

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144645.10 - phase: 0 /pseudo
(70 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
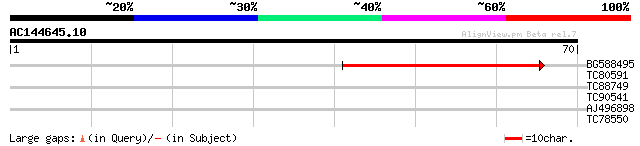
Score E
Sequences producing significant alignments: (bits) Value
BG588495 42 6e-05
TC80591 36 0.003
TC88749 29 0.30
TC90541 homologue to GP|11862963|dbj|BAB19344. putative curly le... 28 0.88
AJ496898 similar to GP|23171295|gb CG3731-PA {Drosophila melanog... 25 5.7
TC78550 weakly similar to GP|21689669|gb|AAM67456.1 unknown prot... 24 9.8
>BG588495
Length = 540
Score = 41.6 bits (96), Expect = 6e-05
Identities = 19/25 (76%), Positives = 22/25 (88%)
Frame = +1
Query: 42 LSRKIEKNCLNGHVVKQIRWDLQLL 66
LSRKIE NCL+G V+KQIR DLQL+
Sbjct: 13 LSRKIEMNCLSGRVIKQIRRDLQLM 87
>TC80591
Length = 734
Score = 35.8 bits (81), Expect = 0.003
Identities = 17/20 (85%), Positives = 17/20 (85%)
Frame = +1
Query: 1 VEAKPNTDGASVHVEASKYV 20
VEAK N DGA VHVEASKYV
Sbjct: 664 VEAKSNIDGAFVHVEASKYV 723
>TC88749
Length = 1221
Score = 29.3 bits (64), Expect = 0.30
Identities = 16/33 (48%), Positives = 21/33 (63%)
Frame = +3
Query: 37 FFQTTLSRKIEKNCLNGHVVKQIRWDLQLLHKD 69
++ T L KI+K CL + KQI DLQL+ KD
Sbjct: 531 YYTTKLHGKIKKTCLVRYGDKQIGPDLQLV*KD 629
>TC90541 homologue to GP|11862963|dbj|BAB19344. putative curly leaf protein
{Oryza sativa (japonica cultivar-group)}, partial (36%)
Length = 788
Score = 27.7 bits (60), Expect = 0.88
Identities = 17/59 (28%), Positives = 30/59 (50%), Gaps = 9/59 (15%)
Frame = +2
Query: 10 ASVHVEASKYVDSSVL-----PLQAHEVDTARF----FQTTLSRKIEKNCLNGHVVKQI 59
+++HV+A++ V+ SVL P VD R F+ + R++ +N H + QI
Sbjct: 101 STIHVDANQLVERSVLVF*MEPAVRSTVDAPRVVKIDFEAVIVRRVNAEVVNVHALLQI 277
>AJ496898 similar to GP|23171295|gb CG3731-PA {Drosophila melanogaster},
partial (35%)
Length = 698
Score = 25.0 bits (53), Expect = 5.7
Identities = 18/64 (28%), Positives = 25/64 (38%), Gaps = 9/64 (14%)
Frame = -1
Query: 13 HVEASKYVDSSVLPLQAHEVDTARFFQTTLSRKIEKNCL---------NGHVVKQIRWDL 63
H+ SK S +PL+ + F SR + CL NGH Q +W L
Sbjct: 320 HLVLSKKCTSFSMPLECSRPYSQLSFHLKQSRLPKGPCLCSRY*MIGKNGHNYPQQQWQL 141
Query: 64 QLLH 67
+H
Sbjct: 140 SQMH 129
>TC78550 weakly similar to GP|21689669|gb|AAM67456.1 unknown protein
{Arabidopsis thaliana}, partial (69%)
Length = 1486
Score = 24.3 bits (51), Expect = 9.8
Identities = 15/51 (29%), Positives = 29/51 (56%), Gaps = 1/51 (1%)
Frame = -1
Query: 2 EAKPNTDGASVHVEASKYVDSSVLPLQAHEV-DTARFFQTTLSRKIEKNCL 51
EA+P+++ ++ +SKY ++S L LQ + D + ++ L + I CL
Sbjct: 847 EARPSSEV*YQYLHSSKYPEASNLHLQHRTIPDWLNYSESKLKQHIML*CL 695
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.130 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,143,190
Number of Sequences: 36976
Number of extensions: 20912
Number of successful extensions: 90
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 90
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 90
length of query: 70
length of database: 9,014,727
effective HSP length: 46
effective length of query: 24
effective length of database: 7,313,831
effective search space: 175531944
effective search space used: 175531944
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 51 (24.3 bits)
Medicago: description of AC144645.10


