

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144726.3 - phase: 0
(151 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
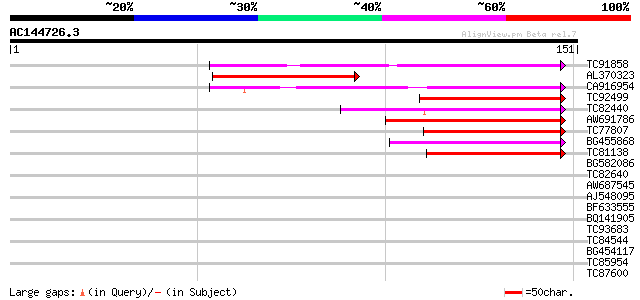
Score E
Sequences producing significant alignments: (bits) Value
TC91858 similar to PIR|T52592|T52592 squamosa-promoter binding p... 72 1e-13
AL370323 similar to GP|19698937|gb| 4-alpha-glucanotransferase {... 71 2e-13
CA916954 similar to PIR|T52592|T525 squamosa-promoter binding pr... 58 1e-09
TC92499 homologue to PIR|T52594|T52594 squamosa promoter binding... 51 2e-07
TC82440 similar to PIR|H96793|H96793 unknown protein F14G6.18 [i... 51 2e-07
AW691786 similar to PIR|T52298|T52 squamosa promoter binding pro... 50 3e-07
TC77807 similar to PIR|T52601|T52601 squamosa promoter binding p... 48 1e-06
BG455868 similar to SP|Q38741|SBP1_ Squamosa-promoter binding pr... 46 5e-06
TC81138 similar to PIR|T52606|T52606 squamosa promoter binding p... 40 5e-04
BG582086 similar to PIR|T52603|T526 squamosa promoter binding pr... 39 0.001
TC82640 similar to GP|10177002|dbj|BAB10252. gene_id:K9E15.9~unk... 30 0.48
AW687545 similar to GP|16904813|gb|A S100Z protein {Homo sapiens... 30 0.48
AJ548095 30 0.48
BF633555 30 0.48
BQ141905 weakly similar to SP|P24613|RK21_ 50S ribosomal protein... 29 0.82
TC93683 weakly similar to GP|7296138|gb|AAF51432.1| BcDNA:GH1214... 28 1.8
TC84544 similar to GP|13591696|gb|AAK31318.1 histone acetyltrans... 27 3.1
BG454117 similar to GP|6437558|gb putative ATPase (ISW2-like) {A... 27 3.1
TC85954 homologue to GP|10719739|gb|AAG17942.1 putative pyridoxi... 27 4.1
TC87600 similar to GP|16226915|gb|AAL16297.1 At1g79440/T8K14_14 ... 27 4.1
>TC91858 similar to PIR|T52592|T52592 squamosa-promoter binding protein 6
[imported] - Arabidopsis thaliana, partial (19%)
Length = 1130
Score = 71.6 bits (174), Expect = 1e-13
Identities = 42/95 (44%), Positives = 57/95 (59%)
Frame = +3
Query: 54 VDLKLGHVGNSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADGC 113
+DLKLG G+ G SK G T ++SS SST ++ AI++ QI GC
Sbjct: 816 IDLKLGRFGDHRDGIGTPFSK---GTTILSSSGSSTPSKRVRSSAIHS--QIAYCQVYGC 980
Query: 114 QADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
DLS+ + YH RHKVC++HSKTS+V + G +QRF
Sbjct: 981 NKDLSSCKDYHKRHKVCEVHSKTSKVIVNGIEQRF 1085
>AL370323 similar to GP|19698937|gb| 4-alpha-glucanotransferase {Arabidopsis
thaliana}, partial (8%)
Length = 477
Score = 70.9 bits (172), Expect = 2e-13
Identities = 35/39 (89%), Positives = 37/39 (94%)
Frame = -3
Query: 55 DLKLGHVGNSSAESGLTKSKDVGGFTKITSSSSSTSGSS 93
DLKLGHVGNSS +SGLTKSKDVGGFTK+ SSSSSTSGSS
Sbjct: 472 DLKLGHVGNSSTKSGLTKSKDVGGFTKMKSSSSSTSGSS 356
>CA916954 similar to PIR|T52592|T525 squamosa-promoter binding protein 6
[imported] - Arabidopsis thaliana, partial (16%)
Length = 775
Score = 58.2 bits (139), Expect = 1e-09
Identities = 36/98 (36%), Positives = 52/98 (52%), Gaps = 3/98 (3%)
Frame = +3
Query: 54 VDLKLGHV---GNSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLA 110
+DLKLG GN+S + +K G ++S SST + ++ Q+
Sbjct: 429 IDLKLGRFDDHGNASIDVAFSK----GAAPILSSCESSTPPKRVRVHSMTAYCQVY---- 584
Query: 111 DGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
GC DLS+ + YH RHKVC++HSKT V + G +QRF
Sbjct: 585 -GCNKDLSSCKDYHKRHKVCEVHSKTPIVIVNGIEQRF 695
>TC92499 homologue to PIR|T52594|T52594 squamosa promoter binding protein 8
[imported] - Arabidopsis thaliana, partial (22%)
Length = 805
Score = 51.2 bits (121), Expect = 2e-07
Identities = 22/39 (56%), Positives = 26/39 (66%)
Frame = +1
Query: 110 ADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
A+GC ADLS + YH RHKVC+ HSK + V G QRF
Sbjct: 28 AEGCNADLSQAKHYHRRHKVCEFHSKAATVVAAGLTQRF 144
>TC82440 similar to PIR|H96793|H96793 unknown protein F14G6.18 [imported] -
Arabidopsis thaliana, partial (12%)
Length = 1244
Score = 50.8 bits (120), Expect = 2e-07
Identities = 25/64 (39%), Positives = 36/64 (56%), Gaps = 4/64 (6%)
Frame = +1
Query: 89 TSGSSKKARAINNGTQIVSSL----ADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGH 144
++G + + I +G+ +S D C+ DLS + YH RHKVC+ HSK S+ LG
Sbjct: 286 STGLVRPNKRIRSGSPTSASYPMCQVDNCKEDLSKAKDYHRRHKVCEAHSKASKALLGNQ 465
Query: 145 KQRF 148
QRF
Sbjct: 466 MQRF 477
>AW691786 similar to PIR|T52298|T52 squamosa promoter binding protein-homolog
4 [imported] - garden snapdragon (fragment), partial
(42%)
Length = 649
Score = 50.4 bits (119), Expect = 3e-07
Identities = 21/48 (43%), Positives = 32/48 (65%)
Frame = +3
Query: 101 NGTQIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
N +Q +GC+ DL++ + Y+ RHKVC +HSK+ VT+ G +QRF
Sbjct: 195 NSSQPPRCQVEGCKLDLTDAKAYYSRHKVCSMHSKSPTVTVSGLQQRF 338
>TC77807 similar to PIR|T52601|T52601 squamosa promoter binding protein 1
[imported] - Arabidopsis thaliana, partial (55%)
Length = 2923
Score = 48.1 bits (113), Expect = 1e-06
Identities = 20/38 (52%), Positives = 26/38 (67%)
Frame = +3
Query: 111 DGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
+ C ADLS + YH RHKVC++HSK S+ +G QRF
Sbjct: 27 EDCGADLSRGKDYHRRHKVCEMHSKASRALVGNAMQRF 140
>BG455868 similar to SP|Q38741|SBP1_ Squamosa-promoter binding protein 1.
[Garden snapdragon] {Antirrhinum majus}, partial (70%)
Length = 656
Score = 46.2 bits (108), Expect = 5e-06
Identities = 22/47 (46%), Positives = 27/47 (56%)
Frame = +1
Query: 102 GTQIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
G+ S D C ADLS + YH RHKVC+ HSK V + +QRF
Sbjct: 247 GSVTPSCQVDNCNADLSAAKQYHKRHKVCENHSKAHSVLISELQQRF 387
>TC81138 similar to PIR|T52606|T52606 squamosa promoter binding protein 7
[imported] - Arabidopsis thaliana (fragment), partial
(32%)
Length = 743
Score = 39.7 bits (91), Expect = 5e-04
Identities = 15/37 (40%), Positives = 23/37 (61%)
Frame = +2
Query: 112 GCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
GC+ D+S +GYH RH+VC + + V L G +R+
Sbjct: 440 GCEVDISELKGYHRRHRVCLRCANAATVVLDGDVKRY 550
>BG582086 similar to PIR|T52603|T526 squamosa promoter binding protein 2
[imported] - Arabidopsis thaliana, partial (18%)
Length = 805
Score = 38.5 bits (88), Expect = 0.001
Identities = 14/38 (36%), Positives = 26/38 (67%)
Frame = +1
Query: 111 DGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
+GC +LS+ + YH + +VC+ H+K+ V + G ++RF
Sbjct: 223 EGCGLNLSSAKDYHCKRRVCESHAKSPMVVIDGLERRF 336
>TC82640 similar to GP|10177002|dbj|BAB10252. gene_id:K9E15.9~unknown
protein {Arabidopsis thaliana}, partial (22%)
Length = 894
Score = 29.6 bits (65), Expect = 0.48
Identities = 8/19 (42%), Positives = 16/19 (84%)
Frame = +1
Query: 27 FWRPWHDSIWNILYGFVCF 45
F+RP HD++W++++G + F
Sbjct: 730 FFRPNHDTVWSLIFGLIFF 786
>AW687545 similar to GP|16904813|gb|A S100Z protein {Homo sapiens}, partial
(16%)
Length = 312
Score = 29.6 bits (65), Expect = 0.48
Identities = 15/40 (37%), Positives = 28/40 (69%)
Frame = +1
Query: 57 KLGHVGNSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKA 96
KL H +++ ++ +++++ V GFTK +S+SS GSSK +
Sbjct: 7 KLLHDLDANKDNEISENEFVDGFTKWINSNSSKPGSSKSS 126
>AJ548095
Length = 593
Score = 29.6 bits (65), Expect = 0.48
Identities = 12/24 (50%), Positives = 16/24 (66%)
Frame = -2
Query: 4 PMLCLFNVCSILYIVLFLNNDEVF 27
PML LFN+C Y ++F +D VF
Sbjct: 160 PMLLLFNICRSCYNLVFFLDDHVF 89
>BF633555
Length = 513
Score = 29.6 bits (65), Expect = 0.48
Identities = 18/33 (54%), Positives = 21/33 (63%), Gaps = 1/33 (3%)
Frame = +3
Query: 51 EFFVDLKLGHVGNSSA-ESGLTKSKDVGGFTKI 82
EF VDLKLG VGNS+ +S L S D +KI
Sbjct: 339 EFSVDLKLGQVGNSATDQSPLPLSNDAVVVSKI 437
>BQ141905 weakly similar to SP|P24613|RK21_ 50S ribosomal protein L21
chloroplast precursor (CL21) (CS-L7). [Spinach]
{Spinacia oleracea}, partial (10%)
Length = 897
Score = 28.9 bits (63), Expect = 0.82
Identities = 18/62 (29%), Positives = 31/62 (49%), Gaps = 3/62 (4%)
Frame = +1
Query: 73 SKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADG---CQADLSNYRGYHMRHKV 129
S D+ F+ S +S SS +AR + Q+ S ++ Q+DL+N+R M +
Sbjct: 673 SYDIDCFSTTDSDHTSHVNSSSRARRYPHSHQLYSDISVASQFAQSDLTNHRSSDMITII 852
Query: 130 CK 131
C+
Sbjct: 853 CR 858
>TC93683 weakly similar to GP|7296138|gb|AAF51432.1| BcDNA:GH12144 gene
product {Drosophila melanogaster}, partial (3%)
Length = 476
Score = 27.7 bits (60), Expect = 1.8
Identities = 25/85 (29%), Positives = 37/85 (43%), Gaps = 6/85 (7%)
Frame = +1
Query: 13 SILYIVLFLNNDEVFWRPWHDSIWNILYGFVCFSMCANEFFVDLKLG------HVGNSSA 66
S + + LFL+ E+ P+H Y VCFS+ + + F L LG + N
Sbjct: 109 SEICVFLFLSLSEI---PFHKIS*LFFYLIVCFSLFSIKLFESLHLGLFLLIIEMDNWRK 279
Query: 67 ESGLTKSKDVGGFTKITSSSSSTSG 91
G++ VG + SSSS G
Sbjct: 280 NQGISNHHQVGHW----KSSSSYKG 342
>TC84544 similar to GP|13591696|gb|AAK31318.1 histone acetyltransferase GCN5
{Arabidopsis thaliana}, partial (30%)
Length = 816
Score = 26.9 bits (58), Expect = 3.1
Identities = 11/37 (29%), Positives = 19/37 (50%)
Frame = -1
Query: 7 CLFNVCSILYIVLFLNNDEVFWRPWHDSIWNILYGFV 43
C++ +C+ V+ L N W P H+ N+L G +
Sbjct: 813 CIWTICNASNNVIRLYNHYRLWAPIHNKPDNVLLGHI 703
>BG454117 similar to GP|6437558|gb putative ATPase (ISW2-like) {Arabidopsis
thaliana}, partial (16%)
Length = 658
Score = 26.9 bits (58), Expect = 3.1
Identities = 21/71 (29%), Positives = 33/71 (45%), Gaps = 3/71 (4%)
Frame = -2
Query: 63 NSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAIN---NGTQIVSSLADGCQADLSN 119
+SS S T + + GGF+ TSSSSS + +A A N + + ++SS C +
Sbjct: 234 SSSPSSPATSASESGGFSPATSSSSSE--LTARATASNSS*SSSSLISSFT--CSSSDKE 67
Query: 120 YRGYHMRHKVC 130
+ G C
Sbjct: 66 FTGSSSEEGCC 34
>TC85954 homologue to GP|10719739|gb|AAG17942.1 putative pyridoxine
biosynthetic enzyme {Phaseolus vulgaris}, partial (94%)
Length = 1373
Score = 26.6 bits (57), Expect = 4.1
Identities = 15/40 (37%), Positives = 22/40 (54%)
Frame = -1
Query: 51 EFFVDLKLGHVGNSSAESGLTKSKDVGGFTKITSSSSSTS 90
+FF +L++GH N++ S DVGG T +STS
Sbjct: 335 DFFDELRVGHTSNTAL------STDVGGDTLQGHDGASTS 234
>TC87600 similar to GP|16226915|gb|AAL16297.1 At1g79440/T8K14_14 {Arabidopsis
thaliana}, partial (93%)
Length = 2016
Score = 26.6 bits (57), Expect = 4.1
Identities = 16/33 (48%), Positives = 20/33 (60%), Gaps = 2/33 (6%)
Frame = -1
Query: 16 YIVLFLNNDEVFWR-PWHDS-IWNILYGFVCFS 46
Y +LFLN V W P H+S I N+LY F C +
Sbjct: 1974 YKILFLNQANVLWEXPHHNSFIXNLLY-FECLT 1879
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.137 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,281,381
Number of Sequences: 36976
Number of extensions: 102300
Number of successful extensions: 978
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 965
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 975
length of query: 151
length of database: 9,014,727
effective HSP length: 88
effective length of query: 63
effective length of database: 5,760,839
effective search space: 362932857
effective search space used: 362932857
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 54 (25.4 bits)
Medicago: description of AC144726.3


