

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
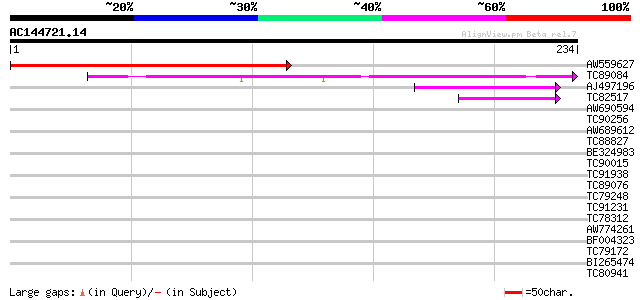
Query= AC144721.14 - phase: 0
(234 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW559627 228 2e-60
TC89084 weakly similar to GP|9759369|dbj|BAB09828.1 gene_id:MYJ2... 54 5e-08
AJ497196 weakly similar to SP|O60683|PEXA_ Peroxisome assembly p... 47 5e-06
TC82517 homologue to GP|7688065|emb|CAB89694.1 constitutively ph... 42 2e-04
AW690594 similar to GP|23498163|emb hypothetical protein {Plasmo... 40 0.001
TC90256 similar to GP|22296435|dbj|BAC10202. contains ESTs AU055... 40 0.001
AW689612 weakly similar to GP|19698963|gb| unknown protein {Arab... 40 0.001
TC88827 similar to GP|22165059|gb|AAM93676.1 unknown protein {Or... 39 0.002
BE324983 similar to PIR|T48442|T48 hypothetical protein T32M21.6... 39 0.002
TC90015 similar to PIR|T01393|T01393 apoptosis inhibitor homolog... 38 0.003
TC91938 similar to PIR|E96612|E96612 probable transcription fact... 38 0.004
TC89076 similar to GP|18481708|gb|AAL73530.1 hypothetical protei... 37 0.005
TC79248 similar to GP|22165059|gb|AAM93676.1 unknown protein {Or... 37 0.007
TC91231 similar to GP|9502381|gb|AAF88088.1| T12C24.29 {Arabidop... 37 0.009
TC78312 similar to GP|19698885|gb|AAL91178.1 putative protein {A... 37 0.009
AW774261 weakly similar to EGAD|146423|156 vitellogenin {Anolis ... 36 0.011
BF004323 similar to GP|4337011|gb| zinc-binding peroxisomal inte... 36 0.015
TC79172 similar to GP|9502381|gb|AAF88088.1| T12C24.29 {Arabidop... 36 0.015
BI265474 weakly similar to PIR|T48058|T480 RING-H2 zinc finger p... 35 0.032
TC80941 weakly similar to PIR|T39903|T39903 serine-rich protein ... 34 0.042
>AW559627
Length = 434
Score = 228 bits (580), Expect = 2e-60
Identities = 116/116 (100%), Positives = 116/116 (100%)
Frame = +1
Query: 1 MENRQRITLYDQMTSTSNTRSSLASLIQNDAVLSNQKNTNQTLLDIIRENEPNNVVVNVS 60
MENRQRITLYDQMTSTSNTRSSLASLIQNDAVLSNQKNTNQTLLDIIRENEPNNVVVNVS
Sbjct: 82 MENRQRITLYDQMTSTSNTRSSLASLIQNDAVLSNQKNTNQTLLDIIRENEPNNVVVNVS 261
Query: 61 VTNSSSSSNNNVVKDRKSWKAFKELLLFKRRTSDESTHQQQQNEDIPTNSDPSEFL 116
VTNSSSSSNNNVVKDRKSWKAFKELLLFKRRTSDESTHQQQQNEDIPTNSDPSEFL
Sbjct: 262 VTNSSSSSNNNVVKDRKSWKAFKELLLFKRRTSDESTHQQQQNEDIPTNSDPSEFL 429
>TC89084 weakly similar to GP|9759369|dbj|BAB09828.1
gene_id:MYJ24.10~ref|NP_055178.1~similar to unknown
protein {Arabidopsis thaliana}, partial (3%)
Length = 1200
Score = 53.9 bits (128), Expect = 5e-08
Identities = 45/210 (21%), Positives = 87/210 (41%), Gaps = 8/210 (3%)
Frame = +1
Query: 33 LSNQKNTNQTLLDIIRENEPNNVVVNVSVTNSSSSSNNNVVKDRKSWKAFKELLLFKRRT 92
+S+ + Q +D+I + ++ VT + SS K + KE L+
Sbjct: 604 MSDHQQVKQLCIDLILK-------LHQGVTQAFLSSQVFSFKSVSASSPLKETLVHDSPI 762
Query: 93 SD-------ESTHQQQQNEDIPTNSDPSEFLPGGEFSDETAQV-SMSLMDLLEESEIELE 144
S+ ES+ + + N +P+ + S + E ++ S + +++ E + L+
Sbjct: 763 SNRSRVEFPESSGRGETNSQVPSLREQSGKVSAAELVHAVNEILSAAGINMDAEKQALLQ 942
Query: 145 RISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGHT 204
+ +D ++++E+ E+E+ E E K C VC+ + +PCGH
Sbjct: 943 K---TIDLQENLKESQASLLLEQEKVERSTREADTAKAAWTCRVCLSSEVDITIVPCGHV 1113
Query: 205 FCRMCSRELWVSRGNCPLCNHFILEVLDIF 234
CR CS + CP C + + IF
Sbjct: 1114LCRKCSSAV----SKCPFCRLSSTKAIRIF 1191
>AJ497196 weakly similar to SP|O60683|PEXA_ Peroxisome assembly protein 10
(Peroxin-10). [Human] {Homo sapiens}, partial (11%)
Length = 582
Score = 47.4 bits (111), Expect = 5e-06
Identities = 20/60 (33%), Positives = 28/60 (46%)
Frame = +1
Query: 168 EREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRGNCPLCNHFI 227
E EE+G + +V + C +C+ PCGH FC C +R CPLC F+
Sbjct: 256 ELEEDGPPKSDSVLTYYRCSLCLEARTQDTATPCGHVFCWTCISGWLQTRAQCPLCRDFV 435
>TC82517 homologue to GP|7688065|emb|CAB89694.1 constitutively
photomorphogenic 1 protein {Pisum sativum}, partial
(22%)
Length = 671
Score = 42.0 bits (97), Expect = 2e-04
Identities = 16/42 (38%), Positives = 23/42 (54%)
Frame = +3
Query: 186 CCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRGNCPLCNHFI 227
C +CM K A CGH+FC MC ++ +CP C H++
Sbjct: 159 CPICMQIIKDAFLTSCGHSFCYMCIITHLRNKSDCPCCGHYL 284
>AW690594 similar to GP|23498163|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (10%)
Length = 633
Score = 39.7 bits (91), Expect = 0.001
Identities = 30/91 (32%), Positives = 43/91 (46%)
Frame = +3
Query: 90 RRTSDESTHQQQQNEDIPTNSDPSEFLPGGEFSDETAQVSMSLMDLLEESEIELERISDV 149
+ T ES+ ++++ E+ D E E DE + D EE E E E
Sbjct: 3 KMTMAESSWKRRKVEEHNEEEDEGEV---EEEEDEHDEEEEDEHDEEEEEEEEEEE---- 161
Query: 150 VDEGDDVEENNEKREEEEEREEEGEGEGSNV 180
DE DD EE E+ EEEEE +++ EGE +
Sbjct: 162 EDEHDDEEEEEEEEEEEEEEDDDEEGEEDEI 254
Score = 35.0 bits (79), Expect = 0.025
Identities = 20/47 (42%), Positives = 26/47 (54%)
Frame = +3
Query: 137 EESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVE 183
+E E+E E DE D+ EE+ EEEEE EEE E E + + E
Sbjct: 66 DEGEVEEEE-----DEHDEEEEDEHDEEEEEEEEEEEEDEHDDEEEE 191
Score = 30.4 bits (67), Expect = 0.61
Identities = 26/98 (26%), Positives = 44/98 (44%)
Frame = +3
Query: 88 FKRRTSDESTHQQQQNEDIPTNSDPSEFLPGGEFSDETAQVSMSLMDLLEESEIELERIS 147
+KRR +E H ++++E GE +E + D +E E E
Sbjct: 27 WKRRKVEE--HNEEEDE--------------GEVEEEEDEHDEEEEDEHDEEEEE----- 143
Query: 148 DVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHN 185
+E ++ E+ ++ EEEEE EEE E E + + E +
Sbjct: 144 ---EEEEEEEDEHDDEEEEEEEEEEEEEEDDDEEGEED 248
>TC90256 similar to GP|22296435|dbj|BAC10202. contains ESTs AU055804(S20063)
AU055998(S20213) AU055803(S20063)~similar to ankyrin
homolog, partial (16%)
Length = 812
Score = 39.7 bits (91), Expect = 0.001
Identities = 21/75 (28%), Positives = 32/75 (42%), Gaps = 1/75 (1%)
Frame = +1
Query: 161 EKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGHTF-CRMCSRELWVSRGN 219
EK EE + G G C +C+ A IPCGH C C E+ +
Sbjct: 172 EKLPNEEGKTAGGSGS--------TCVICLDAPAEGACIPCGHVAGCMSCLNEVKTKKWG 327
Query: 220 CPLCNHFILEVLDIF 234
CP+C I +++ ++
Sbjct: 328 CPVCRAKIDQIIKLY 372
>AW689612 weakly similar to GP|19698963|gb| unknown protein {Arabidopsis
thaliana}, partial (2%)
Length = 653
Score = 39.7 bits (91), Expect = 0.001
Identities = 21/83 (25%), Positives = 35/83 (41%)
Frame = +1
Query: 142 ELERISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPC 201
EL++++ + + G + NN K + + +E C +C+ K +PC
Sbjct: 16 ELQKLTKMENSGKSIMNNNTKLLNPWMLHFQ------KLALELKCPLCLNLFKKPVLLPC 177
Query: 202 GHTFCRMCSRELWVSRGNCPLCN 224
H FC C + R C LCN
Sbjct: 178 NHLFCSSCLADSTSIRSECALCN 246
>TC88827 similar to GP|22165059|gb|AAM93676.1 unknown protein {Oryza sativa
(japonica cultivar-group)}, complete
Length = 1221
Score = 38.9 bits (89), Expect = 0.002
Identities = 19/83 (22%), Positives = 35/83 (41%), Gaps = 2/83 (2%)
Frame = +3
Query: 150 VDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHN--CCVCMVRDKGAAFIPCGHTFCR 207
V + +D ++ E R+++ + S++ +E + C +CM + C H C
Sbjct: 474 VTDSEDKKQKAVCMERYRRRDDDDCRQSSDIDIERDDECGICMEMNSKIVLPNCNHVMCL 653
Query: 208 MCSRELWVSRGNCPLCNHFILEV 230
C RE +CP C + V
Sbjct: 654 KCYREWRTRSQSCPFCRDSLKRV 722
>BE324983 similar to PIR|T48442|T48 hypothetical protein T32M21.60 -
Arabidopsis thaliana, partial (13%)
Length = 676
Score = 38.5 bits (88), Expect = 0.002
Identities = 19/47 (40%), Positives = 23/47 (48%), Gaps = 1/47 (2%)
Frame = +1
Query: 186 CCVCMVRDKGAAFIPCGHTF-CRMCSRELWVSRGNCPLCNHFILEVL 231
CCVC + CGH C C+ EL G CPLC I+EV+
Sbjct: 298 CCVCCDNHIDSLLYRCGHMCTCSKCASELIRGGGKCPLCRAPIVEVV 438
>TC90015 similar to PIR|T01393|T01393 apoptosis inhibitor homolog T4I9.12 -
Arabidopsis thaliana, partial (8%)
Length = 757
Score = 38.1 bits (87), Expect = 0.003
Identities = 20/60 (33%), Positives = 29/60 (48%), Gaps = 3/60 (5%)
Frame = +1
Query: 177 GSNVKVEHNCCVCMVRDKGAAFIPCGH-TFCRMCSRELWVSRG--NCPLCNHFILEVLDI 233
G VK E C +C+ + F+PC H C C+ EL +G +CP C I E + +
Sbjct: 424 GGGVKRERECVMCLSEEMSVVFLPCAHQVVCTKCN-ELHEKQGMQDCPSCRSPIQERISV 600
>TC91938 similar to PIR|E96612|E96612 probable transcription factor
F12K22.14 [imported] - Arabidopsis thaliana, partial
(11%)
Length = 685
Score = 37.7 bits (86), Expect = 0.004
Identities = 25/85 (29%), Positives = 34/85 (39%), Gaps = 19/85 (22%)
Frame = +3
Query: 158 ENNEKREEEEEREEEGEGEGSNVKVEH-----------------NCCVCMVRDKGAAFIP 200
EN+ EEE+R++ E G ++K + NC C+ + P
Sbjct: 312 ENDPSLTEEEKRKKRQELHGGSLKEKDEVHVRRSGVLDIFDGSLNCSFCVKLPERPVTTP 491
Query: 201 CGHTFCRMCSRELWVSRG--NCPLC 223
CGH FC C E WV G C C
Sbjct: 492 CGHNFCLKCF-EKWVGLGKRTCSNC 563
>TC89076 similar to GP|18481708|gb|AAL73530.1 hypothetical protein {Sorghum
bicolor}, partial (65%)
Length = 1326
Score = 37.4 bits (85), Expect = 0.005
Identities = 19/49 (38%), Positives = 24/49 (48%)
Frame = +2
Query: 186 CCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRGNCPLCNHFILEVLDIF 234
C VC+ K AF CGH CR C L +CP+C H I L ++
Sbjct: 1043 CPVCLTNAKDLAF-GCGHMTCRDCGSRL----RHCPICRHRITSRLRVY 1174
>TC79248 similar to GP|22165059|gb|AAM93676.1 unknown protein {Oryza sativa
(japonica cultivar-group)}, partial (97%)
Length = 1272
Score = 37.0 bits (84), Expect = 0.007
Identities = 21/81 (25%), Positives = 31/81 (37%), Gaps = 5/81 (6%)
Frame = +2
Query: 155 DVEENNEK---REEEEEREEEGEGEGSNVKVEHN--CCVCMVRDKGAAFIPCGHTFCRMC 209
D E+ +K E RE+E + S++ E C +CM + C H C C
Sbjct: 452 DAEDKKQKVVCMERYRRREDEEHKQFSDIDFEREEECGICMEMNSKIVLPNCNHVMCLKC 631
Query: 210 SRELWVSRGNCPLCNHFILEV 230
E +CP C + V
Sbjct: 632 YHEWRARSQSCPFCRDSLKRV 694
>TC91231 similar to GP|9502381|gb|AAF88088.1| T12C24.29 {Arabidopsis
thaliana}, partial (17%)
Length = 728
Score = 36.6 bits (83), Expect = 0.009
Identities = 18/47 (38%), Positives = 24/47 (50%), Gaps = 3/47 (6%)
Frame = +1
Query: 186 CCVCM-VRDKGAAF--IPCGHTFCRMCSRELWVSRGNCPLCNHFILE 229
CC+C+ D GA +PCGH F C + CPLC + +LE
Sbjct: 94 CCICLSAYDDGAELRQLPCGHHFHCTCVDKWLHINATCPLCKYNMLE 234
>TC78312 similar to GP|19698885|gb|AAL91178.1 putative protein {Arabidopsis
thaliana}, partial (14%)
Length = 1317
Score = 36.6 bits (83), Expect = 0.009
Identities = 27/115 (23%), Positives = 52/115 (44%), Gaps = 2/115 (1%)
Frame = +3
Query: 97 THQQQQNEDIPTNSDPSEFL--PGGEFSDETAQVSMSLMDLLEESEIELERISDVVDEGD 154
+H + +N I N+ FL G + + S++ ++ +EE E L+ + +
Sbjct: 993 SHLKHRNISIELNN----FLVNSGNGIGLQAGEGSVAAVERMEEEEAPLK----ACENNN 1148
Query: 155 DVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMC 209
V E+ ++ +++ E+ G G+G +C +C+ K CGH FC C
Sbjct: 1149 GVMEDETLQKNKDDVEKAGGGDGDFF----DCNICLDLAKEPVLTCCGHLFCWQC 1301
Score = 27.7 bits (60), Expect = 4.0
Identities = 10/19 (52%), Positives = 16/19 (83%)
Frame = -3
Query: 156 VEENNEKREEEEEREEEGE 174
+E+ +++E+EEE EEEGE
Sbjct: 223 IEKKRKQKEKEEEEEEEGE 167
>AW774261 weakly similar to EGAD|146423|156 vitellogenin {Anolis pulchellus},
partial (40%)
Length = 495
Score = 36.2 bits (82), Expect = 0.011
Identities = 29/97 (29%), Positives = 42/97 (42%)
Frame = +1
Query: 80 KAFKELLLFKRRTSDESTHQQQQNEDIPTNSDPSEFLPGGEFSDETAQVSMSLMDLLEES 139
KA L+ K E T + + D+ +DP E M+ +E
Sbjct: 217 KANCSLVAKKSEPEPEKTVESDEKIDLEEENDPEEE-----------------MEEIEYE 345
Query: 140 EIELERISDVVDEGDDVEENNEKREEEEEREEEGEGE 176
E+E E + ++E +VEEN E EEEEE E+ E E
Sbjct: 346 EVEEEEEVEEIEE--EVEENEEDAEEEEEE*EKEEEE 450
>BF004323 similar to GP|4337011|gb| zinc-binding peroxisomal integral
membrane protein {Arabidopsis thaliana}, partial (32%)
Length = 513
Score = 35.8 bits (81), Expect = 0.015
Identities = 19/71 (26%), Positives = 27/71 (37%), Gaps = 7/71 (9%)
Frame = -1
Query: 160 NEKREEEEEREEEGEGEGSNVKVEHN-------CCVCMVRDKGAAFIPCGHTFCRMCSRE 212
NE+ ++G + EHN C +C+ + CGH FC C E
Sbjct: 354 NEEGNLASPEADKGSWVPGSSSSEHNATNGVSKCTLCLSNRQHPTATSCGHVFCWNCITE 175
Query: 213 LWVSRGNCPLC 223
+ CPLC
Sbjct: 174 WCNEKPECPLC 142
>TC79172 similar to GP|9502381|gb|AAF88088.1| T12C24.29 {Arabidopsis
thaliana}, partial (71%)
Length = 1589
Score = 35.8 bits (81), Expect = 0.015
Identities = 25/98 (25%), Positives = 42/98 (42%), Gaps = 22/98 (22%)
Frame = +2
Query: 154 DDVEENNE----KREEEEEREEEGEGEGSNVK--------VEH-------NCCVCMVR-D 193
+D+E+ ++ K E E++ + +G + +EH CC+C+ D
Sbjct: 1034 EDIEQLSKFKFRKVESNEKQTDNNQGPVGGIMTECRADSPIEHVLAEEDAECCICLSSYD 1213
Query: 194 KGAAF--IPCGHTFCRMCSRELWVSRGNCPLCNHFILE 229
G +PCGH F C + CPLC + IL+
Sbjct: 1214 DGVELRELPCGHHFHCACVDKWLYINATCPLCKYNILK 1327
>BI265474 weakly similar to PIR|T48058|T480 RING-H2 zinc finger protein ATL5
- Arabidopsis thaliana, partial (19%)
Length = 617
Score = 34.7 bits (78), Expect = 0.032
Identities = 15/44 (34%), Positives = 20/44 (45%), Gaps = 4/44 (9%)
Frame = +3
Query: 184 HNCCVCMVR----DKGAAFIPCGHTFCRMCSRELWVSRGNCPLC 223
H+C VC+ D+ C H F +C + S NCPLC
Sbjct: 342 HDCAVCLSEFTDGDECLTLPNCNHDFHSLCVEPWFASHSNCPLC 473
>TC80941 weakly similar to PIR|T39903|T39903 serine-rich protein - fission
yeast (Schizosaccharomyces pombe), partial (15%)
Length = 557
Score = 34.3 bits (77), Expect = 0.042
Identities = 23/103 (22%), Positives = 48/103 (46%), Gaps = 7/103 (6%)
Frame = +2
Query: 77 KSWKAFKELLLFKRRTS-------DESTHQQQQNEDIPTNSDPSEFLPGGEFSDETAQVS 129
K WK ++LL R E+T +++ E I + + E ++ +
Sbjct: 17 KKWKRMTKMLLNLRMRKWMLMXEVKETTEDKEEKEKIEAEKEAEGSIEEKEEGNDEEKTE 196
Query: 130 MSLMDLLEESEIELERISDVVDEGDDVEENNEKREEEEEREEE 172
+ + +++E + E ++ +EGD+ E+ + EEEEE +E+
Sbjct: 197 VEEKEEGDDNE-KSEEETEEKEEGDEKEKTEAETEEEEEADED 322
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.128 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,419,693
Number of Sequences: 36976
Number of extensions: 110838
Number of successful extensions: 1566
Number of sequences better than 10.0: 218
Number of HSP's better than 10.0 without gapping: 1219
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1403
length of query: 234
length of database: 9,014,727
effective HSP length: 93
effective length of query: 141
effective length of database: 5,575,959
effective search space: 786210219
effective search space used: 786210219
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC144721.14


