

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144731.10 + phase: 0 /pseudo
(222 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
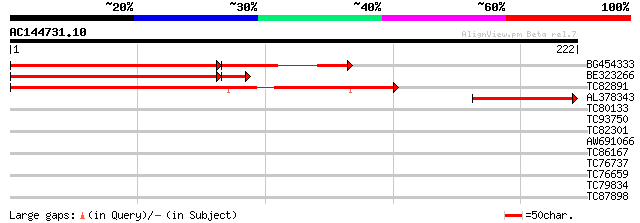
BG454333 similar to PIR|T48571|T48 hypothetical protein T31B5.60... 166 4e-56
BE323266 similar to PIR|T48571|T485 hypothetical protein T31B5.6... 166 2e-43
TC82891 similar to PIR|T48571|T48571 hypothetical protein T31B5.... 168 2e-42
AL378343 weakly similar to GP|21689797|gb| unknown protein {Arab... 87 5e-18
TC80133 similar to PIR|S52728|S52728 H+-transporting ATPase (EC ... 32 0.25
TC93750 weakly similar to PIR|T00534|T00534 S-receptor kinase (E... 30 0.96
TC82301 GP|23496378|gb|AAN36035.1 hypothetical protein {Plasmodi... 28 2.1
AW691066 similar to GP|18491209|gb putative tyrosine decarboxyla... 28 3.7
TC86167 similar to PIR|H96751|H96751 probable casein kinase I F2... 28 3.7
TC76737 homologue to SP|P06379|RK2_TOBAC Chloroplast 50S ribosom... 27 4.8
TC76659 weakly similar to PIR|T09127|T09127 probable erythrocyte... 27 4.8
TC79834 similar to GP|16323504|gb|AAL15246.1 unknown protein {Ar... 27 4.8
TC87898 weakly similar to GP|1174199|gb|AAA86652.1| S25-PR6 {Nic... 27 8.2
>BG454333 similar to PIR|T48571|T48 hypothetical protein T31B5.60 -
Arabidopsis thaliana, partial (43%)
Length = 487
Score = 166 bits (421), Expect(2) = 4e-56
Identities = 83/83 (100%), Positives = 83/83 (100%)
Frame = +2
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 60
MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS
Sbjct: 98 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 277
Query: 61 DNDSPSPVDSLSSPVDSLSARTS 83
DNDSPSPVDSLSSPVDSLSARTS
Sbjct: 278 DNDSPSPVDSLSSPVDSLSARTS 346
Score = 68.6 bits (166), Expect(2) = 4e-56
Identities = 36/51 (70%), Positives = 36/51 (70%)
Frame = +3
Query: 84 PAGRHWFT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQ 134
PAGRHWFT*FLPSITCIQIMIS VLSRFLKPTCLKPQ
Sbjct: 342 PAGRHWFT*FLPSITCIQIMIS---------------VLSRFLKPTCLKPQ 449
>BE323266 similar to PIR|T48571|T485 hypothetical protein T31B5.60 -
Arabidopsis thaliana, partial (26%)
Length = 395
Score = 166 bits (421), Expect(2) = 2e-43
Identities = 83/83 (100%), Positives = 83/83 (100%)
Frame = +2
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 60
MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS
Sbjct: 101 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 280
Query: 61 DNDSPSPVDSLSSPVDSLSARTS 83
DNDSPSPVDSLSSPVDSLSARTS
Sbjct: 281 DNDSPSPVDSLSSPVDSLSARTS 349
Score = 26.2 bits (56), Expect(2) = 2e-43
Identities = 10/11 (90%), Positives = 10/11 (90%)
Frame = +3
Query: 84 PAGRHWFT*FL 94
PAGRHWFT* L
Sbjct: 345 PAGRHWFT*VL 377
>TC82891 similar to PIR|T48571|T48571 hypothetical protein T31B5.60 -
Arabidopsis thaliana, partial (56%)
Length = 637
Score = 168 bits (425), Expect = 2e-42
Identities = 103/166 (62%), Positives = 112/166 (67%), Gaps = 14/166 (8%)
Frame = +1
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 60
MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS
Sbjct: 130 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 309
Query: 61 DNDSPSPVDSLSSPVDSLSARTSP--------AGRHWFT*FLPSITCIQIMISAQLKLTS 112
DNDSPSPVDSLSSPVDSLSARTS A H + + S + A T
Sbjct: 310 DNDSPSPVDSLSSPVDSLSARTSRKTLVYLVLALYHMYPDYDFSA------VKAHQFFTE 471
Query: 113 FSQRNHGTVLSRFLKPTCLK------PQSLLDALFKAVDEVVSLDD 152
S + + ++ + SLLDALFKAVD VSL +
Sbjct: 472 ESWDSFKQIFEAYMFEASKEWAETFGGVSLLDALFKAVDXGVSLXE 609
>AL378343 weakly similar to GP|21689797|gb| unknown protein {Arabidopsis
thaliana}, partial (18%)
Length = 483
Score = 87.0 bits (214), Expect = 5e-18
Identities = 41/41 (100%), Positives = 41/41 (100%)
Frame = +1
Query: 182 NRKLKRIVTFRLSCFSNLIADGFSLDEICDEYDEEIFADMD 222
NRKLKRIVTFRLSCFSNLIADGFSLDEICDEYDEEIFADMD
Sbjct: 1 NRKLKRIVTFRLSCFSNLIADGFSLDEICDEYDEEIFADMD 123
>TC80133 similar to PIR|S52728|S52728 H+-transporting ATPase (EC 3.6.1.35) -
kidney bean, partial (20%)
Length = 788
Score = 31.6 bits (70), Expect = 0.25
Identities = 15/40 (37%), Positives = 25/40 (62%)
Frame = -3
Query: 47 SLSNEILDYLGKSSDNDSPSPVDSLSSPVDSLSARTSPAG 86
+++N LD + ++ +ND+ P DS SP +SL + SP G
Sbjct: 279 AIANCKLDNVKRNIENDTVKPDDSCPSPTNSLDSCKSPVG 160
>TC93750 weakly similar to PIR|T00534|T00534 S-receptor kinase (EC 2.7.1.-)
T20K24.15 precursor - Arabidopsis thaliana, partial (7%)
Length = 558
Score = 29.6 bits (65), Expect = 0.96
Identities = 25/63 (39%), Positives = 34/63 (53%), Gaps = 3/63 (4%)
Frame = -2
Query: 96 SITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLD-ALFKAVDEVVS--LDD 152
+I ++I AQL +T +RNH V S+F CLK L+D LF A +E V L
Sbjct: 383 TIKYLKIKTMAQLIVTQAIERNH*NVSSKF----CLKD*VLVDEMLFLA*EEAVETLLFT 216
Query: 153 CEI 155
CE+
Sbjct: 215 CEL 207
>TC82301 GP|23496378|gb|AAN36035.1 hypothetical protein {Plasmodium
falciparum 3D7}, partial (3%)
Length = 712
Score = 28.5 bits (62), Expect = 2.1
Identities = 17/56 (30%), Positives = 30/56 (53%), Gaps = 4/56 (7%)
Frame = +3
Query: 103 MISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAV----DEVVSLDDCE 154
M S + ++ + + +L+RFL+ +CL P+ LL LF + D V +D C+
Sbjct: 345 MFSFPIAFSAATTDSRSFILTRFLRGSCLSPKWLL-TLFSLITFEADWTVKIDCCD 509
>AW691066 similar to GP|18491209|gb putative tyrosine decarboxylase
{Arabidopsis thaliana}, partial (41%)
Length = 648
Score = 27.7 bits (60), Expect = 3.7
Identities = 19/68 (27%), Positives = 32/68 (46%), Gaps = 8/68 (11%)
Frame = +3
Query: 55 YLGKSSDNDSPSPVDSLSSPVDSLSARTSPAGRHW----FT*FLPSITCIQ----IMISA 106
YLGK + +P+ +SL ++ + + P HW + + PS + I M+SA
Sbjct: 171 YLGKLLPDSAPTHPESLQHVLNDVQEKILPGVTHWQSPNYFAYFPSNSSIAGFLGEMLSA 350
Query: 107 QLKLTSFS 114
L + FS
Sbjct: 351 GLNIVGFS 374
>TC86167 similar to PIR|H96751|H96751 probable casein kinase I F28P22.10
[imported] - Arabidopsis thaliana, partial (74%)
Length = 1967
Score = 27.7 bits (60), Expect = 3.7
Identities = 31/88 (35%), Positives = 41/88 (46%), Gaps = 3/88 (3%)
Frame = -1
Query: 2 KFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGA---DKKLSISLSNEILDYLGK 58
+ LE AP RL D L L I ++ + +GA DK S ++ E L L
Sbjct: 1433 RLLETAPT-RLVDPLGRKKLELDNITS-LDPFESLGTGAFCLDKFPSFAIGPEFL--LRA 1266
Query: 59 SSDNDSPSPVDSLSSPVDSLSARTSPAG 86
SD D P+ + S SP D LSA T+ G
Sbjct: 1265 GSDEDQPACLPS--SPPDGLSALTTAGG 1188
>TC76737 homologue to SP|P06379|RK2_TOBAC Chloroplast 50S ribosomal protein
L2. [Common tobacco] {Nicotiana tabacum}, partial (95%)
Length = 5466
Score = 27.3 bits (59), Expect = 4.8
Identities = 12/41 (29%), Positives = 21/41 (50%)
Frame = +2
Query: 160 PDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLI 200
P + +P+ ++WS N Y++ RIV +S F L+
Sbjct: 2009 P*EKEEPIHRSISLWSKNICVYSQDKARIVIHNISIFKGLL 2131
>TC76659 weakly similar to PIR|T09127|T09127 probable erythrocyte-binding
protein MAEBL - Plasmodium yoelii, partial (2%)
Length = 843
Score = 27.3 bits (59), Expect = 4.8
Identities = 19/60 (31%), Positives = 31/60 (51%)
Frame = +1
Query: 13 SDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSSDNDSPSPVDSLS 72
SD +SNL +G + + ++ + GAD + S+ + D L K D+DS S +S S
Sbjct: 172 SDLISNLGIGPFST---LTLHTPFYGGADDDTIGATSSTVKDELVKEVDSDSESESESES 342
>TC79834 similar to GP|16323504|gb|AAL15246.1 unknown protein {Arabidopsis
thaliana}, partial (49%)
Length = 731
Score = 27.3 bits (59), Expect = 4.8
Identities = 14/33 (42%), Positives = 19/33 (57%)
Frame = -1
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAY 33
+K + A L R+S L NLG + I GC+ AY
Sbjct: 638 VKTADIAKLMRISFSLDLANLGPQGITGCLSAY 540
>TC87898 weakly similar to GP|1174199|gb|AAA86652.1| S25-PR6 {Nicotiana
tabacum}, partial (25%)
Length = 1300
Score = 26.6 bits (57), Expect = 8.2
Identities = 11/21 (52%), Positives = 15/21 (71%)
Frame = -1
Query: 35 CKHSGADKKLSISLSNEILDY 55
C H A ++LS +LSN+I DY
Sbjct: 610 CDHKQAQQRLSSTLSNQIGDY 548
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,317,251
Number of Sequences: 36976
Number of extensions: 105259
Number of successful extensions: 557
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 554
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 556
length of query: 222
length of database: 9,014,727
effective HSP length: 92
effective length of query: 130
effective length of database: 5,612,935
effective search space: 729681550
effective search space used: 729681550
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144731.10


