

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144724.5 - phase: 0 /pseudo
(534 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
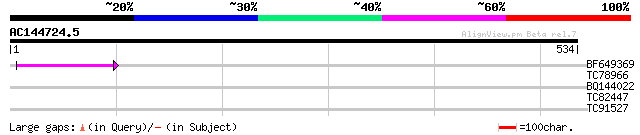
Score E
Sequences producing significant alignments: (bits) Value
BF649369 55 7e-08
TC78966 weakly similar to GP|16649051|gb|AAL24377.1 Unknown prot... 30 1.8
BQ144022 homologue to GP|22831315|dbj hypothetical protein~predi... 30 2.3
TC82447 similar to GP|14589311|emb|CAC43238. calcium binding pro... 29 5.2
TC91527 weakly similar to PIR|T45786|T45786 receptor-protein kin... 28 8.8
>BF649369
Length = 631
Score = 55.1 bits (131), Expect = 7e-08
Identities = 39/96 (40%), Positives = 54/96 (55%)
Frame = +1
Query: 7 WKVQPYTGLIS*WKQRTICPGKSSRKG*LRAMEGGDWKILSKNWQH*NRVEMLKNLWRPL 66
WKV G *WKQRTI PG++ R+ * DW+I SKN+QH ++E LK+L +
Sbjct: 244 WKVLRSIGSTY*WKQRTIYPGRN*RRR*SLVTTVEDWRIHSKNYQHFVKLEALKSLSKLS 423
Query: 67 NFCPRR*DVFLKTST*GTL*VD*SHRFVGGFVL*IL 102
+ RR + + +++ T *VD* FV *IL
Sbjct: 424 SCYRRRLEGYRRSNIWVTS*VD*KLTSEDEFVR*IL 531
>TC78966 weakly similar to GP|16649051|gb|AAL24377.1 Unknown protein
{Arabidopsis thaliana}, partial (33%)
Length = 1022
Score = 30.4 bits (67), Expect = 1.8
Identities = 20/46 (43%), Positives = 25/46 (53%), Gaps = 4/46 (8%)
Frame = +1
Query: 393 CNSCMKEKW-*RCWGKVASRRNTV--ISTHFWGTNK-VGWEWIGGG 434
CN C KE++ * C V + R IS + NK VGW W+GGG
Sbjct: 496 CNVCDKEEYD*CCCNCVKATRECT*NISINEKAYNKEVGWSWLGGG 633
>BQ144022 homologue to GP|22831315|dbj hypothetical protein~predicted by
GlimmerM etc. {Oryza sativa (japonica cultivar-group)},
partial (16%)
Length = 1286
Score = 30.0 bits (66), Expect = 2.3
Identities = 20/60 (33%), Positives = 27/60 (44%)
Frame = -3
Query: 368 VWTWC*ACRG*IPWVRY*WIGRLSLCNSCMKEKW*RCWGKVASRRNTVISTHFWGTNKVG 427
VW C C + VR W+G L LC +EK+ W K + V+S WG +G
Sbjct: 180 VWVGCPRCFL-LRAVRLSWVGLLVLCFLFSEEKFCGVWPK----KRWVVSVWGWGAGALG 16
>TC82447 similar to GP|14589311|emb|CAC43238. calcium binding protein
{Sesbania rostrata}, partial (79%)
Length = 660
Score = 28.9 bits (63), Expect = 5.2
Identities = 10/23 (43%), Positives = 15/23 (64%)
Frame = -3
Query: 399 EKW*RCWGKVASRRNTVISTHFW 421
E+W*RCW +V N +S H++
Sbjct: 166 EEW*RCWSRVGFNWNGFVSCHWF 98
>TC91527 weakly similar to PIR|T45786|T45786 receptor-protein kinase-like
protein - Arabidopsis thaliana, partial (17%)
Length = 1205
Score = 28.1 bits (61), Expect = 8.8
Identities = 13/45 (28%), Positives = 30/45 (65%), Gaps = 1/45 (2%)
Frame = +2
Query: 177 LYVITSFYWKKKERTRQ-DLRSGKGSATSTTTRWRKEGPRAYASS 220
L ++ F +++K +T + + ++ K S +STT++W GP ++A++
Sbjct: 629 LSIVVFFLYRRKRQTEEKNSKTTKDSKSSTTSKW---GPLSFATT 754
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.357 0.157 0.630
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,995,007
Number of Sequences: 36976
Number of extensions: 306510
Number of successful extensions: 3139
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1650
Number of HSP's successfully gapped in prelim test: 106
Number of HSP's that attempted gapping in prelim test: 1447
Number of HSP's gapped (non-prelim): 1837
length of query: 534
length of database: 9,014,727
effective HSP length: 101
effective length of query: 433
effective length of database: 5,280,151
effective search space: 2286305383
effective search space used: 2286305383
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144724.5


