame = +1
Query: 583 VFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQN 642
+FN + + I S + F + F ++ G+ D++T+ VLKA + +
Sbjct: 73 IFNHTLHPSLFLYNLLIKSFFKRNSFQTLISLFNQLRLNGLYPDNYTYPFVLKAVAFIAD 252
Query: 643 RGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAML 702
G ++HA K GLDSD YV S + MY G + +F+ +ER+ S N M+
Sbjct: 253 FRQ-GTKIHAFVFKTGLDSDYYVSNSFMDMYAELGRIDFVRKLFDEI-SERDSVSWNVMI 426
Query: 703 MGYIQNGLYIEAVKFVYQMK 722
G ++ + EAV+ +M+
Sbjct: 427 SGCVKCRRFEEAVEVFQRMR 486
Score = 52.0 bits (123), Expect = 8e-07
Identities = 32/131 (24%), Positives = 62/131 (46%)
Frame = +1
Query: 424 YTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMS 483
Y ++K D ++H + G++ + N + M+ G ++ R++FD +S
Sbjct: 214 YPFVLKAVAFIADFRQGTKIHAFVFKTGLDSDYYVSNSFMDMYAELGRIDFVRKLFDEIS 393
Query: 484 VRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVP 543
RD SW + + +E A++VF M ++D + S L ACA + NV
Sbjct: 394 ERDSVSWNVMISGCVKCRRFEEAVEVFQRM--RVDSNEKISEATVVSS-LTACAASRNVE 564
Query: 544 LGMQVHGCLLK 554
+G ++HG +++
Sbjct: 565 VGKEIHGFIIE 597
>TC80535 weakly similar to GP|14335136|gb|AAK59848.1 At2g15690/F9O13.24
{Arabidopsis thaliana}, partial (55%)
Length = 1727
Score = 54.7 bits (130), Expect = 1e-07
Identities = 41/164 (25%), Positives = 68/164 (41%)
Frame = +2
Query: 532 LLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHN 591
L C + +V +VH L+ + + +I YG K + DA VF+ + N
Sbjct: 299 LFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRN 478
Query: 592 TLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVH 651
+W I E L F++M +G++ S T +VL ACG + +
Sbjct: 479 MDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLE 658
Query: 652 ADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNV 695
+ K G++ L+ + G+SG L++AE E E V
Sbjct: 659 SMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTV 790
Score = 47.8 bits (112), Expect = 2e-05
Identities = 53/217 (24%), Positives = 82/217 (37%), Gaps = 18/217 (8%)
Frame = +2
Query: 338 PPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIH--------PSSQP 389
PPPP P R+ PP+ P+ + NP+ NL+ P+ Q
Sbjct: 5 PPPPQNPNFRQPIRQNPNFQPPNG--PNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQE 178
Query: 390 ITPPKKS--KRRRKCDT--TSHILPLM------DALHFPITIDIYTSLVKECTLSTDPET 439
PP S R C L LM DA F I D+ C S E
Sbjct: 179 QAPPPPSIVDLTRFCQEGKVKEALELMEKGIKADANCFEILFDL-------CGKSKSVED 337
Query: 440 AIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYE 499
A ++H + + N+++ M+ +C + +ARRVFD M R+ SW + Y
Sbjct: 338 AKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYAN 517
Query: 500 NGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKAC 536
+ + + +F Q++ +G +L AC
Sbjct: 518 STMGDEGLQLFE----QMNELGLEITSETMLAVLSAC 616
>TC82019 weakly similar to GP|11994382|dbj|BAB02341.
gb|AAB82628.1~gene_id:MIL23.2~similar to unknown protein
{Arabidopsis thaliana}, partial (7%)
Length = 737
Score = 53.5 bits (127), Expect = 3e-07
Identities = 38/119 (31%), Positives = 58/119 (47%)
Frame = +1
Query: 622 GVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRD 681
G D F S+LK C +++ G++VH + + + LI +Y + G ++D
Sbjct: 316 GAFADYSDFLSLLKLCEDLKSL-ELGKRVHEFLRRSKFGGNVELCNRLIGLYVKCGSVKD 492
Query: 682 AELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIAC 740
A VF+ +RNV SLN M+ GY NGL I+ + QM+ GV P E + C
Sbjct: 493 ARKVFDKMP-DRNVGSLNLMIGGYNVNGLRIDGLLVFKQMRQQGVVPDEETFALVLAVC 666
Score = 40.4 bits (93), Expect = 0.003
Identities = 39/147 (26%), Positives = 64/147 (43%), Gaps = 9/147 (6%)
Frame = +1
Query: 376 KPKNLSLIHPSSQPITPPKKSKRRRKCDTTSH---------ILPLMDALHFPITIDIYTS 426
K + L + +PS QP ++ + K +H +L LM F D + S
Sbjct: 172 KTQPLRMGNPSIQPKLNHHQAPHQHKNVNFAHFLQEGNVNQVLELMGQGAFADYSD-FLS 348
Query: 427 LVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRD 486
L+K C E +H + + L NR++ ++V CG +++AR+VFD M R+
Sbjct: 349 LLKLCEDLKSLELGKRVHEFLRRSKFGGNVELCNRLIGLYVKCGSVKDARKVFDKMPDRN 528
Query: 487 FHSWATLFVSYYENGEYENAIDVFVSM 513
S + Y NG + + VF M
Sbjct: 529 VGSLNLMIGGYNVNGLRIDGLLVFKQM 609
Score = 39.7 bits (91), Expect = 0.004
Identities = 31/106 (29%), Positives = 49/106 (45%)
Frame = +1
Query: 532 LLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHN 591
LLK C ++ LG +VH L + +V + + LI Y + ++DA VF+++ N
Sbjct: 349 LLKLCEDLKSLELGKRVHEFLRRSKFGGNVELCNRLIGLYVKCGSVKDARKVFDKMPDRN 528
Query: 592 TLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKAC 637
+ I + L FK+M + GV D TF+ VL C
Sbjct: 529 VGSLNLMIGGYNVNGLRIDGLLVFKQMRQQGVVPDEETFALVLAVC 666
>TC92997 similar to GP|21743256|dbj|BAC03254. contains ESTs AU030758(E60201)
AU091411(E60201)~similar to Arabidopsis thaliana
chromosome 5, partial (13%)
Length = 788
Score = 50.4 bits (119), Expect = 2e-06
Identities = 43/213 (20%), Positives = 82/213 (38%), Gaps = 26/213 (12%)
Frame = +2
Query: 23 IERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSLWSHPS----YLTQSLSSTTVSL 78
I R E T F N P ++ +W + SLW+ + YL + S+ V
Sbjct: 35 IRRYEEVLTSNDFESLIEAQNVPAVLCGCTKNWTAFSLWNPRNDGLNYLQDRVGSSVVEA 214
Query: 79 HLTPTG-------SADSLTPLPSSPS-SLCFASAHVQ-----NLPFPEALRLINSSNPSQ 125
++ + + PLP S LC H+Q +L + S+
Sbjct: 215 MISSSAPVFYGDLGSHQRVPLPFSTFLDLCKKRMHMQTQQQQHLDNDHCVASQTDSSQHD 394
Query: 126 CVAYAQQQNDCFRSEYDSIVKDCDQHIAWATEAFGLEP---------EAVNLWIGNKHSS 176
C+++ + ++ + + + + T ++ ++NLW+ N S
Sbjct: 395 CLSFEDIPEQIYLAQVPIMNSNRQEKVQLETLREDIQTPPILGAKDLSSINLWMNNAQSR 574
Query: 177 TWFHKDHYENLYAVVTGQKHFLLFPPTDVHRFY 209
+ H D + NL +V+G+K +L+PP+ Y
Sbjct: 575 SSTHYDPHHNLLCIVSGRKQVVLWPPSASSSLY 673
>TC92948 weakly similar to GP|15623829|dbj|BAB67888. hypothetical
protein~similar to Arabidopsis thaliana chromosome 5
MDH9.15, partial (17%)
Length = 692
Score = 49.7 bits (117), Expect = 4e-06
Identities = 48/227 (21%), Positives = 85/227 (37%), Gaps = 5/227 (2%)
Frame = +1
Query: 373 RNPKPKNLSLIHPSSQPITPPKKSKRRRKCDTTSHILPLMDALHFPITIDIYTSLVKECT 432
R+P P+ S P++ K S + +I+ HFP CT
Sbjct: 82 RHPPPEATSFSLVIPPPVSDMKTSIQHHTKTPIKNIISRFSTPHFPF-----------CT 228
Query: 433 LSTDPETAIE----LHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFH 488
T + + LHT G + + V+C + + + V V +
Sbjct: 229 YQTKTHHSHDKTNFLHTGFQPNGSDS--------ITFDVACNMFDEMPELLTVGLVTE-- 378
Query: 489 SWATLFVSYYENGEYENAIDVFVSMLCQ-LDVMGFSFPPWIWSCLLKACACTMNVPLGMQ 547
+ S+ + +E+AI +F ML + F+F +L V +G Q
Sbjct: 379 ----IITSFSKQSRHEDAIYLFSRMLASTIRPNEFTF-----GTVLNTSTRLGKVGVGKQ 531
Query: 548 VHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLT 594
+HGC +K C +V + S+L+ Y + +E+A F N ++
Sbjct: 532 IHGCAIKTSLCSNVFVGSALVDLYVKLSSIEEAQKAFEDTEYXNVVS 672
Score = 45.8 bits (107), Expect = 6e-05
Identities = 28/94 (29%), Positives = 49/94 (51%)
Frame = +1
Query: 596 TAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAI 655
T I S ++ +A+ F +M ++ + FTF +VL R+ G G+Q+H AI
Sbjct: 373 TEIITSFSKQSRHEDAIYLFSRMLASTIRPNEFTFGTVLNTSTRLGKVG-VGKQIHGCAI 549
Query: 656 KLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMT 689
K L S+ +V +L+ +Y + + +A+ FE T
Sbjct: 550 KTSLCSNVFVGSALVDLYVKLSSIEEAQKAFEDT 651
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.136 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,493,119
Number of Sequences: 36976
Number of extensions: 572598
Number of successful extensions: 7817
Number of sequences better than 10.0: 267
Number of HSP's better than 10.0 without gapping: 4925
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6501
length of query: 749
length of database: 9,014,727
effective HSP length: 103
effective length of query: 646
effective length of database: 5,206,199
effective search space: 3363204554
effective search space used: 3363204554
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC144759.9


