

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144726.11 + phase: 0
(515 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
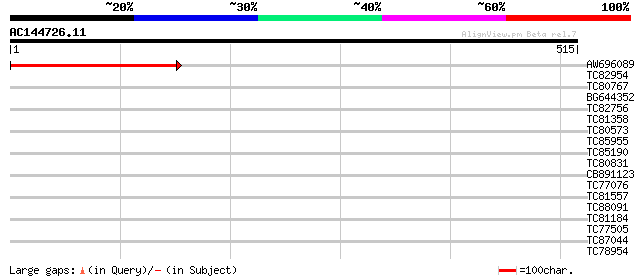
Score E
Sequences producing significant alignments: (bits) Value
AW696089 homologue to GP|1865721|emb| mitochondrial single-subun... 307 6e-84
TC82954 weakly similar to GP|8809591|dbj|BAA97142.1 emb|CAB82814... 33 0.26
TC80767 weakly similar to GP|21593751|gb|AAM65718.1 unknown {Ara... 30 2.9
BG644352 homologue to GP|12231300|gb ripening regulated protein ... 29 3.8
TC82756 similar to GP|17381278|gb|AAL36057.1 AT4g34410/F10M10_18... 29 3.8
TC81358 similar to GP|17473570|gb|AAL38260.1 putative protein {A... 29 3.8
TC80573 similar to GP|9759187|dbj|BAB09724.1 importin beta {Arab... 29 3.8
TC85955 similar to GP|3860319|emb|CAA10127.1 nucleolar protein {... 29 5.0
TC85190 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 28 6.5
TC80831 similar to GP|18542939|gb|AAK00428.2 Unknown protein {Or... 28 6.5
CB891123 GP|4062407|db Heme-binding protein a precursor (hemin-b... 28 6.5
TC77076 similar to PIR|T07734|T07734 homeotic protein VAHOX1 - t... 28 6.5
TC81557 28 8.5
TC88091 homologue to SP|P51819|HS83_PHANI Heat shock protein 83.... 28 8.5
TC81184 similar to GP|13487783|gb|AAK27718.1 ADP-glucose pyropho... 28 8.5
TC77505 MtN29 28 8.5
TC87044 similar to GP|21554145|gb|AAM63225.1 unknown {Arabidopsi... 28 8.5
TC78954 weakly similar to PIR|D96810|D96810 hypothetical protein... 28 8.5
>AW696089 homologue to GP|1865721|emb| mitochondrial single-subunit
DNA-dependent RNA polymerase {Chenopodium album},
partial (2%)
Length = 615
Score = 307 bits (787), Expect = 6e-84
Identities = 152/156 (97%), Positives = 155/156 (98%)
Frame = +3
Query: 1 MWSKLAKRASSRRFILPSQSSTSLRFSHQHNFPEKILRHPFSRFSQVGIFTSEYKLGRSN 60
MWSKLAKRASSRRFILPSQSSTSLRFSHQHNFPEKILRHPFSRFSQVGIFTSEYKLGRSN
Sbjct: 135 MWSKLAKRASSRRFILPSQSSTSLRFSHQHNFPEKILRHPFSRFSQVGIFTSEYKLGRSN 314
Query: 61 FSFSEHPFSFSSISLNHFRTYGSVAAEAIESTDTEDDCSGSEEVVQKLLDQMVIEEEKKN 120
FSFSEHPFSFSSISLNHFRTYGSVAAEAIESTDTEDDCSGSEEVVQKLLDQMVIEEEKKN
Sbjct: 315 FSFSEHPFSFSSISLNHFRTYGSVAAEAIESTDTEDDCSGSEEVVQKLLDQMVIEEEKKN 494
Query: 121 PQLEEKKKNKNNSYKYKMLRRRQIKIETEAWEEAAR 156
PQLEEKKKNKNNSYKYKMLRRRQIKIETEAWE +++
Sbjct: 495 PQLEEKKKNKNNSYKYKMLRRRQIKIETEAWERSSK 602
>TC82954 weakly similar to GP|8809591|dbj|BAA97142.1
emb|CAB82814.1~gene_id:MNB8.8~similar to unknown protein
{Arabidopsis thaliana}, partial (7%)
Length = 783
Score = 33.1 bits (74), Expect = 0.26
Identities = 40/139 (28%), Positives = 59/139 (41%), Gaps = 2/139 (1%)
Frame = +3
Query: 35 KILRHPFSRFSQ--VGIFTSEYKLGRSNFSFSEHPFSFSSISLNHFRTYGSVAAEAIEST 92
K +RH ++ ++ VGIF ++ + S FS LN R + AI +
Sbjct: 324 KKIRHEDAKANEKVVGIFAAQEQ---SWFSERRKLRQQIGALLNELRVFEKKRDLAI--S 488
Query: 93 DTEDDCSGSEEVVQKLLDQMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIETEAWE 152
D E +V++ D+ + EEEKK +LEEK K K E E
Sbjct: 489 DLNQKLKEMEGLVEEK-DKKIEEEEKKRKELEEKAKKAE-------------KDAEELRE 626
Query: 153 EAAREYQELLEDMREQKLA 171
+ RE QE D+R+ K A
Sbjct: 627 SSKREGQEHSSDLRKHKTA 683
>TC80767 weakly similar to GP|21593751|gb|AAM65718.1 unknown {Arabidopsis
thaliana}, partial (35%)
Length = 750
Score = 29.6 bits (65), Expect = 2.9
Identities = 47/166 (28%), Positives = 71/166 (42%), Gaps = 8/166 (4%)
Frame = +3
Query: 7 KRASSRRFILPSQSSTSLRFSHQHNFPE-KILRHPFSRFSQVGIFTSEYKLGRSNFSFSE 65
K+ S FI S+S + L S E K +PF++ ++ I + KL R +F
Sbjct: 135 KKGLSNHFIGKSKSFSDL--SQVTTVTELKKQENPFNKRRRLLIAS---KLSRKSFYSCF 299
Query: 66 HPFSFSSISLNHFRTYGSVAAEAIESTDTEDDCSGSEEVVQKLLDQMVIE--EEKKNPQL 123
+P S +S+N D ++D G +EV K EEKKNPQ
Sbjct: 300 NPKSMPLLSVNE---------------DDDEDQGGEKEVTTKGSPSSSSSSMEEKKNPQE 434
Query: 124 EEKKKNKNN----SYKYKMLRRRQIKIETEAWEEA-AREYQELLED 164
E + NN SY M R R ++ ++ A +E+ E+ ED
Sbjct: 435 EVMMRQFNNRMPQSYANHM-RLRLGSFKSRSFSLADLQEHDEVEED 569
>BG644352 homologue to GP|12231300|gb ripening regulated protein DDTFR10
{Lycopersicon esculentum}, partial (98%)
Length = 726
Score = 29.3 bits (64), Expect = 3.8
Identities = 27/113 (23%), Positives = 45/113 (38%)
Frame = +3
Query: 231 LLMTNSNGVGSARVIQAACQIGEAIEHEGRIYRFMEKTKVKKATSHKSDSDSVPAPVKGE 290
L ++ +G G+ +++ + I EA+ KA++ + + D + GE
Sbjct: 294 LRISGVSGEGAGVIVEGSAPITEAVA--------TPPAADSKASAAEEEDDDDDVDLFGE 449
Query: 291 NLTAEENEKLAKEEKRLRNKVASLIKKQKKQQALGIVKGRDQAKPWGQEAQVK 343
E EK A EE+ K + K+ K L VK PW E +K
Sbjct: 450 ET---EEEKKAAEERAAAMKASGKKKESGKSSVLMDVK------PWDDETDMK 581
>TC82756 similar to GP|17381278|gb|AAL36057.1 AT4g34410/F10M10_180
{Arabidopsis thaliana}, partial (35%)
Length = 630
Score = 29.3 bits (64), Expect = 3.8
Identities = 22/74 (29%), Positives = 33/74 (43%), Gaps = 15/74 (20%)
Frame = +3
Query: 116 EEKKNPQLEEKKKNKNNSYKYKMLR------------RRQIKIETEAW---EEAAREYQE 160
EEKK Q +++K+ KNN Y+ R RR +++ + EEAAR Y
Sbjct: 210 EEKKQKQKQKQKRAKNNKYRGVRQRPWGKWAAEIRDPRRAVRVWLGTFTTAEEAARAYDN 389
Query: 161 LLEDMREQKLAPNL 174
+ R + NL
Sbjct: 390 AAIEFRGPRAKLNL 431
>TC81358 similar to GP|17473570|gb|AAL38260.1 putative protein
{Arabidopsis thaliana}, partial (29%)
Length = 722
Score = 29.3 bits (64), Expect = 3.8
Identities = 27/86 (31%), Positives = 39/86 (44%), Gaps = 9/86 (10%)
Frame = +2
Query: 16 LPSQSSTSLRFSHQHNFPEKILRHPFSRFSQVGIFTSEYKLGRSNFSFS--------EHP 67
+PS SS+SL FSH+ P+ I + P + + S L ++F FS H
Sbjct: 17 IPSCSSSSLHFSHK---PKSIKKIPIFFKTLTLMELSHLSLSENHFFFSTPTSPSSLSHR 187
Query: 68 F-SFSSISLNHFRTYGSVAAEAIEST 92
SF S+S NH R S + + T
Sbjct: 188LPSFLSLSTNHRRRRRSFCSASNSDT 265
>TC80573 similar to GP|9759187|dbj|BAB09724.1 importin beta {Arabidopsis
thaliana}, partial (27%)
Length = 1220
Score = 29.3 bits (64), Expect = 3.8
Identities = 19/76 (25%), Positives = 34/76 (44%), Gaps = 2/76 (2%)
Frame = -1
Query: 157 EYQELLEDMREQKLAPNLPYMKSLFLGWFEPLKNAIAA--DQEICKDTKTRLSHAPYFNE 214
++ E L + + A N+P++K L K + A IC +++ YF++
Sbjct: 449 DFDEFLNPWKIPEYASNIPFLKELMYSVISKSKPDVWAYISAAICS---VFIAYIKYFSK 279
Query: 215 LPPDMMAVITMHKLMG 230
L P A+ MH+ G
Sbjct: 278 LSPIASAISPMHENKG 231
>TC85955 similar to GP|3860319|emb|CAA10127.1 nucleolar protein {Cicer
arietinum}, partial (98%)
Length = 1981
Score = 28.9 bits (63), Expect = 5.0
Identities = 23/84 (27%), Positives = 36/84 (42%), Gaps = 12/84 (14%)
Frame = +2
Query: 84 VAAEAIESTDTEDDCSGSEEVVQKLL----------DQMVIEE--EKKNPQLEEKKKNKN 131
V AIES D +D +EEV K D M +++ E N E+ K K
Sbjct: 1352 VMKSAIESVDNKDTEMETEEVSAKKTKKKKQKAADGDDMAVDKAAEITNGDAEDHKSEKK 1531
Query: 132 NSYKYKMLRRRQIKIETEAWEEAA 155
K K +++++E + E+ A
Sbjct: 1532 KKKKEKRKLDQEVEVEDKVVEDGA 1603
>TC85190 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (50%)
Length = 1496
Score = 28.5 bits (62), Expect = 6.5
Identities = 20/85 (23%), Positives = 38/85 (44%), Gaps = 9/85 (10%)
Frame = +3
Query: 90 ESTDTEDDCSGSEEVVQKLLDQMVIEEEK---------KNPQLEEKKKNKNNSYKYKMLR 140
E E+D +EEVV++ D+ V+EE+K + P++E + K + +
Sbjct: 555 EEKKPEEDEKKTEEVVEEKKDEAVVEEKKVDEEKGSTSEEPKVETAEPEKEEKKVEETVV 734
Query: 141 RRQIKIETEAWEEAAREYQELLEDM 165
KI E+ A+ + + E +
Sbjct: 735 EVVEKIAASTEEDGAKTVEAIQESI 809
>TC80831 similar to GP|18542939|gb|AAK00428.2 Unknown protein {Oryza
sativa}, partial (7%)
Length = 1039
Score = 28.5 bits (62), Expect = 6.5
Identities = 17/64 (26%), Positives = 39/64 (60%), Gaps = 3/64 (4%)
Frame = +2
Query: 111 QMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQ-IKIETEAWEEAAREYQELL--EDMRE 167
+++ EEEK+ + EE+K+ + + K LRR++ +K + + E+ E ++L ++ +
Sbjct: 125 KLLEEEEKEKREEEERKERRRTKEREKKLRRKERLKGKEKDREKICSESNDILCTSEISK 304
Query: 168 QKLA 171
++LA
Sbjct: 305 EELA 316
>CB891123 GP|4062407|db Heme-binding protein a precursor (hemin-binding
lipoprotein). {Escherichia coli}, partial (33%)
Length = 733
Score = 28.5 bits (62), Expect = 6.5
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = -1
Query: 17 PSQSSTSLRFSHQHNFPEKILRHPFSRFSQVGI 49
P+ ST+L SH H+ +K+L+ + +QVGI
Sbjct: 196 PNGFSTTLWSSHNHSTAQKVLQFTQQQLAQVGI 98
>TC77076 similar to PIR|T07734|T07734 homeotic protein VAHOX1 - tomato,
partial (43%)
Length = 1664
Score = 28.5 bits (62), Expect = 6.5
Identities = 15/64 (23%), Positives = 33/64 (51%)
Frame = +1
Query: 123 LEEKKKNKNNSYKYKMLRRRQIKIETEAWEEAAREYQELLEDMREQKLAPNLPYMKSLFL 182
L+E +++ + Y ++ ++R++ ++ + E + E LE R+ KLA +L
Sbjct: 589 LDENGEDEMDEYFHQSEKKRRLSVDQVQFLEKSFEEDNKLEPERKTKLAKDLGLQPRQVA 768
Query: 183 GWFE 186
WF+
Sbjct: 769 IWFQ 780
>TC81557
Length = 626
Score = 28.1 bits (61), Expect = 8.5
Identities = 16/52 (30%), Positives = 26/52 (49%)
Frame = +2
Query: 109 LDQMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIETEAWEEAAREYQE 160
LD + ++KN L K KN ++ YK+ + I ETE +E + +E
Sbjct: 176 LDFNELVPKEKNTMLVAKTKNIKHTNNYKLKKLEDIPEETELEDETHPQLEE 331
>TC88091 homologue to SP|P51819|HS83_PHANI Heat shock protein 83. [Violet
Japanese morning glory] {Pharbitis nil}, partial (77%)
Length = 1813
Score = 28.1 bits (61), Expect = 8.5
Identities = 21/80 (26%), Positives = 41/80 (51%), Gaps = 6/80 (7%)
Frame = +1
Query: 94 TEDDCSGSEEVVQKLLDQMVIEEEKKNPQLEEKKKN--KNNSYKYKMLRRRQ----IKIE 147
TE + S E+ K ++ V+EE ++ + + KKK K S++++++ +++ K E
Sbjct: 181 TEKEISDDEDDEPKKEEEGVVEEVDEDKEKDAKKKKKIKEVSHEWELINKQKPIWLRKPE 360
Query: 148 TEAWEEAAREYQELLEDMRE 167
+E A Y+ L D E
Sbjct: 361 EITKDEYAAFYKSLTNDWEE 420
>TC81184 similar to GP|13487783|gb|AAK27718.1 ADP-glucose pyrophosphorylase
{Cicer arietinum}, partial (29%)
Length = 683
Score = 28.1 bits (61), Expect = 8.5
Identities = 12/28 (42%), Positives = 19/28 (67%)
Frame = +3
Query: 264 FMEKTKVKKATSHKSDSDSVPAPVKGEN 291
F +T+ K++ + S+S SVP PV G+N
Sbjct: 102 FQTRTRRKRSFINNSNSHSVPPPVTGKN 185
>TC77505 MtN29
Length = 889
Score = 28.1 bits (61), Expect = 8.5
Identities = 14/54 (25%), Positives = 33/54 (60%), Gaps = 1/54 (1%)
Frame = +3
Query: 132 NSYKYKMLRRRQIKIETEAWEEAAREYQ-ELLEDMREQKLAPNLPYMKSLFLGW 184
+S ++ + R+ +K++T + +E + + +E M+E+ L + Y+K L++GW
Sbjct: 66 SSLLWQSIHRKLVKLKTPKTKLG*KELRMDTMETMKEEML---MDYLKILYIGW 218
>TC87044 similar to GP|21554145|gb|AAM63225.1 unknown {Arabidopsis
thaliana}, complete
Length = 1403
Score = 28.1 bits (61), Expect = 8.5
Identities = 22/59 (37%), Positives = 29/59 (48%), Gaps = 2/59 (3%)
Frame = -2
Query: 16 LPSQSSTSLRFSHQHN--FPEKILRHPFSRFSQVGIFTSEYKLGRSNFSFSEHPFSFSS 72
+PS TS + + FP KI P +RFS +S L R+ FS E F+FSS
Sbjct: 625 IPSLEKTSASCAEPRSNMFPAKI---PLTRFSTRLTSSSMASLTRTTFSNREMAFAFSS 458
>TC78954 weakly similar to PIR|D96810|D96810 hypothetical protein T11I11.6
[imported] - Arabidopsis thaliana, partial (45%)
Length = 2329
Score = 28.1 bits (61), Expect = 8.5
Identities = 11/32 (34%), Positives = 18/32 (55%)
Frame = +1
Query: 133 SYKYKMLRRRQIKIETEAWEEAAREYQELLED 164
+Y MLRR + E WE A ++Y+ L+ +
Sbjct: 1702 NYSKAMLRRADCNAKLERWEAAIQDYEMLIRE 1797
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,759,592
Number of Sequences: 36976
Number of extensions: 195037
Number of successful extensions: 1253
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 1234
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1250
length of query: 515
length of database: 9,014,727
effective HSP length: 100
effective length of query: 415
effective length of database: 5,317,127
effective search space: 2206607705
effective search space used: 2206607705
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144726.11


