

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144617.6 - phase: 0
(178 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
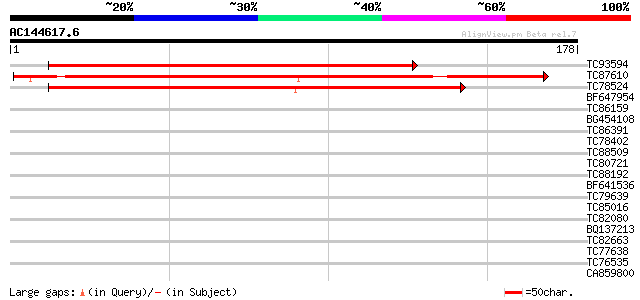
Score E
Sequences producing significant alignments: (bits) Value
TC93594 weakly similar to GP|20466452|gb|AAM20543.1 unknown prot... 168 9e-43
TC87610 similar to GP|9294102|dbj|BAB01954.1 contains similarity... 148 1e-36
TC78524 similar to PIR|D84561|D84561 probable AAA-type ATPase [i... 99 9e-22
BF647954 homologue to GP|11094192|db 26S proteasome regulatory p... 37 0.005
TC86159 homologue to GP|21387177|gb|AAM47992.1 26S proteasome AA... 35 0.012
BG454108 similar to GP|9759050|dbj| AAA-type ATPase-like protein... 33 0.060
TC86391 similar to GP|4097571|gb|AAD09514.1| GMFP5 {Glycine max}... 30 0.66
TC78402 weakly similar to PIR|B84853|B84853 hypothetical protein... 29 0.86
TC88509 similar to PIR|T01826|T01826 microfibril-associated prot... 29 1.1
TC80721 similar to GP|21617902|gb|AAM66952.1 unknown {Arabidopsi... 28 2.5
TC88192 weakly similar to PIR|T05682|T05682 monooxygenase (EC 1.... 27 3.3
BF641536 27 3.3
TC79639 similar to GP|10728064|gb|AAF50455.2 CG7060 gene product... 27 4.3
TC85016 27 4.3
TC82080 similar to GP|9757854|dbj|BAB08488.1 contains similarity... 27 4.3
BQ137213 27 4.3
TC82663 similar to PIR|T51505|T51505 hypothetical protein F5E19_... 27 5.6
TC77638 similar to GP|20161455|dbj|BAB90379. ankyrin-like protei... 27 5.6
TC76535 similar to SP|Q9BBN6|YCF1_LOTJA Hypothetical 214.8 kDa p... 26 7.3
CA859800 weakly similar to GP|17473819|gb| putative protein {Ara... 26 7.3
>TC93594 weakly similar to GP|20466452|gb|AAM20543.1 unknown protein
{Arabidopsis thaliana}, partial (40%)
Length = 754
Score = 168 bits (426), Expect = 9e-43
Identities = 89/116 (76%), Positives = 93/116 (79%)
Frame = +1
Query: 13 KNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELPYCC 72
+ NK SQ+TL GLLNFIDGIWSASTGERLIIFTTNY EKLD ALI RGRMDM IEL Y
Sbjct: 406 EKNKASQVTLSGLLNFIDGIWSASTGERLIIFTTNYVEKLDQALIRRGRMDMHIELSYWG 585
Query: 73 FDGFKMLATKYLSLESHFLFDKIACLLVETNMTPADVAENLMPKVDNEDVATPLLR 128
FDGFKMLA YLS+ESH LF+ I LL ETNMTPADVAENLMPKV EDV L R
Sbjct: 586 FDGFKMLAMNYLSIESHPLFETIQRLLEETNMTPADVAENLMPKVAEEDVEASLER 753
>TC87610 similar to GP|9294102|dbj|BAB01954.1 contains similarity to AAA-type
ATPase~gene_id:T20D4.3 {Arabidopsis thaliana}, partial
(53%)
Length = 1765
Score = 148 bits (374), Expect = 1e-36
Identities = 87/171 (50%), Positives = 116/171 (66%), Gaps = 3/171 (1%)
Frame = +3
Query: 2 EKKE--SQAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICR 59
EKK+ +AE KN S++TL GLLNFIDGIWSA ER+IIFTTN+ +KLD ALI R
Sbjct: 1044 EKKDPIKKAEKEEKNE--SKVTLSGLLNFIDGIWSACGSERIIIFTTNFVDKLDPALIRR 1217
Query: 60 GRMDMLIELPYCCFDGFKMLATKYLSLESH-FLFDKIACLLVETNMTPADVAENLMPKVD 118
GRMD IE+ YC + FK+LA YL +E+H LF I LL ETNMTPADVAENLMPK
Sbjct: 1218 GRMDKHIEMSYCSYQAFKVLARNYLDVETHDDLFPIIEKLLGETNMTPADVAENLMPKSI 1397
Query: 119 NEDVATPLLRLIQALRSIEEEAEKEEGTSAKQESDGEDSSAEKKEDAEMVT 169
ED + L LIQ+L E A+K++ AK++ + +++ + +++ + +T
Sbjct: 1398 TEDFESCLKNLIQSL----EIAKKKDEEEAKKKIEDQEAKLKAQKEKQELT 1538
>TC78524 similar to PIR|D84561|D84561 probable AAA-type ATPase [imported] -
Arabidopsis thaliana, partial (15%)
Length = 1697
Score = 99.0 bits (245), Expect = 9e-22
Identities = 54/133 (40%), Positives = 82/133 (61%), Gaps = 2/133 (1%)
Frame = +3
Query: 13 KNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELPYCC 72
KN + TL GLLN++DG+WS+ ER++IFTTN+ +K+D AL+ GRMDM I L +
Sbjct: 1065 KNMPPRKFTLSGLLNYMDGLWSSCGEERILIFTTNHKDKVDPALLRPGRMDMHIHLSFLK 1244
Query: 73 FDGFKMLATKYLSLES--HFLFDKIACLLVETNMTPADVAENLMPKVDNEDVATPLLRLI 130
F++LA YL +E H LF++I LL + +++PA VAE L+ D + L++ +
Sbjct: 1245 AKAFRILAANYLDIEGNHHSLFEQIEELLEKVDVSPAVVAEYLLRSEDPDVALGALVKFL 1424
Query: 131 QALRSIEEEAEKE 143
Q + EE +E
Sbjct: 1425 QDQEIVNEETSQE 1463
>BF647954 homologue to GP|11094192|db 26S proteasome regulatory particle
triple-A ATPase subunit4 {Oryza sativa (japonica
cultivar-group)}, partial (53%)
Length = 649
Score = 36.6 bits (83), Expect = 0.005
Identities = 24/58 (41%), Positives = 32/58 (54%)
Frame = +2
Query: 12 TKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELP 69
T ++ Q TL LLN +DG G+ II TN + LD AL+ GR+D IE+P
Sbjct: 281 TSADREIQRTLMELLNQLDGF--DQLGKVKIIMATNRPDVLDPALLRPGRLDRKIEIP 448
>TC86159 homologue to GP|21387177|gb|AAM47992.1 26S proteasome AAA-ATPase
subunit RPT4a-like protein {Arabidopsis thaliana},
complete
Length = 1640
Score = 35.4 bits (80), Expect = 0.012
Identities = 23/58 (39%), Positives = 32/58 (54%)
Frame = +2
Query: 12 TKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELP 69
T ++ Q TL LLN +DG G+ +I TN + LD AL+ GR+D IE+P
Sbjct: 827 TSADREIQRTLMELLNQLDGF--DQLGKVKMIMATNRPDVLDPALLRPGRLDRKIEIP 994
>BG454108 similar to GP|9759050|dbj| AAA-type ATPase-like protein
{Arabidopsis thaliana}, partial (8%)
Length = 519
Score = 33.1 bits (74), Expect = 0.060
Identities = 21/79 (26%), Positives = 38/79 (47%), Gaps = 11/79 (13%)
Frame = +1
Query: 92 FDKIACLLVETNMTPADVAENLMPKVDNEDVATPLLRLIQALR-----------SIEEEA 140
F +I L+ + +TPA VAE LM D E ++L++ + IE+++
Sbjct: 34 FGEIEGLIEDIQITPAQVAEELMKNEDAEATLEGFVKLLKRKKMEGDVCENNNNKIEQQS 213
Query: 141 EKEEGTSAKQESDGEDSSA 159
+K + KQ+ G +S +
Sbjct: 214 KKRKVVGCKQKRGGGNSKS 270
>TC86391 similar to GP|4097571|gb|AAD09514.1| GMFP5 {Glycine max}, partial
(54%)
Length = 1353
Score = 29.6 bits (65), Expect = 0.66
Identities = 20/66 (30%), Positives = 31/66 (46%), Gaps = 1/66 (1%)
Frame = +1
Query: 100 VETNMTPADVAENLMPKVD-NEDVATPLLRLIQALRSIEEEAEKEEGTSAKQESDGEDSS 158
V+ M P D+ E L K+ N D+ P EEE ++++G K+E ++
Sbjct: 766 VKGTMEPKDLIEYLKEKLKRNVDIVPP---------KKEEEKKEKDGGGEKKEKKEDEKK 918
Query: 159 AEKKED 164
EKK D
Sbjct: 919 EEKKVD 936
>TC78402 weakly similar to PIR|B84853|B84853 hypothetical protein At2g42370
[imported] - Arabidopsis thaliana, partial (14%)
Length = 1982
Score = 29.3 bits (64), Expect = 0.86
Identities = 13/33 (39%), Positives = 19/33 (57%)
Frame = +2
Query: 136 IEEEAEKEEGTSAKQESDGEDSSAEKKEDAEMV 168
+E E E EEG +E +GE E++ED +V
Sbjct: 425 VEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVV 523
>TC88509 similar to PIR|T01826|T01826 microfibril-associated protein homolog
T15F16.8 - Arabidopsis thaliana, partial (83%)
Length = 1306
Score = 28.9 bits (63), Expect = 1.1
Identities = 19/50 (38%), Positives = 33/50 (66%), Gaps = 3/50 (6%)
Frame = +1
Query: 116 KVDN-EDVATPLLRLIQA--LRSIEEEAEKEEGTSAKQESDGEDSSAEKK 162
+VDN E+V R+ QA + +IEEEA+++EG +++ ED+ AE++
Sbjct: 304 RVDNREEVREDHRRIRQAEIVSTIEEEAKRQEGLDLEEQD--EDAMAERR 447
Score = 26.6 bits (57), Expect = 5.6
Identities = 14/32 (43%), Positives = 18/32 (55%)
Frame = +1
Query: 137 EEEAEKEEGTSAKQESDGEDSSAEKKEDAEMV 168
EEE E+EE ++ESD E S E+ MV
Sbjct: 502 EEEEEEEEEEEEEEESDYETDSDEEYTGVAMV 597
>TC80721 similar to GP|21617902|gb|AAM66952.1 unknown {Arabidopsis
thaliana}, partial (49%)
Length = 1094
Score = 27.7 bits (60), Expect = 2.5
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = -1
Query: 125 PLLRLIQALRSIEEEAEKEEGTSAKQESDGED 156
P LRL + E AE+E+G S +E G D
Sbjct: 458 PELRLFETFSPAAEGAEEEDGDSTGEEKTGSD 363
>TC88192 weakly similar to PIR|T05682|T05682 monooxygenase (EC 1.-.-.-) 2 -
Arabidopsis thaliana, partial (61%)
Length = 1557
Score = 27.3 bits (59), Expect = 3.3
Identities = 13/36 (36%), Positives = 18/36 (49%)
Frame = -3
Query: 57 ICRGRMDMLIELPYCCFDGFKMLATKYLSLESHFLF 92
+C R+D + Y CFDG + Y ES FL+
Sbjct: 595 LCNHRVDSITSYQYLCFDGCSINKM*YFK-ESRFLY 491
>BF641536
Length = 645
Score = 27.3 bits (59), Expect = 3.3
Identities = 13/27 (48%), Positives = 18/27 (66%)
Frame = -2
Query: 118 DNEDVATPLLRLIQALRSIEEEAEKEE 144
+ E+ PLL+L Q+ IEEE E+EE
Sbjct: 185 EGEEDVDPLLKLNQSKSCIEEEEEEEE 105
>TC79639 similar to GP|10728064|gb|AAF50455.2 CG7060 gene product
{Drosophila melanogaster}, partial (2%)
Length = 757
Score = 26.9 bits (58), Expect = 4.3
Identities = 18/68 (26%), Positives = 27/68 (39%), Gaps = 10/68 (14%)
Frame = +1
Query: 109 VAENLMPKVDNEDVATPLLRLIQALRSIEEEAEKE----------EGTSAKQESDGEDSS 158
VAEN K +TP ++ + ++IE+ E G S +QE G
Sbjct: 97 VAENNKKKTKKPQTSTPTSKIKKDNKNIEKSKENNNKNKKKLNFTNGNSKEQEESGRKKK 276
Query: 159 AEKKEDAE 166
K +D E
Sbjct: 277 HAKDDDEE 300
>TC85016
Length = 517
Score = 26.9 bits (58), Expect = 4.3
Identities = 16/46 (34%), Positives = 23/46 (49%)
Frame = +1
Query: 53 DHALICRGRMDMLIELPYCCFDGFKMLATKYLSLESHFLFDKIACL 98
D L C + +IELPY K + K+ ++E LFD +CL
Sbjct: 172 DFELFCSSNNETMIELPYKVKLNVKNIDYKHQTIE---LFDPQSCL 300
>TC82080 similar to GP|9757854|dbj|BAB08488.1 contains similarity to
apoptosis antagonizing transcription
factor~gene_id:MFB13.10, partial (37%)
Length = 908
Score = 26.9 bits (58), Expect = 4.3
Identities = 11/39 (28%), Positives = 22/39 (56%)
Frame = +2
Query: 137 EEEAEKEEGTSAKQESDGEDSSAEKKEDAEMVTSSMLRL 175
EEE E +EG+ ++E + ++ S K ++ E + + L
Sbjct: 131 EEEEEDDEGSDEEEEDERQEESRWKDDEMEQLEKEYMDL 247
>BQ137213
Length = 1006
Score = 26.9 bits (58), Expect = 4.3
Identities = 12/36 (33%), Positives = 19/36 (52%)
Frame = +3
Query: 57 ICRGRMDMLIELPYCCFDGFKMLATKYLSLESHFLF 92
I R D +I +PYC F + ++ S+ S+F F
Sbjct: 789 ILRYSCDTIIYIPYCMFVVLRFISLFLFSVSSYFAF 896
>TC82663 similar to PIR|T51505|T51505 hypothetical protein F5E19_70 -
Arabidopsis thaliana, partial (20%)
Length = 914
Score = 26.6 bits (57), Expect = 5.6
Identities = 17/51 (33%), Positives = 23/51 (44%)
Frame = +3
Query: 128 RLIQALRSIEEEAEKEEGTSAKQESDGEDSSAEKKEDAEMVTSSMLRLESY 178
+LI L + EE EK + S + SAE +E E S+ ESY
Sbjct: 345 KLINELDNSREEEEKSKKAMESLASALHEVSAESREAKENFLSTQAERESY 497
>TC77638 similar to GP|20161455|dbj|BAB90379. ankyrin-like protein {Oryza
sativa (japonica cultivar-group)}, partial (62%)
Length = 2345
Score = 26.6 bits (57), Expect = 5.6
Identities = 16/68 (23%), Positives = 34/68 (49%), Gaps = 2/68 (2%)
Frame = +1
Query: 101 ETNMTPADVAENLMPKVDNEDVATPLLRLIQALRSIEEEAEK--EEGTSAKQESDGEDSS 158
++N++ + +E + +ED T E+E +K +EG++ + DGE++S
Sbjct: 580 DSNVSSEEKSEENSTEKSSEDTKT------------EDEGKKTEDEGSNTENNKDGEEAS 723
Query: 159 AEKKEDAE 166
++ E E
Sbjct: 724 TKESESDE 747
>TC76535 similar to SP|Q9BBN6|YCF1_LOTJA Hypothetical 214.8 kDa protein
ycf1. {Lotus japonicus}, partial (16%)
Length = 1658
Score = 26.2 bits (56), Expect = 7.3
Identities = 16/57 (28%), Positives = 25/57 (43%)
Frame = +3
Query: 37 TGERLIIFTTNYAEKLDHALICRGRMDMLIELPYCCFDGFKMLATKYLSLESHFLFD 93
TG+ ++ + YA H + R ++ LPY F+ Y S +SHF D
Sbjct: 417 TGQLMMFISIYYAPL--HLALIRPHTITVLTLPYLFFNFVYKNNKHYYSADSHFYLD 581
>CA859800 weakly similar to GP|17473819|gb| putative protein {Arabidopsis
thaliana}, partial (8%)
Length = 531
Score = 26.2 bits (56), Expect = 7.3
Identities = 14/30 (46%), Positives = 19/30 (62%)
Frame = +2
Query: 134 RSIEEEAEKEEGTSAKQESDGEDSSAEKKE 163
+ +EEE E+EEGT A GE S +KK+
Sbjct: 275 KEVEEENEEEEGTEA---VGGEASKKKKKK 355
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.130 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,488,176
Number of Sequences: 36976
Number of extensions: 57648
Number of successful extensions: 306
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 297
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 305
length of query: 178
length of database: 9,014,727
effective HSP length: 90
effective length of query: 88
effective length of database: 5,686,887
effective search space: 500446056
effective search space used: 500446056
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC144617.6


