

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
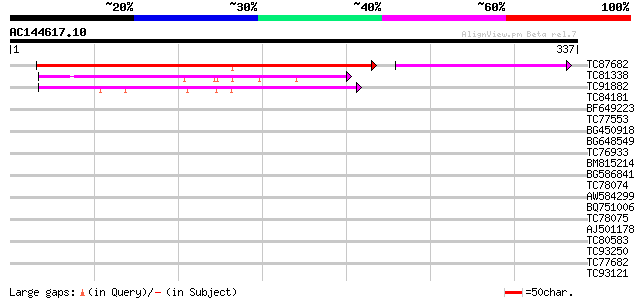
Query= AC144617.10 + phase: 0
(337 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC87682 weakly similar to PIR|T04527|T04527 hypothetical protein... 213 5e-68
TC81338 weakly similar to PIR|B86326|B86326 hypothetical protein... 75 4e-14
TC91882 similar to GP|10176714|dbj|BAB09944. dimethylaniline mon... 62 4e-10
TC84181 similar to SP|Q9SJA7|SOX_ARATH Potential sarcosine oxida... 33 0.12
BF649223 weakly similar to GP|21740857|emb OSJNBb0072N21.6 {Oryz... 33 0.16
TC77553 homologue to GP|1066499|gb|AAB41904.1| NADH-dependent gl... 32 0.35
BG450918 similar to PIR|T00665|T006 hypothetical protein F3I6.28... 30 1.7
BG648549 similar to PIR|E85359|E853 hypothetical protein AT4g307... 29 2.2
TC76933 weakly similar to PIR|JC5703|JC5703 15.5K oleosin - orie... 29 2.2
BM815214 similar to GP|22136992|gb unknown protein {Arabidopsis ... 29 2.9
BG586841 similar to PIR|T07142|T07 beta-glucan-elicitor receptor... 28 3.8
TC78074 similar to GP|20198327|gb|AAB63082.2 putative lipase {Ar... 28 3.8
AW584299 similar to GP|20160694|db hypothetical protein~similar ... 28 3.8
BQ751006 homologue to GP|6069653|dbj| hypothetical protein {Oryz... 28 3.8
TC78075 similar to GP|20198327|gb|AAB63082.2 putative lipase {Ar... 28 3.8
AJ501178 28 5.0
TC80583 squalene monooxygenase 1 [Medicago truncatula] 28 5.0
TC93250 GP|11863369|emb|CAC18791. bA110H4.2 (similar to membrane... 28 6.5
TC77682 phosphate transporter 28 6.5
TC93121 weakly similar to GP|8953375|emb|CAB96648.1 putative pro... 28 6.5
>TC87682 weakly similar to PIR|T04527|T04527 hypothetical protein F16A16.170
- Arabidopsis thaliana, partial (38%)
Length = 1040
Score = 213 bits (543), Expect(2) = 5e-68
Identities = 101/207 (48%), Positives = 144/207 (68%), Gaps = 5/207 (2%)
Frame = +1
Query: 17 IIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFCELPMMSF 76
II+GAG SG+A AACL++Q +P +ILER +C ASLWQN TYDR+ LHL K CELP F
Sbjct: 46 IIIGAGTSGLATAACLTKQSIPFIILERENCFASLWQNYTYDRVHLHLRKQLCELPHFPF 225
Query: 77 PQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEF-DSTSQIWMVRTKEGDF---- 131
P ++P Y K QFI Y+ +Y ++F+I+P +N+ V AE+ D + W V+ +
Sbjct: 226 PPSYPHYVPKKQFIEYLGNYVNNFNINPIYNRAVELAEYVDDDEKKWRVKAENKSSGEVE 405
Query: 132 QYFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLVIGCGNSGME 191
+Y + +L+VA+GE AEP P + G+E+F G V+H++ YK+G E+K++ VLV+G GNSGME
Sbjct: 406 EYSARFLVVASGETAEPRVPVVEGLENFKGKVIHSTRYKNGKEFKDEHVLVVGSGNSGME 585
Query: 192 VSLDLCRHNAMPHLVARNSVHILPRDM 218
++LDL A P ++ R+ VHIL RDM
Sbjct: 586 IALDLANFGAKPSIIVRSPVHILSRDM 666
Score = 62.4 bits (150), Expect(2) = 5e-68
Identities = 33/106 (31%), Positives = 55/106 (51%), Gaps = 1/106 (0%)
Frame = +2
Query: 230 LYKWLPLKLVDKFLLLVSSFFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKS 289
L +L V+K +++ S G+ + YGI P GP +K+ GK PV+DVG + +IKS
Sbjct: 686 LLNYLSPSTVEKLVVIASRIVYGDLSKYGIPFPSEGPFTMKMKYGKFPVIDVGTVKKIKS 865
Query: 290 GNIKVMEG-VKEITRNGAKFLDGQEKEFDAIILATGYKSNVPSWLK 334
G I+V+ ++ I+ N F DG + A ++ + + K
Sbjct: 866 GEIQVLPAEIESISGNQVLFRDGNPTRLTPLFSALVFRDQLKNGFK 1003
>TC81338 weakly similar to PIR|B86326|B86326 hypothetical protein AAF82235.1
[imported] - Arabidopsis thaliana, partial (32%)
Length = 784
Score = 75.1 bits (183), Expect = 4e-14
Identities = 61/222 (27%), Positives = 102/222 (45%), Gaps = 36/222 (16%)
Frame = +2
Query: 18 IVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFCELPMMSFP 77
I+GAG SG+AVA LS ++ E +D I +W++ +Y KL E +P
Sbjct: 77 IIGAGVSGLAVAKQLSHYN--PIVFEATDSIGGVWRHCSYRCTKLQSQTWNYEFSDFPWP 250
Query: 78 QTFPK-YPTKHQFISYMESYADHFHI--HPRFNQTVLSAEFDSTSQ-------------- 120
+ K YP+ + + Y+ YA +F + + +FN V+ +F +
Sbjct: 251 ERESKDYPSHAEILEYLHLYAVYFDLFKYVKFNTKVVEIKFVRDKEGFDFGRLPGDHGNP 430
Query: 121 -----IW--MVRTKEGDF--QYFSPWLIVATGENAE----PVFPTIHGMEHFHGPVVHTS 167
+W V+T E D +Y +++V TG+ + P FP G E F G V+HT
Sbjct: 431 LPERPVWELSVQTNESDAIQKYCFEFVVVCTGKYGDIPLMPTFPCNKGPEVFKGKVLHTI 610
Query: 168 DY------KSGSEYKNKKVLVIGCGNSGMEVSLDLCRHNAMP 203
DY + K KKV+V+G S ++++++ + N P
Sbjct: 611 DYCKLDKEATSDLVKGKKVVVVGYKKSAIDLTMECVQANQRP 736
>TC91882 similar to GP|10176714|dbj|BAB09944. dimethylaniline
monooxygenase-like protein {Arabidopsis thaliana},
partial (47%)
Length = 800
Score = 61.6 bits (148), Expect = 4e-10
Identities = 48/229 (20%), Positives = 102/229 (43%), Gaps = 37/229 (16%)
Frame = +2
Query: 18 IVGAGPSGIAVAACLSEQGVPSLILERSDCIASLW---------------------QNRT 56
++GAGP+G+ A + ++G ++LE++ I W +
Sbjct: 68 VIGAGPAGLVAAREIRKEGHKVIVLEQNHDIGGQWLYDDKNIEGEDPLGRNPFLKVHSSI 247
Query: 57 YDRLKLHLPKH---FCELPMMSFP-QTFPKYPTKHQFISYMESYADHFHIHP--RFNQTV 110
Y+ L+ P+ F + P + + ++P + + Y++ + + F + RFN V
Sbjct: 248 YNSLRTQSPREVMGFTDFPFSTKKGRDMRRFPGHVEILMYLKDFCEWFGLREMIRFNTRV 427
Query: 111 LSAEFDSTSQI------WMVRTKEGD----FQYFSPWLIVATGENAEPVFPTIHGMEHFH 160
+ W+VR+KE + + ++VATG +P P+I GM+ +
Sbjct: 428 NYVGMLDDYGVCGNNLKWIVRSKEKNSDKVVEEVFDAVVVATGHYCQPKLPSIKGMDTWK 607
Query: 161 GPVVHTSDYKSGSEYKNKKVLVIGCGNSGMEVSLDLCRHNAMPHLVARN 209
+H+ Y++ + N+ V+V+G SG +++++L H+ +R+
Sbjct: 608 RKQMHSHIYRTPEPFHNEVVVVVGNSFSGQDIAIELVGVAKEVHISSRS 754
>TC84181 similar to SP|Q9SJA7|SOX_ARATH Potential sarcosine oxidase (EC
1.5.3.1). [Mouse-ear cress] {Arabidopsis thaliana},
partial (20%)
Length = 761
Score = 33.5 bits (75), Expect = 0.12
Identities = 28/100 (28%), Positives = 53/100 (53%), Gaps = 4/100 (4%)
Frame = +3
Query: 17 IIVGAGPSGIAVAACLSEQGVPSLILERSDCI----ASLWQNRTYDRLKLHLPKHFCELP 72
IIVGAG G + A +++G+ +L+LE+ D + +S ++RT + +H+C++
Sbjct: 63 IIVGAGVMGSSTAYYAAKRGLKTLVLEQFDFLHHRGSSHGESRTIH--AAYPQQHYCQMV 236
Query: 73 MMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLS 112
M S+ Q + + + F Y + A HF + + +LS
Sbjct: 237 MKSY-QLWEEAQAQAGFNVYYK--AHHFDMGSSNDPMILS 347
>BF649223 weakly similar to GP|21740857|emb OSJNBb0072N21.6 {Oryza sativa},
partial (11%)
Length = 278
Score = 33.1 bits (74), Expect = 0.16
Identities = 17/49 (34%), Positives = 27/49 (54%)
Frame = +2
Query: 18 IVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPK 66
IVGAG SG+ +++ G ++ E D + LW++ T + KL PK
Sbjct: 83 IVGAGISGLLACKYVTQIGFNPIVFEADDGVGGLWRH-TIESTKLQNPK 226
>TC77553 homologue to GP|1066499|gb|AAB41904.1| NADH-dependent glutamate
synthase {Medicago sativa}, partial (37%)
Length = 3067
Score = 32.0 bits (71), Expect = 0.35
Identities = 16/34 (47%), Positives = 22/34 (64%)
Frame = +1
Query: 18 IVGAGPSGIAVAACLSEQGVPSLILERSDCIASL 51
IVG+GPSG+A A L++ G + ER+D I L
Sbjct: 1387 IVGSGPSGLAAADQLNKMGHIVTVFERADRIGGL 1488
>BG450918 similar to PIR|T00665|T006 hypothetical protein F3I6.28 -
Arabidopsis thaliana, partial (6%)
Length = 646
Score = 29.6 bits (65), Expect = 1.7
Identities = 11/29 (37%), Positives = 21/29 (71%)
Frame = +3
Query: 17 IIVGAGPSGIAVAACLSEQGVPSLILERS 45
+I+GAGP G+ ++ L++ G+ +LER+
Sbjct: 366 LIIGAGPVGLVLSILLTKLGINCTVLERN 452
>BG648549 similar to PIR|E85359|E853 hypothetical protein AT4g30720
[imported] - Arabidopsis thaliana, partial (7%)
Length = 736
Score = 29.3 bits (64), Expect = 2.2
Identities = 13/27 (48%), Positives = 18/27 (66%)
Frame = +2
Query: 18 IVGAGPSGIAVAACLSEQGVPSLILER 44
IVG+GPSG+ A L+E G ++ER
Sbjct: 644 IVGSGPSGLFAALVLAELGADVTLIER 724
>TC76933 weakly similar to PIR|JC5703|JC5703 15.5K oleosin - oriental
sesame, partial (60%)
Length = 907
Score = 29.3 bits (64), Expect = 2.2
Identities = 15/32 (46%), Positives = 19/32 (58%)
Frame = -3
Query: 237 KLVDKFLLLVSSFFLGNTNHYGIKRPKTGPIE 268
KL+ FL VSSFFL + +H+G GP E
Sbjct: 659 KLMKIFLRFVSSFFLRHVDHFGPPELYLGPPE 564
>BM815214 similar to GP|22136992|gb unknown protein {Arabidopsis thaliana},
partial (24%)
Length = 596
Score = 28.9 bits (63), Expect = 2.9
Identities = 21/64 (32%), Positives = 33/64 (50%), Gaps = 1/64 (1%)
Frame = +1
Query: 268 ELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEITRNG-AKFLDGQEKEFDAIILATGYK 326
E+ +AT P+L ++ ++ NI +K + +G F DG DAII TGYK
Sbjct: 127 EVHVATKPNPLLKGMKLENVR--NICFHTLIKCVYEDGLVAFEDGFSTYADAIIHCTGYK 300
Query: 327 SNVP 330
++P
Sbjct: 301 YHIP 312
>BG586841 similar to PIR|T07142|T07 beta-glucan-elicitor receptor - soybean,
partial (32%)
Length = 718
Score = 28.5 bits (62), Expect = 3.8
Identities = 26/107 (24%), Positives = 46/107 (42%), Gaps = 6/107 (5%)
Frame = +1
Query: 106 FNQTVLSAEFDSTSQIWMVRTKEGDF-QYFSPWLIVATGENAEPVFPTIHG-----MEHF 159
F+ +LS + S GD +YF P+LI ++ + +PT + F
Sbjct: 94 FSPNLLSTPLPTNSFFQNFVLNNGDAPEYFHPYLIKSSNSSLSVSYPTRSSNSAVISQVF 273
Query: 160 HGPVVHTSDYKSGSEYKNKKVLVIGCGNSGMEVSLDLCRHNAMPHLV 206
H + TS+ ++ + N K ++ + G V+LD+ N LV
Sbjct: 274 HNDLTITSNQQNTKQSSNGKHIISSYSDLG--VTLDIPSSNLSFFLV 408
>TC78074 similar to GP|20198327|gb|AAB63082.2 putative lipase {Arabidopsis
thaliana}, partial (25%)
Length = 735
Score = 28.5 bits (62), Expect = 3.8
Identities = 19/66 (28%), Positives = 30/66 (44%), Gaps = 1/66 (1%)
Frame = +3
Query: 237 KLVDKFLLLVSSFFLG-NTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVM 295
K++ F S+FF G N+NH + + K T K + I Q++S NIK
Sbjct: 72 KILIAFPKSESTFFFGFNSNHLKFQTTNPQTLTTKKLTTKCTSSQINTITQLQSQNIKQS 251
Query: 296 EGVKEI 301
+ E+
Sbjct: 252 SKLTEL 269
>AW584299 similar to GP|20160694|db hypothetical protein~similar to
Arabidopsis thaliana chromosome 3 At3g56840, partial
(22%)
Length = 549
Score = 28.5 bits (62), Expect = 3.8
Identities = 15/39 (38%), Positives = 24/39 (61%)
Frame = +2
Query: 5 KEKVKCVWIHGPIIVGAGPSGIAVAACLSEQGVPSLILE 43
+E+V CV ++GAG GIAVA L+ +G +++E
Sbjct: 137 RERVDCV------VIGAGVVGIAVARALALKGREVIVIE 235
>BQ751006 homologue to GP|6069653|dbj| hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 694
Score = 28.5 bits (62), Expect = 3.8
Identities = 11/25 (44%), Positives = 15/25 (60%)
Frame = -1
Query: 59 RLKLHLPKHFCELPMMSFPQTFPKY 83
RL+ HL +H LP +FP PK+
Sbjct: 565 RLERHLRRHGTSLPQTAFPDVVPKF 491
>TC78075 similar to GP|20198327|gb|AAB63082.2 putative lipase {Arabidopsis
thaliana}, partial (19%)
Length = 637
Score = 28.5 bits (62), Expect = 3.8
Identities = 19/66 (28%), Positives = 30/66 (44%), Gaps = 1/66 (1%)
Frame = +3
Query: 237 KLVDKFLLLVSSFFLG-NTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVM 295
K++ F S+FF G N+NH + + K T K + I Q++S NIK
Sbjct: 12 KILIAFPKSESTFFFGFNSNHLKFQTTNPQTLTTKKLTTKCTSSQINTITQLQSQNIKQS 191
Query: 296 EGVKEI 301
+ E+
Sbjct: 192 SKLTEL 209
>AJ501178
Length = 630
Score = 28.1 bits (61), Expect = 5.0
Identities = 18/51 (35%), Positives = 24/51 (46%)
Frame = +1
Query: 72 PMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIW 122
P++ P F +P+ HQF S + P+ N LS F STS IW
Sbjct: 73 PLLPLPN-FLLFPSNHQFSSSNRT--------PKINIMHLSKLFHSTSSIW 198
>TC80583 squalene monooxygenase 1 [Medicago truncatula]
Length = 1830
Score = 28.1 bits (61), Expect = 5.0
Identities = 15/28 (53%), Positives = 19/28 (67%)
Frame = +3
Query: 17 IIVGAGPSGIAVAACLSEQGVPSLILER 44
IIVGAG +G A+A L + G LI+ER
Sbjct: 288 IIVGAGVAGSALAYTLGKDGRRVLIIER 371
>TC93250 GP|11863369|emb|CAC18791. bA110H4.2 (similar to membrane protein)
{Homo sapiens}, partial (1%)
Length = 761
Score = 27.7 bits (60), Expect = 6.5
Identities = 14/34 (41%), Positives = 17/34 (49%), Gaps = 3/34 (8%)
Frame = +1
Query: 78 QTFPKYPTKHQFISYMESYADHFH---IHPRFNQ 108
Q P YP ++ F SY A FH IHP F +
Sbjct: 379 QRAPLYPPRNFFSSYSSRLASSFHPQPIHPTFRR 480
>TC77682 phosphate transporter
Length = 2245
Score = 27.7 bits (60), Expect = 6.5
Identities = 13/26 (50%), Positives = 17/26 (65%)
Frame = -1
Query: 158 HFHGPVVHTSDYKSGSEYKNKKVLVI 183
HFH PVV +KS +E+KNK +I
Sbjct: 826 HFHSPVVRQCSWKS-TEHKNKSPNII 752
>TC93121 weakly similar to GP|8953375|emb|CAB96648.1 putative protein
{Arabidopsis thaliana}, partial (27%)
Length = 918
Score = 27.7 bits (60), Expect = 6.5
Identities = 13/49 (26%), Positives = 23/49 (46%)
Frame = -1
Query: 149 VFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLVIGCGNSGMEVSLDLC 197
V+P + + H +V + + + NK + +GCG G SL+ C
Sbjct: 396 VYPYLESLHHLLVALVKSMWIQVNGPFWNKTLFHVGCGLLGPNCSLNQC 250
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.139 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,275,972
Number of Sequences: 36976
Number of extensions: 200006
Number of successful extensions: 956
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 944
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 952
length of query: 337
length of database: 9,014,727
effective HSP length: 97
effective length of query: 240
effective length of database: 5,428,055
effective search space: 1302733200
effective search space used: 1302733200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144617.10


