

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
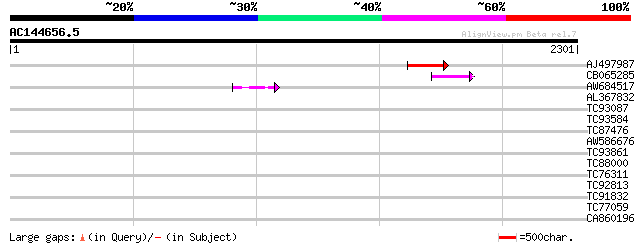
Query= AC144656.5 + phase: 0 /pseudo
(2301 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 179 1e-44
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 142 1e-33
AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thalia... 50 1e-05
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 40 0.010
TC93087 similar to GP|20160511|dbj|BAB89462. hypothetical protei... 36 0.15
TC93584 knotted class I homeodomain KNOX 33 1.7
TC87476 homologue to PIR|T11622|T11622 extensin class 1 precurso... 32 2.2
AW586676 similar to GP|806720|gb|AA arabinogalactan-protein prec... 32 3.7
TC93861 similar to GP|15451102|gb|AAK96822.1 Unknown protein {Ar... 32 3.7
TC88000 weakly similar to GP|13486661|dbj|BAB39898. contains EST... 31 4.8
TC76311 homologue to GP|3204132|emb|CAA07235.1 extensin {Cicer a... 31 4.8
TC92813 31 6.3
TC91832 similar to GP|13925771|gb|AAK49438.1 phytase {Glycine ma... 30 8.3
TC77059 similar to GP|18958499|gb|AAL82617.1 elongation factor 1... 30 8.3
CA860196 weakly similar to GP|1654183|gb|A MYOD-like DNA-binding... 30 8.3
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 179 bits (454), Expect = 1e-44
Identities = 100/167 (59%), Positives = 123/167 (72%)
Frame = -1
Query: 1613 EDLLRTSLGRQTPPSLYD*SYYLVDI*DGSDQVHL*ETRFDRKNCPVADVIIRI*Y*VSF 1672
ED+L +SLG Q PPSLYD*S+YL I*+GS+QVH+*E RFDRK+C VAD I I*Y V F
Sbjct: 636 EDMLCSSLGCQAPPSLYD*SHYLAGI*NGSNQVHI*EARFDRKDCTVADATIGI*YRVPF 457
Query: 1673 PESNQRQYPC*SLGSPTD*RLSAYQVRLS**RDHVFESERLR*TTAWGRSRSKIKMGFNI 1732
PE N RQY C*SLGSPT *RLS YQV LS**RD+V + ERL T RS S+ +GF+I
Sbjct: 456 PEGN*RQYSC*SLGSPTT*RLSTYQV*LS**RDYVSKDERL*RATIRRRS*SRFSVGFDI 277
Query: 1733 *RGC*CLW*RNWGNHYYS*GCTHPLHCQVDVRLYKQHGRVRSMYHGY 1779
* GC C+ R+ G+ YS* C+HP H + V L++QH +RS++HG+
Sbjct: 276 *WGCQCIRQRHRGSTPYS*RCSHPFHSKTPV*LHEQHR*IRSLHHGH 136
Score = 38.5 bits (88), Expect = 0.030
Identities = 19/24 (79%), Positives = 20/24 (83%)
Frame = +1
Query: 1578 GLCARTTRRNWKKGACHLLFEQEV 1601
GLCARTTR N KK A HLLF+QEV
Sbjct: 34 GLCARTTR*NRKKRARHLLFKQEV 105
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 142 bits (358), Expect = 1e-33
Identities = 83/176 (47%), Positives = 104/176 (58%)
Frame = -1
Query: 1711 ERLR*TTAWGRSRSKIKMGFNI*RGC*CLW*RNWGNHYYS*GCTHPLHCQVDVRLYKQHG 1770
+RLR T W RSRS+ +MGF++* C CLW*RNW +H G HP + VR+YKQ+G
Sbjct: 586 KRLRRTVDW*RSRSQ*QMGFSL*WCCQCLW*RNWSSHCIPAGALHPFYRPDSVRMYKQYG 407
Query: 1771 RVRSMYHGYRRSHRSENQEPRHLW*LSPCNQSNHMRMGDSSPRIGSLQRLCKTITDLLQQ 1830
V SMY R S+ ENQ PRHLW C+Q + MGD + SL RLC+ DL +
Sbjct: 406 *V*SMYLWDRGSN*HENQTPRHLWRFCTCHQPDQG*MGDPPCQSDSLPRLCEAFADLFYK 227
Query: 1831 SRATPYSS**EPDGGCSGYFVFDVPSKSLE*GAIGPYQTSRETRSCIRH*RSC*WQ 1886
S A PYSS* EP+G CSGY + V SK LE* A T +T +C+ C W+
Sbjct: 226 S*AAPYSS*REPNG*CSGYSILHVSSKPLE*CANNQSATP*KTFTCV-----CYWE 74
>AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thaliana chromosome
II BAC F26H6; putative retroelement pol polyprotein,
partial (1%)
Length = 488
Score = 49.7 bits (117), Expect = 1e-05
Identities = 55/192 (28%), Positives = 80/192 (41%), Gaps = 4/192 (2%)
Frame = +2
Query: 905 FK*WTFRHHIVAYWADPGFMRLEQ*RLHCIKS*SS*GMGS**L*MVKRLC**AICRLSLS 964
F+ W I W D G+M L +* L +S*+ * M
Sbjct: 8 FRSWISMLRIAVSWEDLGYMTLAR*PLPYTRS*NL*RMD--------------------- 124
Query: 965 LGLIVLKGRLFRDLPWKKRVPGRVRPLFLL*RMLRK*YKLEGLQAGES*LNFQKTNAEKG 1024
+L+ LF+ WK + P R+ +LL*R R+ ++ + L G S* N + + K
Sbjct: 125 ----LLRELLFKGCLWKVQSPRRLELQWLL*RTPRRLFRRDRLPTGAS*SNSVRISERKV 292
Query: 1025 WVSSHQLICPRQRPSSSR*GVLFIVLGSSMQSPK----MILKECHEAL*HKEGLAAIGLL 1080
S + PR+ LFIV G S+ K + L C L*++E L IG+
Sbjct: 293 SDSPQHRVSPRE---------LFIVPGLSIHLLKRWLDLFLGPC---L*YQEALRKIGMP 436
Query: 1081 *MFLLLLICPSN 1092
MF +CP N
Sbjct: 437 LMFPQSCMCPXN 472
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 40.0 bits (92), Expect = 0.010
Identities = 20/52 (38%), Positives = 30/52 (57%)
Frame = -3
Query: 1267 VSAGQAEIKKNPS*YGPQD*RGSAETDRCRFPRYIKIPSMASQHSACSEEGW 1318
+S+ Q EI+K+ S YG QD* ++ D C F + M Q+ C++EGW
Sbjct: 157 MSSRQVEIEKDSSRYGSQD*E*GSKAD*CGFSHDS*VS*MGCQYCTCAKEGW 2
>TC93087 similar to GP|20160511|dbj|BAB89462. hypothetical protein {Oryza
sativa (japonica cultivar-group)}, partial (5%)
Length = 668
Score = 36.2 bits (82), Expect = 0.15
Identities = 33/94 (35%), Positives = 47/94 (49%), Gaps = 8/94 (8%)
Frame = +3
Query: 24 LRL*LKLRLMPKPKHLLHLLS-ELRSKHLP-------LLSLNGQYVPTLRHTLLHNVPRL 75
L L L +P+ HL HLL+ +L + LP LL+L + R LL ++ L
Sbjct: 315 LELQLNALRLPRQNHLCHLLALQLNALRLPRQNLLHHLLALQLNVLRLPRQNLLRHLLAL 494
Query: 76 GSHLLLLAKYSVPLLVKLRCLLLNMQLMFLRLLR 109
S++L L + + LRC LL +QL LRL R
Sbjct: 495 QSNVLRLPRQN------LRCHLLVLQLNVLRLPR 578
>TC93584 knotted class I homeodomain KNOX
Length = 1161
Score = 32.7 bits (73), Expect = 1.7
Identities = 20/76 (26%), Positives = 34/76 (44%), Gaps = 9/76 (11%)
Frame = +1
Query: 398 LLPMLFNHQIVSLDSHNILNKILNNNIPNQHTSNAHINN---------NPINNNSLDHKK 448
L P + +V+ HN + I +NN N + +N + NN N NNN+ +
Sbjct: 76 LCPPMMMMPLVTSSHHNAHHPINSNNNNNNNNNNTNANNTTGLFLPIPNSTNNNNNHYTN 255
Query: 449 CKLTQSLLLTQSYSQD 464
C S ++ Q+ Q+
Sbjct: 256 CNNNTSSIMLQNNHQN 303
>TC87476 homologue to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (48%)
Length = 939
Score = 32.3 bits (72), Expect = 2.2
Identities = 32/104 (30%), Positives = 39/104 (36%), Gaps = 7/104 (6%)
Frame = +1
Query: 36 PKHLLHLLSELRSKHLPLLSLNGQYVPTLRHTLLHNVPRLGSHLLLLAKYSVPLLVKLRC 95
P HLLH L + HL LL L H LLH L HL L
Sbjct: 463 PHHLLHPLHPMSISHLHLLQL---------HHLLHTTTNLHLHL-------------LHH 576
Query: 96 LLLNMQLMFLRLLRQFLKL-------L*HTQHLRFMLSRKITNP 132
LL +M + LLR +L L H HL ++ T+P
Sbjct: 577 LLHHMAISHRHLLRLHHRLLIFTSLHLHHLHHLLHIIPTSTTHP 708
>AW586676 similar to GP|806720|gb|AA arabinogalactan-protein precursor
{Nicotiana alata}, partial (22%)
Length = 590
Score = 31.6 bits (70), Expect = 3.7
Identities = 29/73 (39%), Positives = 34/73 (45%), Gaps = 1/73 (1%)
Frame = +3
Query: 20 HKPRLRL*LKLRLMPKPKHLLHLLSEL-RSKHLPLLSLNGQYVPTLRHTLLHNVPRLGSH 78
H P L LR +P H HLL L R+ HL L + TL H LL P L
Sbjct: 99 HLPPYPLLPHLRSLPPLTHRRHLLPYL*RTLHLLHLQPS-----TLHHLLLLLHPTLPLS 263
Query: 79 LLLLAKYSVPLLV 91
LLLL + PLL+
Sbjct: 264 LLLLLHHPFPLLL 302
>TC93861 similar to GP|15451102|gb|AAK96822.1 Unknown protein {Arabidopsis
thaliana}, partial (35%)
Length = 913
Score = 31.6 bits (70), Expect = 3.7
Identities = 28/74 (37%), Positives = 37/74 (49%), Gaps = 6/74 (8%)
Frame = +2
Query: 33 MPKPKHLLHLLSELRSKHLPLLSLNGQ------YVPTLRHTLLHNVPRLGSHLLLLAKYS 86
M + + L HL +L K LP+ L+GQ +V L L V + S L L +S
Sbjct: 233 MHRKQDLAHLCMKLPRKGLPMNLLSGQQ*L*MMFVMILWLDFL-TVKQQSSQLQRLL*WS 409
Query: 87 VPLLVKLRCLLLNM 100
P LVKL LLLN+
Sbjct: 410 WPKLVKLTSLLLNL 451
>TC88000 weakly similar to GP|13486661|dbj|BAB39898. contains EST
AU081362(R10239)~unknown protein {Oryza sativa (japonica
cultivar-group)}, partial (20%)
Length = 1096
Score = 31.2 bits (69), Expect = 4.8
Identities = 19/60 (31%), Positives = 29/60 (47%)
Frame = +2
Query: 39 LLHLLSELRSKHLPLLSLNGQYVPTLRHTLLHNVPRLGSHLLLLAKYSVPLLVKLRCLLL 98
LLH+L + HLP L G + H L+ ++ L LL Y++ LV+L L+
Sbjct: 500 LLHVLPQTHHHHLPKLLQEGGVF*VMVHLLISQEKKMKMLLHLLQHYALVFLVQLALRLI 679
>TC76311 homologue to GP|3204132|emb|CAA07235.1 extensin {Cicer arietinum},
partial (77%)
Length = 973
Score = 31.2 bits (69), Expect = 4.8
Identities = 28/87 (32%), Positives = 43/87 (49%)
Frame = +3
Query: 51 LPLLSLNGQYVPTLRHTLLHNVPRLGSHLLLLAKYSVPLLVKLRCLLLNMQLMFLRLLRQ 110
LPL L+ +P H LLH++P +HL ++ + LL L L++ L L +
Sbjct: 9 LPLPLLHTIIIPHHHHRLLHHLPTTTNHLHHHPRH-LHLLTITSHLHLHLHLHHLLIT-- 179
Query: 111 FLKLL*HTQHLRFMLSRKITNPFSIQR 137
+K+L H HL ++ TNP QR
Sbjct: 180 -IKVLLH--HLPLLILPTTTNPLHHQR 251
>TC92813
Length = 784
Score = 30.8 bits (68), Expect = 6.3
Identities = 17/42 (40%), Positives = 23/42 (54%)
Frame = +1
Query: 38 HLLHLLSELRSKHLPLLSLNGQYVPTLRHTLLHNVPRLGSHL 79
H H + LRSKHLP +S G ++P + LH R +HL
Sbjct: 325 HPFHSMVSLRSKHLPYVSA-GVWIPFIFQKGLHLGRRASNHL 447
>TC91832 similar to GP|13925771|gb|AAK49438.1 phytase {Glycine max}, partial
(49%)
Length = 1204
Score = 30.4 bits (67), Expect = 8.3
Identities = 19/50 (38%), Positives = 30/50 (60%)
Frame = +1
Query: 66 HTLLHNVPRLGSHLLLLAKYSVPLLVKLRCLLLNMQLMFLRLLRQFLKLL 115
HT+ H + + S ++L+ Y + +LV L C+LL M L+ + L FL LL
Sbjct: 895 HTIRHPLSII*SAIILILFYWLEMLVMLTCILL-MALVQIATLVHFLILL 1041
>TC77059 similar to GP|18958499|gb|AAL82617.1 elongation factor 1-gamma
{Glycine max}, partial (96%)
Length = 1582
Score = 30.4 bits (67), Expect = 8.3
Identities = 27/81 (33%), Positives = 42/81 (51%), Gaps = 2/81 (2%)
Frame = +3
Query: 42 LLSELRSKHLPLLSLNGQYVPTL-RHTLLHNVPRLGSHLLLLAKYSVPLLVKLRCLLLNM 100
LL L + +PLL++ +V T+ LL P+L S L+ L +P+L C L++
Sbjct: 261 LLMVLSLRVMPLLAMLPDWVTTICLDHLLFIRPKLTSGLIFLLWRLMPIL*SCTCQDLDL 440
Query: 101 QLMFLRLLR-QFLKLL*HTQH 120
FL+ R QFL+ H +H
Sbjct: 441 PPTFLQ*RRMQFLRSREHLRH 503
>CA860196 weakly similar to GP|1654183|gb|A MYOD-like DNA-binding protein
{Trichinella spiralis}, partial (7%)
Length = 398
Score = 30.4 bits (67), Expect = 8.3
Identities = 18/46 (39%), Positives = 24/46 (52%), Gaps = 4/46 (8%)
Frame = +2
Query: 405 HQIVSLDSHNI----LNKILNNNIPNQHTSNAHINNNPINNNSLDH 446
H V + +NI LN ++NNN+ N H N + N IN N DH
Sbjct: 59 HLTVKVKVNNIIMEDLNNMVNNNMFNSHMVNNNSIVNNINLNRTDH 196
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.358 0.158 0.577
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 78,786,859
Number of Sequences: 36976
Number of extensions: 1232529
Number of successful extensions: 14377
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 4367
Number of HSP's successfully gapped in prelim test: 763
Number of HSP's that attempted gapping in prelim test: 9480
Number of HSP's gapped (non-prelim): 6275
length of query: 2301
length of database: 9,014,727
effective HSP length: 112
effective length of query: 2189
effective length of database: 4,873,415
effective search space: 10667905435
effective search space used: 10667905435
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 66 (30.0 bits)
Medicago: description of AC144656.5


