

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144721.3 - phase: 0 /pseudo
(750 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
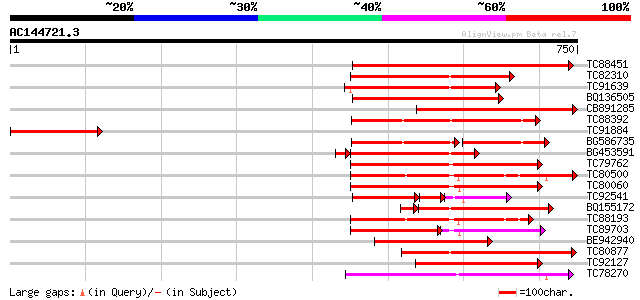
Score E
Sequences producing significant alignments: (bits) Value
TC88451 similar to GP|836954|gb|AAC23542.1|| receptor protein ki... 319 3e-87
TC82310 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1... 291 8e-79
TC91639 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1... 279 2e-75
BQ136505 similar to PIR|T05754|T05 S-receptor kinase (EC 2.7.1.-... 267 1e-71
CB891285 weakly similar to PIR|T14470|T14 receptor-like kinase (... 265 4e-71
TC88392 similar to GP|11275527|dbj|BAB18292. putative receptor-l... 254 7e-68
TC91884 similar to GP|4741219|emb|CAB41879.1 SRK15 protein {Bras... 249 2e-66
BG586735 similar to GP|11275527|db putative receptor-like protei... 171 7e-65
BG453591 similar to PIR|T05181|T05 S-receptor kinase (EC 2.7.1.-... 238 3e-63
TC79762 similar to PIR|H96731|H96731 hypothetical protein F5A18.... 236 2e-62
TC80500 similar to PIR|G96602|G96602 probable receptor protein k... 232 5e-61
TC80060 similar to PIR|D96574|D96574 hypothetical protein T3F20.... 232 5e-61
TC92541 similar to PIR|T02153|T02153 protein kinase homolog T1F1... 123 7e-60
BQ155172 weakly similar to GP|6686398|gb F1E22.15 {Arabidopsis t... 210 4e-56
TC88193 similar to PIR|G96602|G96602 probable receptor protein k... 213 2e-55
TC89703 similar to GP|11994595|dbj|BAB02650. receptor-like serin... 133 6e-55
BE942940 weakly similar to GP|13506749|gb receptor-like protein ... 208 5e-54
TC80877 weakly similar to GP|5814093|gb|AAD52097.1| receptor-lik... 208 5e-54
TC92127 weakly similar to GP|16040952|dbj|BAB69683. receptor kin... 203 2e-52
TC78270 similar to GP|16649103|gb|AAL24403.1 Unknown protein {Ar... 203 2e-52
>TC88451 similar to GP|836954|gb|AAC23542.1|| receptor protein kinase
{Ipomoea trifida}, partial (28%)
Length = 1276
Score = 319 bits (817), Expect = 3e-87
Identities = 159/293 (54%), Positives = 214/293 (72%)
Frame = +3
Query: 454 GQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSL 513
GQEIAVKRLSK SGQGI EF NE+++I +LQH+NLV+LLGCC+ E++L+YE+MPNKSL
Sbjct: 294 GQEIAVKRLSKTSGQGIVEFKNELLLICELQHKNLVQLLGCCIHEEERILIYEYMPNKSL 473
Query: 514 DAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKI 573
D +LFD +KK LDW+KR NI+EGIA+G++YLH+ SRLKIIHRDLKASNILLD+ M PKI
Sbjct: 474 DFYLFDCTKKKLLDWKKRFNIIEGIAQGLLYLHKYSRLKIIHRDLKASNILLDENMNPKI 653
Query: 574 SDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRR 633
+DFG+AR+ E NT R+VGTYGYM PEYAM G+ S KSDVYSFGVLLLEIV G +
Sbjct: 654 ADFGMARMFTQQE-SVVNTNRIVGTYGYMSPEYAMEGVCSTKSDVYSFGVLLLEIVCGIK 830
Query: 634 NNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKE 693
NNSFY + L+L+G AW+LW + + L+D + D + RC+H+GLLCV++ +
Sbjct: 831 NNSFYDVDRPLNLIGHAWELWNDGEYLKLMDPTLNDTFVPDEVKRCIHVGLLCVEQYAND 1010
Query: 694 RPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVTLS 746
RP++S V+ +L ++ P K AF + E++ + +++ + T+S
Sbjct: 1011RPTMSEVISVLTNKYVLTNLPRKPAFYVRREIFEGETTSKGQDTDTYSTTTIS 1169
>TC82310 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1.-)
T6K22.120 precursor - Arabidopsis thaliana, partial
(24%)
Length = 696
Score = 291 bits (744), Expect = 8e-79
Identities = 141/217 (64%), Positives = 173/217 (78%)
Frame = +1
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ D +EIAVKRLSK S QG+EE NE+++I+KLQHRNLVRLL CC+E+ EK+L+YE++PN
Sbjct: 37 LADEKEIAVKRLSKTSSQGVEELKNEIILIAKLQHRNLVRLLACCIEQNEKLLIYEYLPN 216
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
SLD LFD ++ +L WR+R NI+ GIA+G++YLH DSRL++IHRDLKASNILLD EM
Sbjct: 217 SSLDFHLFDMVKGAQLAWRQRLNIINGIAKGLLYLHEDSRLRVIHRDLKASNILLDQEMN 396
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFGLAR GG+ DEANT RVVGTYGYM PEYAM GLFS KSDV+SFGVLLLEI+S
Sbjct: 397 PKISDFGLARTF-GGDQDEANTIRVVGTYGYMAPEYAMEGLFSVKSDVFSFGVLLLEIIS 573
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREV 667
GR+N+ FY +E SL FAW LW + L+D +
Sbjct: 574 GRKNSKFYLSEHGQSLPIFAWNLWCKRKGFELMDPSI 684
>TC91639 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1.-)
T6K22.120 precursor - Arabidopsis thaliana, partial
(28%)
Length = 1313
Score = 279 bits (714), Expect = 2e-75
Identities = 144/210 (68%), Positives = 171/210 (80%), Gaps = 4/210 (1%)
Frame = +3
Query: 444 QGRFWP----GIQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERG 499
QG F P + G+EIAVKRLS+ SGQG++EF NE+ + ++LQHRNLV+L+GC +E
Sbjct: 432 QGGFGPVYKGKLPSGEEIAVKRLSRRSGQGLDEFKNEMRLFAQLQHRNLVKLMGCSIEGD 611
Query: 500 EKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLK 559
EK+LVYEFM NKSLD FLFDPI+K +LDW +R I+EGIARG++YLHRDSRL+IIHRDLK
Sbjct: 612 EKLLVYEFMLNKSLDRFLFDPIKKTQLDWARRYEIIEGIARGLLYLHRDSRLRIIHRDLK 791
Query: 560 ASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVY 619
ASNILLD+ M PKISDFGLARI GG +E N +VVGTYGYM PEYAM GL S KSDVY
Sbjct: 792 ASNILLDENMNPKISDFGLARIF-GGNQNEENATKVVGTYGYMSPEYAMEGLVSVKSDVY 968
Query: 620 SFGVLLLEIVSGRRNNSFYQNEDSLSLVGF 649
SFGVLLLEIVSGRRN SF ++DS SL+G+
Sbjct: 969 SFGVLLLEIVSGRRNTSFRHSDDS-SLIGY 1055
>BQ136505 similar to PIR|T05754|T05 S-receptor kinase (EC 2.7.1.-) M4I22.110
precursor - Arabidopsis thaliana, partial (23%)
Length = 618
Score = 267 bits (683), Expect = 1e-71
Identities = 128/200 (64%), Positives = 165/200 (82%)
Frame = +2
Query: 454 GQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSL 513
G+E+A+KR SK S QG+EEF NEV++I+KLQHRNLV+LLGCC+ R EK+L+YE+MPN+SL
Sbjct: 20 GKEVAIKRNSKMSDQGLEEFKNEVLLIAKLQHRNLVKLLGCCIHREEKLLIYEYMPNRSL 199
Query: 514 DAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKI 573
D F+FD + K LDW KRS+I+ G+ARG++YLH+DSRL+IIHRDLK SNILLD M PKI
Sbjct: 200 DYFIFDETRSKLLDWSKRSHIIAGVARGLLYLHQDSRLRIIHRDLKLSNILLDALMNPKI 379
Query: 574 SDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRR 633
SDFGLAR G + EA T+++VGTYGYMPPEYA+ G +S KSDV+SFGV++LEI+SG++
Sbjct: 380 SDFGLARTFCGDQ-VEAKTRKLVGTYGYMPPEYAVHGRYSMKSDVFSFGVIVLEIISGKK 556
Query: 634 NNSFYQNEDSLSLVGFAWKL 653
FY SL+L+G AW+L
Sbjct: 557 IKVFYDPXHSLNLLGHAWRL 616
>CB891285 weakly similar to PIR|T14470|T14 receptor-like kinase (EC 2.7.1.-)
SFR2 - wild cabbage, partial (18%)
Length = 753
Score = 265 bits (678), Expect = 4e-71
Identities = 131/212 (61%), Positives = 167/212 (77%)
Frame = +2
Query: 539 ARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGT 598
ARG++YLHRDS+L+IIHRDLK SNILLD+E+ PKISDFG+ARI +G E D NT RVVGT
Sbjct: 2 ARGLLYLHRDSKLRIIHRDLKTSNILLDEELNPKISDFGMARIFEGRE-DTENTIRVVGT 178
Query: 599 YGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEEN 658
YGY+ PEYAM GLFSEKSDV+SFGVLLLEI+SGRRN+SFY NE +L+L+GF W W E N
Sbjct: 179 YGYISPEYAMQGLFSEKSDVFSFGVLLLEIISGRRNSSFYDNEHALTLLGFVWIQWKEGN 358
Query: 659 TISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVA 718
+S I+ E++D S ++RC+HIGLLCVQEL +RP+++ V+ ML SE LPPP + A
Sbjct: 359 ILSFINTEIYDPSHHKYVVRCIHIGLLCVQELAVDRPNMAAVISMLNSEAELLPPPSQPA 538
Query: 719 FVHNQNSRSTESSQQSHRSNSNNNVTLSDVIG 750
F+ QN ST S ++SHR S N+V+++D+ G
Sbjct: 539 FILRQNMLSTMSHEESHRRYSINSVSITDISG 634
>TC88392 similar to GP|11275527|dbj|BAB18292. putative receptor-like protein
kinase {Oryza sativa (japonica cultivar-group)}, partial
(61%)
Length = 1017
Score = 254 bits (650), Expect = 7e-68
Identities = 129/250 (51%), Positives = 179/250 (71%)
Frame = +1
Query: 453 DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKS 512
+G+E+A+K+LS S QGI EF NEV ++ ++QH+NLV LLGCC E EKMLVYE++PNKS
Sbjct: 280 NGEEVAIKKLSMESRQGIREFTNEVRLLLRIQHKNLVTLLGCCAEGPEKMLVYEYLPNKS 459
Query: 513 LDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
LD FLFD +K+ LDW R IV GIARG++YLH ++ +IIHRD+KASNILLD+++ PK
Sbjct: 460 LDHFLFD--KKRSLDWMTRFRIVTGIARGLLYLHEEAPERIIHRDIKASNILLDEKLNPK 633
Query: 573 ISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGR 632
ISDFGLAR+ GE T R+ T+GYM PEYA+ G S K+DV+S+GVL+LEIVSGR
Sbjct: 634 ISDFGLARLFP-GEDTHVQTFRISRTHGYMAPEYALRGYLSVKTDVFSYGVLVLEIVSGR 810
Query: 633 RNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPK 692
+N+ + + L+ +AWKL+ + LID+ + + + + + C+ +GLLC Q
Sbjct: 811 KNHDLKLDAEKADLLSYAWKLYQGRKIMDLIDQNIGKYNGDEAAM-CIQLGLLCCQASLV 987
Query: 693 ERPSISTVVL 702
ERP +++V L
Sbjct: 988 ERPDMNSVNL 1017
>TC91884 similar to GP|4741219|emb|CAB41879.1 SRK15 protein {Brassica
oleracea}, partial (3%)
Length = 1156
Score = 249 bits (637), Expect = 2e-66
Identities = 117/123 (95%), Positives = 118/123 (95%)
Frame = +2
Query: 1 MDCKQRSTTQRFKWNCHNSQRWKSCNIEQTKWYHHLVNKHFIFNKFNS*T**CRELDSKR 60
MDCKQRSTTQRFKWNCHNSQ+WKSCN EQTKW HHLVNKHFIFNKFNS*T**CRELDSKR
Sbjct: 773 MDCKQRSTTQRFKWNCHNSQKWKSCNFEQTKWQHHLVNKHFIFNKFNS*T**CRELDSKR 952
Query: 61 HQFWSNNLG*FHTPCRFSCSIDENCI**SYR*ANCFCSKKK*Q*SFFGTFYN*C*KIGCT 120
HQFWSNNLG*FHTP RFSCSIDENCI* SYR*ANCFCSKKK*Q*SFFGTFYN*C*KIGCT
Sbjct: 953 HQFWSNNLG*FHTPFRFSCSIDENCI*QSYR*ANCFCSKKK*Q*SFFGTFYN*C*KIGCT 1132
Query: 121 RSF 123
R F
Sbjct: 1133RKF 1141
>BG586735 similar to GP|11275527|db putative receptor-like protein kinase
{Oryza sativa (japonica cultivar-group)}, partial (57%)
Length = 794
Score = 171 bits (432), Expect(2) = 7e-65
Identities = 85/143 (59%), Positives = 110/143 (76%)
Frame = +3
Query: 453 DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKS 512
+G+E+A+K+LS S QGI EF NEV ++ ++QH+NLV LLGCC E EKMLVYE++PNKS
Sbjct: 21 NGEEVAIKKLSMESRQGIREFTNEVRLLLRIQHKNLVTLLGCCAEGPEKMLVYEYLPNKS 200
Query: 513 LDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
LD FLFD +K+ LDW R IV GIARG++YLH ++ +IIHRD+KASNILLD+++ PK
Sbjct: 201 LDHFLFD--KKRSLDWMTRFRIVTGIARGLLYLHEEAPERIIHRDIKASNILLDEKLNPK 374
Query: 573 ISDFGLARIVKGGEGDEANTKRV 595
ISDFGLAR+ GE T R+
Sbjct: 375 ISDFGLARLFP-GEDTHVQTFRI 440
Score = 95.9 bits (237), Expect(2) = 7e-65
Identities = 49/115 (42%), Positives = 74/115 (63%)
Frame = +2
Query: 600 GYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENT 659
GYM PEYA+ G S K+DV+S+GVL+LEIVSGR+N+ + + L+ +AWKL+
Sbjct: 446 GYMAPEYALRGYLSVKTDVFSYGVLVLEIVSGRKNHDLKLDAEKADLLSYAWKLYQGGKI 625
Query: 660 ISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLMLISEITHLPPP 714
+ LID+ + + + + + C+ +GLLC Q ERP +++V LML S+ LP P
Sbjct: 626 MDLIDQNIGKYNGDEAAM-CIQLGLLCCQASLVERPDMNSVNLMLSSDSFTLPKP 787
>BG453591 similar to PIR|T05181|T05 S-receptor kinase (EC 2.7.1.-) T6K22.120
precursor - Arabidopsis thaliana, partial (24%)
Length = 640
Score = 238 bits (608), Expect(2) = 3e-63
Identities = 118/171 (69%), Positives = 143/171 (83%)
Frame = +3
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ D +EIAVKRLS +S QG EF NEV++ SKLQHRNLV++LGCC++ EKML+YE+MPN
Sbjct: 129 VLDRREIAVKRLSGSSKQGTREFKNEVILCSKLQHRNLVKVLGCCIQGEEKMLIYEYMPN 308
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
+SLD+FLFD QKK LDW KR NI+ GIARG++YLH+DSRL+IIHRDLK SNILLD++M
Sbjct: 309 RSLDSFLFDQAQKKLLDWSKRFNIICGIARGLIYLHQDSRLRIIHRDLKPSNILLDNDMN 488
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSF 621
PKISDFGLA+I G + E NT RVVGT+GYM PEYA+ GLFS KSDV+SF
Sbjct: 489 PKISDFGLAKIC-GDDQVEGNTNRVVGTHGYMAPEYAIDGLFSIKSDVFSF 638
Score = 22.7 bits (47), Expect(2) = 3e-63
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = +2
Query: 432 CHKQFSF**YAWQGRFWPGI 451
CH+ ** AW+ FW GI
Sbjct: 56 CHQ*LLQ**QAWRRWFWTGI 115
>TC79762 similar to PIR|H96731|H96731 hypothetical protein F5A18.8
[imported] - Arabidopsis thaliana, partial (55%)
Length = 1564
Score = 236 bits (603), Expect = 2e-62
Identities = 116/256 (45%), Positives = 182/256 (70%), Gaps = 2/256 (0%)
Frame = +3
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG+E+AVK+LS+ S QG +EFMNE +++++QH+N+V LLG CV EK+LVYE++P+
Sbjct: 441 LSDGREVAVKKLSQTSNQGKKEFMNEAKLLARVQHKNVVNLLGYCVHGTEKILVYEYVPH 620
Query: 511 KSLDAFLFDPIQKK-KLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEM 569
+SLD FLF +K+ +LDW++R I+ G+A+G++YLH DS IIHRD+KASNILLDD+
Sbjct: 621 ESLDKFLFKEAEKREQLDWKRRFGIITGVAKGLLYLHEDSHNCIIHRDIKASNILLDDKW 800
Query: 570 IPKISDFGLARIVKGGEGDEANTK-RVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEI 628
KI+DFG+AR+ D++ K RV GT GYM PEY M G S K+DV+S+GVL+LE+
Sbjct: 801 TAKIADFGMARLF---PEDQSQVKTRVAGTNGYMAPEYMMHGRLSVKADVFSYGVLVLEL 971
Query: 629 VSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQ 688
++G+RN+SF + +L+ +A+K++ + ++ ++D + + C+ + LLC+Q
Sbjct: 972 ITGQRNSSFNLXVEEHNLLDWAYKMYKKGRSLEIVDSALASTVLTEQVDMCIQLALLCIQ 1151
Query: 689 ELPKERPSISTVVLML 704
P+ RP++ +V+ L
Sbjct: 1152GDPQLRPTMRRIVVKL 1199
>TC80500 similar to PIR|G96602|G96602 probable receptor protein kinase
F14G9.24 [imported] - Arabidopsis thaliana, partial
(13%)
Length = 1015
Score = 232 bits (591), Expect = 5e-61
Identities = 137/309 (44%), Positives = 199/309 (64%), Gaps = 9/309 (2%)
Frame = +1
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG+++AVK+LS S QG +F+ E+ IS +QHRNLV+L GCC+E +++LVYE++ N
Sbjct: 28 LNDGRDVAVKQLSIGSHQGKSQFVAEIATISAVQHRNLVKLYGCCIEGSKRLLVYEYLEN 207
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD LF + L+W R +I G+ARG+ YLH +SRL+I+HRD+KASNILLD E++
Sbjct: 208 KSLDQALFGNVLF--LNWSTRYDICMGVARGLTYLHEESRLRIVHRDVKASNILLDSELV 381
Query: 571 PKISDFGLARIVKGGEGDEANT---KRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLE 627
PKISDFGLA++ D+ T RV GT GY+ PEYAM G +EK+DV+SFGV+ LE
Sbjct: 382 PKISDFGLAKLY-----DDKKTHISTRVAGTIGYLAPEYAMRGHLTEKADVFSFGVVALE 546
Query: 628 IVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTIS-LIDREVWDASFESSMLRCMHIGLLC 686
+VSGR N+ + + L+ +AW+L E NTI+ LID + + + E + R + I LLC
Sbjct: 547 LVSGRPNSDSTLEGEKMYLLEWAWQLH-ERNTINELIDPRLSEFNKE-EVQRLVGIALLC 720
Query: 687 VQELPKERPSISTVVLMLISEI---THLPPPGKVA--FVHNQNSRSTESSQQSHRSNSNN 741
Q P RPS+S VV ML +I T PG + + +S T+ S + +++ N
Sbjct: 721 TQTSPTLRPSMSRVVAMLSGDIEVGTVTSRPGYLTDWKFDDVSSIMTDVSAKGLDTSNYN 900
Query: 742 NVTLSDVIG 750
+ + ++G
Sbjct: 901 STASTSIVG 927
>TC80060 similar to PIR|D96574|D96574 hypothetical protein T3F20.24
[imported] - Arabidopsis thaliana, partial (32%)
Length = 1547
Score = 232 bits (591), Expect = 5e-61
Identities = 125/258 (48%), Positives = 167/258 (64%), Gaps = 4/258 (1%)
Frame = +1
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG IAVK+LS S QG EF+NE+ +IS LQH NLV+L GCC+E + +LVYE+M N
Sbjct: 250 LSDGAVIAVKQLSSKSKQGNREFVNEIGMISALQHPNLVKLYGCCIEGNQLLLVYEYMEN 429
Query: 511 KSLDAFLFD-PIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEM 569
SL LF P Q+ LDWR R I GIARG+ YLH +SRLKI+HRD+KA+N+LLD +
Sbjct: 430 NSLARALFGKPEQRLNLDWRTRMKICVGIARGLAYLHEESRLKIVHRDIKATNVLLDKNL 609
Query: 570 IPKISDFGLARIVKGGEGDEANTK---RVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLL 626
KISDFGLA++ +E NT R+ GT GYM PEYAM G ++K+DVYSFGV+ L
Sbjct: 610 NAKISDFGLAKL-----DEEENTHISTRIAGTIGYMAPEYAMRGYLTDKADVYSFGVVAL 774
Query: 627 EIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLC 686
EIVSG N ++ E+ + L+ +A+ L + N + L+D + +R + + LLC
Sbjct: 775 EIVSGMSNTNYRPKEEFVYLLDWAYVLQEQGNLLELVDPTLGSKYSSEEAMRMLQLALLC 954
Query: 687 VQELPKERPSISTVVLML 704
P RP +S+VV ML
Sbjct: 955 TNPSPTLRPPMSSVVSML 1008
>TC92541 similar to PIR|T02153|T02153 protein kinase homolog T1F15.1 -
Arabidopsis thaliana, partial (22%)
Length = 791
Score = 123 bits (309), Expect(3) = 7e-60
Identities = 57/89 (64%), Positives = 74/89 (83%)
Frame = +2
Query: 454 GQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSL 513
GQEIAVKRLSK S QGI EF NE+V+I +LQH NLV+LLGCC+ E++L+YE+M NKSL
Sbjct: 86 GQEIAVKRLSKTSRQGIVEFKNELVLICELQHTNLVQLLGCCIHEEERILIYEYMSNKSL 265
Query: 514 DAFLFDPIQKKKLDWRKRSNIVEGIARGI 542
D +LFD ++K LDW+KR NI+EGI++G+
Sbjct: 266 DFYLFDSTRRKCLDWKKRLNIIEGISQGL 352
Score = 88.2 bits (217), Expect(3) = 7e-60
Identities = 51/113 (45%), Positives = 63/113 (55%), Gaps = 25/113 (22%)
Frame = +1
Query: 576 FGLARIVKGGEGDEANTKRVVGT-------------------------YGYMPPEYAMGG 610
FG+AR+ E NT R+VGT GYM PEYAM G
Sbjct: 454 FGMARMFTQQES-VVNTNRIVGT**VL*TFLMIT*RKRF*FFK*FFLSSGYMSPEYAMEG 630
Query: 611 LFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLI 663
+ S KSDVYSFGVLLLEI+ GRRNNSFY + L+L+G AW+LW + + L+
Sbjct: 631 ICSTKSDVYSFGVLLLEIICGRRNNSFYDVDRPLNLIGHAWELWNDGEYLQLM 789
Score = 59.3 bits (142), Expect(3) = 7e-60
Identities = 28/34 (82%), Positives = 31/34 (90%)
Frame = +3
Query: 543 MYLHRDSRLKIIHRDLKASNILLDDEMIPKISDF 576
+YLH+ SRLKIIHRDLKASNILLD+ M PKISDF
Sbjct: 354 LYLHKYSRLKIIHRDLKASNILLDENMNPKISDF 455
>BQ155172 weakly similar to GP|6686398|gb F1E22.15 {Arabidopsis thaliana},
partial (10%)
Length = 637
Score = 210 bits (535), Expect(2) = 4e-56
Identities = 104/178 (58%), Positives = 136/178 (75%)
Frame = +3
Query: 542 IMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGY 601
++YLH+DSRL++IHRDLK SNILLD+EM PKISDFGLARIV GG+ EA+T+RVVGTYGY
Sbjct: 78 LLYLHQDSRLRVIHRDLKTSNILLDEEMQPKISDFGLARIV-GGKETEASTERVVGTYGY 254
Query: 602 MPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTIS 661
M PEYA+ G FS KSDV+SFGV+LLEI+SG++N FY++++ SL+G+AW+LW E+
Sbjct: 255 MSPEYALDGYFSTKSDVFSFGVVLLEIISGKKNTGFYRSKEISSLLGYAWRLWREDKLQD 434
Query: 662 LIDREVWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVAF 719
L+D + D + +RC IGLLCVQ+ P +RP +S VV ML +E T L P + F
Sbjct: 435 LMDPTLSDTYNVNQFIRCSQIGLLCVQDEPDDRPHMSNVVTMLDNETTTLLTPKQPTF 608
Score = 26.9 bits (58), Expect(2) = 4e-56
Identities = 12/25 (48%), Positives = 15/25 (60%)
Frame = +2
Query: 517 LFDPIQKKKLDWRKRSNIVEGIARG 541
+FD + LDW R I+ GIARG
Sbjct: 2 IFDSTKSVILDWPMRFEIILGIARG 76
>TC88193 similar to PIR|G96602|G96602 probable receptor protein kinase
F14G9.24 [imported] - Arabidopsis thaliana, partial
(20%)
Length = 1815
Score = 213 bits (542), Expect = 2e-55
Identities = 116/246 (47%), Positives = 168/246 (68%), Gaps = 4/246 (1%)
Frame = +3
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG+ +AVK+LS S QG +F+ E+ IS +QHRNLV+L GCC+E +++LVYE++ N
Sbjct: 633 LNDGRFVAVKQLSIGSHQGKSQFIAEIATISAVQHRNLVKLYGCCIEGNKRLLVYEYLEN 812
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD LF + L+W R ++ G+ARG+ YLH +SRL+I+HRD+KASNILLD E++
Sbjct: 813 KSLDQALFGNVLF--LNWSTRYDVCMGVARGLTYLHEESRLRIVHRDVKASNILLDSELV 986
Query: 571 PKISDFGLARIVKGGEGDEANT---KRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLE 627
PK+SDFGLA++ D+ T RV GT GY+ PEYAM G +EK+DV+SFGV+ LE
Sbjct: 987 PKLSDFGLAKLY-----DDKKTHISTRVAGTIGYLAPEYAMRGRLTEKADVFSFGVVALE 1151
Query: 628 IVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTIS-LIDREVWDASFESSMLRCMHIGLLC 686
+VSGR N+ ED + L+ +AW+L E N I+ LID + + + E + R + IG++
Sbjct: 1152LVSGRPNSDSSLEEDKMYLLDWAWQLH-ERNCINDLIDPRLSEFNME-EVERLVGIGIVV 1325
Query: 687 VQELPK 692
++ K
Sbjct: 1326HSDITK 1343
Score = 32.7 bits (73), Expect = 0.53
Identities = 15/26 (57%), Positives = 18/26 (68%)
Frame = +1
Query: 683 GLLCVQELPKERPSISTVVLMLISEI 708
GLLC Q P RPS+S VV ML+ +I
Sbjct: 1315 GLLCTQTSPNLRPSMSRVVAMLLGDI 1392
>TC89703 similar to GP|11994595|dbj|BAB02650. receptor-like serine/threonine
kinase {Arabidopsis thaliana}, partial (26%)
Length = 1212
Score = 133 bits (335), Expect(2) = 6e-55
Identities = 66/123 (53%), Positives = 87/123 (70%), Gaps = 1/123 (0%)
Frame = +2
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG IAVK LS S QG EF+NE+ +IS LQH LV+L GCCVE + ML+YE++ N
Sbjct: 89 LSDGTLIAVKLLSSKSKQGNREFLNEIGMISALQHPQLVKLYGCCVEGDQLMLIYEYLEN 268
Query: 511 KSLDAFLFDPIQKK-KLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEM 569
SL LF P + + +LDW R I GIARG+ YLH +SRLK++HRD+KA+N+LLD ++
Sbjct: 269 NSLARALFGPAEHQIRLDWPTRYKICVGIARGLAYLHEESRLKVVHRDIKATNVLLDKDL 448
Query: 570 IPK 572
PK
Sbjct: 449 NPK 457
Score = 100 bits (248), Expect(2) = 6e-55
Identities = 57/142 (40%), Positives = 84/142 (59%), Gaps = 3/142 (2%)
Frame = +3
Query: 570 IPKISDFGLARIVKGGEGDEANTK---RVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLL 626
I KISDFGLA++ +E NT R+ GTYGYM PEYAM G ++K+DVYSFG++ L
Sbjct: 450 IRKISDFGLAKL-----DEEENTHISTRIAGTYGYMAPEYAMHGYLTDKADVYSFGIVAL 614
Query: 627 EIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLC 686
EI+ G N Q E++ L+ +A L + N I L+D+ + + + +++ LLC
Sbjct: 615 EILHGSNNTILRQKEEAFHLLDWAHILKEKGNEIELVDKRLGSNFNKEEAMLMINVALLC 794
Query: 687 VQELPKERPSISTVVLMLISEI 708
RP++S+VV ML +I
Sbjct: 795 TNVTSSLRPAMSSVVSMLEGKI 860
>BE942940 weakly similar to GP|13506749|gb receptor-like protein kinase 6
{Arabidopsis thaliana}, partial (23%)
Length = 466
Score = 208 bits (530), Expect = 5e-54
Identities = 105/156 (67%), Positives = 127/156 (81%)
Frame = +1
Query: 483 LQHRNLVRLLGCCVERGEKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGI 542
LQHRNLV +G VE EK+L+YE++ NKSLD FLFD Q+K L W +R NI+ GIARGI
Sbjct: 1 LQHRNLVTFIGFWVEGHEKILIYEYVSNKSLDYFLFDSHQQKLLTWVERFNIIGGIARGI 180
Query: 543 MYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYM 602
+YLH SRLK+IHRDLK SNILLD+ MIPKISDFGLAR+V+ + +E +T R+VGTYGYM
Sbjct: 181 LYLHEHSRLKVIHRDLKPSNILLDENMIPKISDFGLARMVEISQ-EEGSTNRIVGTYGYM 357
Query: 603 PPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFY 638
PEYAM G FSEKSDVYSFGV++LEIV+G++N S Y
Sbjct: 358 SPEYAMFGQFSEKSDVYSFGVMILEIVAGKKNISSY 465
>TC80877 weakly similar to GP|5814093|gb|AAD52097.1| receptor-like kinase
CHRK1 {Nicotiana tabacum}, partial (26%)
Length = 1229
Score = 208 bits (530), Expect = 5e-54
Identities = 105/232 (45%), Positives = 154/232 (66%), Gaps = 1/232 (0%)
Frame = +1
Query: 519 DPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGL 578
DP + LDWRKR N++EGI +G++YL S IIHRD+KASN+LLD EM PKISDFG+
Sbjct: 244 DPRKSILLDWRKRVNVIEGITQGLLYLQEYSNFTIIHRDIKASNVLLDHEMNPKISDFGM 423
Query: 579 ARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFY 638
ARI G EANT R+VGTYGY+PPEY G++S K DVYSFGVLLL+I+SG+R + +Y
Sbjct: 424 ARIF-GKYELEANTSRIVGTYGYVPPEYVKKGIYSPKYDVYSFGVLLLQIISGKRTSHYY 600
Query: 639 QNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSIS 698
++++L+ +A++LW+E + D + D++ ++RCM + LLCVQE +RPS+
Sbjct: 601 GTHENMNLLEYAYELWIEGRGMEFFDPSLDDSTSHCKIMRCMQVALLCVQENSSDRPSML 780
Query: 699 TVVLMLISEITHLPPPGKVAF-VHNQNSRSTESSQQSHRSNSNNNVTLSDVI 749
V +L +E ++ P AF + ++S + +S N+VT+S ++
Sbjct: 781 EVDSLLKNEGAYVGTPNVPAFSMKKHEDDKGDTSNSGFKFSSINDVTISQMV 936
Score = 34.3 bits (77), Expect = 0.18
Identities = 13/18 (72%), Positives = 17/18 (94%)
Frame = +1
Query: 501 KMLVYEFMPNKSLDAFLF 518
K+L+YE++PNKSLD FLF
Sbjct: 28 KLLIYEYLPNKSLDHFLF 81
>TC92127 weakly similar to GP|16040952|dbj|BAB69683. receptor kinase 5
{Brassica rapa}, partial (18%)
Length = 507
Score = 203 bits (517), Expect = 2e-52
Identities = 101/167 (60%), Positives = 130/167 (77%)
Frame = +3
Query: 538 IARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVG 597
IA+G++YLH+ SRL+IIHRDLKASNILLD+ M PKISDFG+AR+ E +ANT R+VG
Sbjct: 3 IAQGLLYLHKYSRLRIIHRDLKASNILLDENMNPKISDFGVARMFTRQE-TKANTNRIVG 179
Query: 598 TYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEE 657
TYGYM PEYAM G+FS KSDVYSFGVLLLEI++G +NNSFY + L+LVG AW+LW E
Sbjct: 180 TYGYMSPEYAMEGVFSTKSDVYSFGVLLLEIINGEKNNSFYCEDRPLNLVGHAWELWKEG 359
Query: 658 NTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLML 704
+ L+D + ++ E +LRC+H GLLCV+E +RP++S V+ ML
Sbjct: 360 VVLELVDPLLNESFSEDEVLRCVHAGLLCVEENADDRPTMSNVIAML 500
>TC78270 similar to GP|16649103|gb|AAL24403.1 Unknown protein {Arabidopsis
thaliana}, partial (80%)
Length = 1463
Score = 203 bits (516), Expect = 2e-52
Identities = 118/308 (38%), Positives = 177/308 (57%), Gaps = 6/308 (1%)
Frame = +3
Query: 445 GRFWPGIQDGQEIA-VKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKML 503
G + G+ G ++A +K LS S QG++EF+ E+ VIS+++H NLV L GCCVE ++L
Sbjct: 345 GSVYKGVLKGGKLAAIKVLSTESKQGVKEFLTEINVISEIKHENLVILYGCCVEGDHRIL 524
Query: 504 VYEFMPNKSLDAFLFDPIQKK-KLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASN 562
VY ++ N SL L DW+ R I G+ARG+ +LH + I+HRD+KASN
Sbjct: 525 VYNYLENNSLSQTLLAGGHSNIYFDWQTRRRICLGVARGLAFLHEEVLPHIVHRDIKASN 704
Query: 563 ILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFG 622
ILLD ++ PKISDFGLA+++ + RV GT GY+ PEYA+ G + K+D+YSFG
Sbjct: 705 ILLDKDLTPKISDFGLAKLIPSYMTHVST--RVAGTIGYLAPEYAIRGQLTRKADIYSFG 878
Query: 623 VLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHI 682
VLL+EIVSGR N + ++ W+L+ + L+D + + + I
Sbjct: 879 VLLVEIVSGRSNTNTRLPIADQYILETTWQLYERKELAQLVDISLNGEFDAEEACKILKI 1058
Query: 683 GLLCVQELPKERPSISTVVLMLISEI----THLPPPGKVAFVHNQNSRSTESSQQSHRSN 738
LLC Q+ PK RP++S+VV ML E+ T + PG ++ V + R + + +
Sbjct: 1059ALLCTQDTPKLRPTMSSVVKMLTGEMDINETKITKPGLISDVMDLKIREPKKNINMGTAP 1238
Query: 739 SNNNVTLS 746
S+ N + S
Sbjct: 1239SSYNASSS 1262
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.358 0.159 0.601
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,951,608
Number of Sequences: 36976
Number of extensions: 437222
Number of successful extensions: 6786
Number of sequences better than 10.0: 664
Number of HSP's better than 10.0 without gapping: 2417
Number of HSP's successfully gapped in prelim test: 285
Number of HSP's that attempted gapping in prelim test: 3636
Number of HSP's gapped (non-prelim): 2985
length of query: 750
length of database: 9,014,727
effective HSP length: 103
effective length of query: 647
effective length of database: 5,206,199
effective search space: 3368410753
effective search space used: 3368410753
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC144721.3


