

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
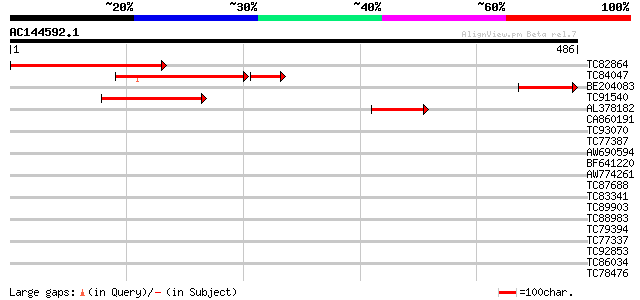
Query= AC144592.1 - phase: 0 /pseudo
(486 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC82864 similar to GP|3695061|gb|AAC62625.1| rac GTPase activati... 271 5e-73
TC84047 homologue to GP|3695063|gb|AAC62626.1| rac GTPase activa... 164 3e-51
BE204083 similar to GP|3695061|gb|A rac GTPase activating protei... 108 5e-24
TC91540 similar to GP|9757821|dbj|BAB08339.1 rac GTPase activati... 97 1e-20
AL378182 similar to GP|3695063|gb|A rac GTPase activating protei... 50 2e-06
CA860191 homologue to GP|19697342|gb hypothetical protein {Dicty... 40 0.002
TC93070 weakly similar to GP|17065230|gb|AAL32769.1 Unknown prot... 38 0.008
TC77387 similar to GP|15983396|gb|AAL11566.1 At1g51440/F5D21_19 ... 38 0.010
AW690594 similar to GP|23498163|emb hypothetical protein {Plasmo... 38 0.010
BF641220 weakly similar to PIR|G86203|G86 probable N-arginine di... 37 0.013
AW774261 weakly similar to EGAD|146423|156 vitellogenin {Anolis ... 37 0.017
TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~un... 37 0.022
TC83341 weakly similar to GP|8978076|dbj|BAA98104.1 gene_id:K14A... 37 0.022
TC89903 similar to GP|15810167|gb|AAL06985.1 At1g64110/F22C12_22... 37 0.022
TC88983 similar to SP|P93231|VP41_LYCES Vacuolar assembly protei... 36 0.029
TC79394 weakly similar to PIR|T49019|T49019 probable RNA binding... 36 0.038
TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Atel... 35 0.065
TC92853 similar to GP|20268748|gb|AAM14077.1 unknown protein {Ar... 35 0.084
TC86034 similar to GP|6815067|dbj|BAA90354.1 ESTs AU082210(C5365... 35 0.084
TC78476 homologue to SP|O22437|CHLD_PEA Magnesium-chelatase subu... 35 0.084
>TC82864 similar to GP|3695061|gb|AAC62625.1| rac GTPase activating protein
2 {Lotus japonicus}, partial (14%)
Length = 583
Score = 271 bits (692), Expect = 5e-73
Identities = 134/134 (100%), Positives = 134/134 (100%)
Frame = +2
Query: 1 MTRLFRSKSCNIAGFSVSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEI 60
MTRLFRSKSCNIAGFSVSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEI
Sbjct: 182 MTRLFRSKSCNIAGFSVSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEI 361
Query: 61 YSSNNASRQFTKGSNNNHQNQFAILDIVMAALKKSIVTCSVEREDVSSLDISWPTEVRHV 120
YSSNNASRQFTKGSNNNHQNQFAILDIVMAALKKSIVTCSVEREDVSSLDISWPTEVRHV
Sbjct: 362 YSSNNASRQFTKGSNNNHQNQFAILDIVMAALKKSIVTCSVEREDVSSLDISWPTEVRHV 541
Query: 121 SHVTFDRFNGFLGL 134
SHVTFDRFNGFLGL
Sbjct: 542 SHVTFDRFNGFLGL 583
>TC84047 homologue to GP|3695063|gb|AAC62626.1| rac GTPase activating
protein 3 {Lotus japonicus}, partial (34%)
Length = 630
Score = 164 bits (416), Expect(2) = 3e-51
Identities = 84/120 (70%), Positives = 97/120 (80%), Gaps = 6/120 (5%)
Frame = +3
Query: 91 ALKKSIVTCSVER-EDVSS-----LDISWPTEVRHVSHVTFDRFNGFLGLPSEFQPEVPT 144
ALKKS+V CSVE +DV S ++I WPT V+HV+HVTFDRFNGFLGLP E + VP
Sbjct: 156 ALKKSMVACSVESPDDVISAVHHPMEIGWPTNVKHVNHVTFDRFNGFLGLPLELEVHVPA 335
Query: 145 RVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQE 204
VPSASV VFGVSA+SMQCSYD +GNSVPTILL MQ++LYS+GGLKAEGIFRI EN +E
Sbjct: 336 PVPSASVSVFGVSAESMQCSYDSKGNSVPTILLLMQERLYSQGGLKAEGIFRINPENGEE 515
Score = 55.8 bits (133), Expect(2) = 3e-51
Identities = 23/30 (76%), Positives = 28/30 (92%)
Frame = +1
Query: 207 VRDQLNKGVVPHGIDVHCLSGLIKAWFREL 236
+R+QLN G+VP+ IDVHCL+GLIKAWFREL
Sbjct: 523 LREQLNSGIVPNDIDVHCLAGLIKAWFREL 612
>BE204083 similar to GP|3695061|gb|A rac GTPase activating protein 2 {Lotus
japonicus}, partial (12%)
Length = 468
Score = 108 bits (270), Expect = 5e-24
Identities = 50/50 (100%), Positives = 50/50 (100%)
Frame = -2
Query: 437 NSYSYRRRYDSEHWSRLRNGVRKLCRHPVFQLSKPSKKPASLGIVNTREG 486
NSYSYRRRYDSEHWSRLRNGVRKLCRHPVFQLSKPSKKPASLGIVNTREG
Sbjct: 467 NSYSYRRRYDSEHWSRLRNGVRKLCRHPVFQLSKPSKKPASLGIVNTREG 318
>TC91540 similar to GP|9757821|dbj|BAB08339.1 rac GTPase activating protein
{Arabidopsis thaliana}, partial (13%)
Length = 645
Score = 97.4 bits (241), Expect = 1e-20
Identities = 48/91 (52%), Positives = 64/91 (69%), Gaps = 1/91 (1%)
Frame = +1
Query: 79 QNQFAILDIVMAALKKSIVTC-SVEREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPSE 137
Q F + ++++ +KS+ S +D+ ++DIS PT VRHV+HVTFDRFNGFLGLP E
Sbjct: 373 QPHFPLFELLVTLFRKSLFPFKSSGNKDLCNMDISPPTNVRHVAHVTFDRFNGFLGLPDE 552
Query: 138 FQPEVPTRVPSASVKVFGVSAKSMQCSYDDR 168
F+P+ P R PSAS VF VS KS+Q SYD +
Sbjct: 553 FEPDFPRRPPSASATVFEVSTKSIQFSYDSK 645
>AL378182 similar to GP|3695063|gb|A rac GTPase activating protein 3 {Lotus
japonicus}, partial (12%)
Length = 218
Score = 50.1 bits (118), Expect = 2e-06
Identities = 27/50 (54%), Positives = 33/50 (66%), Gaps = 1/50 (2%)
Frame = +3
Query: 311 PLTALIHAVQVMNFLKTLILKMLREREESIDNA-RLLSHSMDFASCNDEF 359
P+TAL+HAVQVMN LKTLI+K LREREE+ +S F DE+
Sbjct: 3 PVTALMHAVQVMNLLKTLIMKTLREREETATGGYSPMSFRTSFRRSEDEY 152
>CA860191 homologue to GP|19697342|gb hypothetical protein {Dictyostelium
discoideum}, partial (1%)
Length = 504
Score = 40.0 bits (92), Expect = 0.002
Identities = 26/93 (27%), Positives = 45/93 (47%), Gaps = 6/93 (6%)
Frame = +2
Query: 3 RLFRSKSCNIAGFSVSEFNPAAQ---VTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
R F + S N G S+ V +KK ++ +Q +++EEE E D+ + D+
Sbjct: 119 RFFSAISANAEGKSLDRSPVDIMKDIVELDKKMQQGLDQTLQNDEEEEEEDEEEEDEFTT 298
Query: 60 IYSSNNASRQFTKGSNN---NHQNQFAILDIVM 89
++SNN + +NN NH N+F D ++
Sbjct: 299 TFNSNNNNNNSNNNNNNNEDNHHNKFNSEDAML 397
>TC93070 weakly similar to GP|17065230|gb|AAL32769.1 Unknown protein
{Arabidopsis thaliana}, partial (19%)
Length = 679
Score = 38.1 bits (87), Expect = 0.008
Identities = 14/57 (24%), Positives = 35/57 (60%)
Frame = +1
Query: 3 RLFRSKSCNIAGFSVSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
++ ++ + G +S ++ ++ ++++ E E+EDE+++ + DD+D DDDD+
Sbjct: 166 KILKTNRKSTYGKFLSPYDSDDEIEEMDFEDDEDEDEDEDEDDDDDDDDDDEDDDDD 336
Score = 34.7 bits (78), Expect = 0.084
Identities = 15/49 (30%), Positives = 31/49 (62%)
Frame = +1
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSNNNHQNQ 81
+E +E + ED+E+E E +D D+DDDD+ ++ F + ++ N +++
Sbjct: 229 DEIEEMDFEDDEDEDEDEDEDDDDDDDDDDEDDDDDGFAEPTDLNAKDK 375
>TC77387 similar to GP|15983396|gb|AAL11566.1 At1g51440/F5D21_19
{Arabidopsis thaliana}, partial (75%)
Length = 1793
Score = 37.7 bits (86), Expect = 0.010
Identities = 21/71 (29%), Positives = 41/71 (57%)
Frame = +3
Query: 27 TNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSNNNHQNQFAILD 86
T N+ EE++E+EEEDE+EE E ++ + +++ + + R+ +G N+ +LD
Sbjct: 306 TMNQLHEEEEEEEEEDEDEEEEEEEEEEEEESPLPPLSEVWREI-QGENDWE----GLLD 470
Query: 87 IVMAALKKSIV 97
+ L+K I+
Sbjct: 471 PMDPILRKEII 503
Score = 29.6 bits (65), Expect = 2.7
Identities = 10/37 (27%), Positives = 25/37 (67%)
Frame = +3
Query: 10 CNIAGFSVSEFNPAAQVTNNKKKEEQQEQEEEDEEEE 46
C+ + ++++ + + + ++E++E+EEE+EEEE
Sbjct: 288 CSSSSSTMNQLHEEEEEEEEEDEDEEEEEEEEEEEEE 398
Score = 29.3 bits (64), Expect = 3.5
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = +3
Query: 16 SVSEFNPAAQVTNNKKKEEQQEQEEEDEEEELE 48
S S N + +++E++ E+EEE+EEEE E
Sbjct: 297 SSSTMNQLHEEEEEEEEEDEDEEEEEEEEEEEE 395
>AW690594 similar to GP|23498163|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (10%)
Length = 633
Score = 37.7 bits (86), Expect = 0.010
Identities = 12/32 (37%), Positives = 29/32 (90%)
Frame = +3
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+++++E++ ++EEE+EEEE E D++D+++++E
Sbjct: 99 HDEEEEDEHDEEEEEEEEEEEEDEHDDEEEEE 194
Score = 37.0 bits (84), Expect = 0.017
Identities = 13/30 (43%), Positives = 24/30 (79%)
Frame = +3
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+++EE+ E ++E+EEEE E ++ + DDD+E
Sbjct: 147 EEEEEEDEHDDEEEEEEEEEEEEEEDDDEE 236
Score = 37.0 bits (84), Expect = 0.017
Identities = 14/29 (48%), Positives = 23/29 (79%)
Frame = +3
Query: 32 KEEQQEQEEEDEEEELESDDNDNDDDDEI 60
+ + +E+EEE+EEEE E DD++ ++DEI
Sbjct: 168 EHDDEEEEEEEEEEEEEEDDDEEGEEDEI 254
Score = 35.8 bits (81), Expect = 0.038
Identities = 12/30 (40%), Positives = 24/30 (80%)
Frame = +3
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+++EE++E E +DEEEE E ++ + ++DD+
Sbjct: 141 EEEEEEEEDEHDDEEEEEEEEEEEEEEDDD 230
Score = 35.4 bits (80), Expect = 0.049
Identities = 11/30 (36%), Positives = 25/30 (82%)
Frame = +3
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+++EE++++ +++EEEE E ++ + +DDDE
Sbjct: 144 EEEEEEEDEHDDEEEEEEEEEEEEEEDDDE 233
Score = 35.0 bits (79), Expect = 0.065
Identities = 13/41 (31%), Positives = 27/41 (65%)
Frame = +3
Query: 19 EFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
E + + +++E++ + EEE+EEEE E ++ D+D++ E
Sbjct: 120 EHDEEEEEEEEEEEEDEHDDEEEEEEEEEEEEEEDDDEEGE 242
Score = 34.7 bits (78), Expect = 0.084
Identities = 12/32 (37%), Positives = 25/32 (77%)
Frame = +3
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+ ++++E E+EEE+EEEE E + +D ++++E
Sbjct: 102 DEEEEDEHDEEEEEEEEEEEEDEHDDEEEEEE 197
Score = 34.7 bits (78), Expect = 0.084
Identities = 13/43 (30%), Positives = 27/43 (62%)
Frame = +3
Query: 17 VSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
V E N +++E++ ++EEEDE +E E ++ + +++DE
Sbjct: 42 VEEHNEEEDEGEVEEEEDEHDEEEEDEHDEEEEEEEEEEEEDE 170
Score = 33.5 bits (75), Expect = 0.19
Identities = 11/30 (36%), Positives = 25/30 (82%)
Frame = +3
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
++++E ++EEE+EEEE E +++D+++ +E
Sbjct: 156 EEEDEHDDEEEEEEEEEEEEEEDDDEEGEE 245
Score = 32.3 bits (72), Expect = 0.42
Identities = 11/30 (36%), Positives = 23/30 (76%)
Frame = +3
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
++ E +E+EEE+EEEE + D++ ++++E
Sbjct: 111 EEDEHDEEEEEEEEEEEEDEHDDEEEEEEE 200
>BF641220 weakly similar to PIR|G86203|G86 probable N-arginine dibasic
convertase [imported] - Arabidopsis thaliana, partial
(5%)
Length = 634
Score = 37.4 bits (85), Expect = 0.013
Identities = 15/41 (36%), Positives = 24/41 (57%)
Frame = +2
Query: 19 EFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
E P + E+ +E+++EDEE++ E DD DD+DE
Sbjct: 320 EIYPEGAPKDGSIDEDDEEEDDEDEEDDEEDDDEGEDDEDE 442
Score = 31.6 bits (70), Expect = 0.71
Identities = 10/32 (31%), Positives = 23/32 (71%)
Frame = +2
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+ ++ +E +E +EED++E + +D + +D+DE
Sbjct: 368 DEEEDDEDEEDDEEDDDEGEDDEDEEXEDEDE 463
>AW774261 weakly similar to EGAD|146423|156 vitellogenin {Anolis pulchellus},
partial (40%)
Length = 495
Score = 37.0 bits (84), Expect = 0.017
Identities = 18/37 (48%), Positives = 26/37 (69%), Gaps = 2/37 (5%)
Frame = +1
Query: 25 QVTNNKKKEEQQEQEEEDEEEELESDD--NDNDDDDE 59
+V N++ E++E+E E EEEE+E DD + DDDDE
Sbjct: 385 EVEENEEDAEEEEEE*EKEEEEVEEDDTMQNLDDDDE 495
Score = 30.0 bits (66), Expect = 2.1
Identities = 12/40 (30%), Positives = 25/40 (62%)
Frame = +1
Query: 19 EFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDD 58
E +V +++ E+ E++ E+EEEE E ++ + ++DD
Sbjct: 346 EVEEEEEVEEIEEEVEENEEDAEEEEEE*EKEEEEVEEDD 465
>TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~unknown
protein {Arabidopsis thaliana}, partial (46%)
Length = 2077
Score = 36.6 bits (83), Expect = 0.022
Identities = 18/39 (46%), Positives = 24/39 (61%), Gaps = 7/39 (17%)
Frame = +2
Query: 28 NNKKKEEQQEQEEEDEEE-------ELESDDNDNDDDDE 59
N +K EE+ E +E+ EE+ E E DD D+DDDDE
Sbjct: 629 NYEKGEERDEADEDSEEDDELEKAGEFEMDDEDDDDDDE 745
Score = 31.2 bits (69), Expect = 0.93
Identities = 12/24 (50%), Positives = 19/24 (79%)
Frame = +2
Query: 30 KKKEEQQEQEEEDEEEELESDDND 53
KK+ E +E+E +D +EEL+ DD+D
Sbjct: 356 KKRVELEEEESDDFDEELDDDDDD 427
Score = 30.0 bits (66), Expect = 2.1
Identities = 11/27 (40%), Positives = 21/27 (77%)
Frame = +2
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
E + + EEED+E+E S+D D+++D++
Sbjct: 467 EGEDDFEEEDDEDEEGSEDEDDEEDEK 547
Score = 28.5 bits (62), Expect = 6.0
Identities = 12/30 (40%), Positives = 19/30 (63%)
Frame = +2
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
KK + E E++ EEE+ E ++ D+DDE
Sbjct: 446 KKAKGGSEGEDDFEEEDDEDEEGSEDEDDE 535
>TC83341 weakly similar to GP|8978076|dbj|BAA98104.1 gene_id:K14A3.4~unknown
protein {Arabidopsis thaliana}, partial (80%)
Length = 1253
Score = 36.6 bits (83), Expect = 0.022
Identities = 16/39 (41%), Positives = 28/39 (71%)
Frame = +1
Query: 22 PAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEI 60
P ++ N+ +E++E+EEE+EEEE E DD+D + D+ +
Sbjct: 550 PRSEWIGNEGLQEEEEEEEEEEEEE-EEDDDDVEKDETV 663
Score = 35.8 bits (81), Expect = 0.038
Identities = 15/34 (44%), Positives = 24/34 (70%)
Frame = +1
Query: 26 VTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+ N +EE++E+EEE+EEEE + DD + D+ E
Sbjct: 565 IGNEGLQEEEEEEEEEEEEEEEDDDDVEKDETVE 666
>TC89903 similar to GP|15810167|gb|AAL06985.1 At1g64110/F22C12_22
{Arabidopsis thaliana}, partial (20%)
Length = 875
Score = 36.6 bits (83), Expect = 0.022
Identities = 25/99 (25%), Positives = 44/99 (44%), Gaps = 9/99 (9%)
Frame = +1
Query: 5 FRSKSCNIAGFSVSE---------FNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDND 55
F+ + GFS S+ + P ++ ++K+E +++++E E E N ND
Sbjct: 211 FKELATMTEGFSGSDLKNLCITAAYRPVRELIQQERKKEMEKKKKEAESVNSEDASNSND 390
Query: 56 DDDEIYSSNNASRQFTKGSNNNHQNQFAILDIVMAALKK 94
+D+ S + K + N FA VMA LK+
Sbjct: 391 NDEPEIILRPLSMEDMKQAKNQVAASFASEGSVMAELKQ 507
>TC88983 similar to SP|P93231|VP41_LYCES Vacuolar assembly protein VPS41
homolog. [Tomato] {Lycopersicon esculentum}, partial
(17%)
Length = 668
Score = 36.2 bits (82), Expect = 0.029
Identities = 11/28 (39%), Positives = 26/28 (92%)
Frame = +2
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDND 55
+++++E+++++EEEDE+EE+E DD++ +
Sbjct: 146 DDEREEDEEDEEEEDEDEEVEEDDDEEE 229
Score = 36.2 bits (82), Expect = 0.029
Identities = 14/32 (43%), Positives = 24/32 (74%)
Frame = +2
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
N ++++E++EEDEEEE E ++ + DDD+E
Sbjct: 131 NGVDGDDEREEDEEDEEEEDEDEEVEEDDDEE 226
Score = 30.8 bits (68), Expect = 1.2
Identities = 10/27 (37%), Positives = 22/27 (81%)
Frame = +2
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
+E++E EE++EEE+ + + ++DD++E
Sbjct: 149 DEREEDEEDEEEEDEDEEVEEDDDEEE 229
>TC79394 weakly similar to PIR|T49019|T49019 probable RNA binding protein -
Arabidopsis thaliana, partial (74%)
Length = 1909
Score = 35.8 bits (81), Expect = 0.038
Identities = 15/35 (42%), Positives = 24/35 (67%)
Frame = +2
Query: 32 KEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNA 66
+EE +E+EEE+EEEE+E + D++DE +A
Sbjct: 308 EEEVEEEEEEEEEEEVEEESKPLDEEDEADKKKHA 412
Score = 30.0 bits (66), Expect = 2.1
Identities = 13/32 (40%), Positives = 20/32 (61%)
Frame = +2
Query: 25 QVTNNKKKEEQQEQEEEDEEEELESDDNDNDD 56
+V ++EE++E+EEE EEE D+ D D
Sbjct: 302 EVEEEVEEEEEEEEEEEVEEESKPLDEEDEAD 397
>TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Ateline
herpesvirus 3}, partial (8%)
Length = 986
Score = 35.0 bits (79), Expect = 0.065
Identities = 13/27 (48%), Positives = 20/27 (73%)
Frame = +3
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
E +E+++ED +++ E DD D DDDDE
Sbjct: 585 ENGEEEDDEDGDDQDEDDDEDEDDDDE 665
Score = 33.1 bits (74), Expect = 0.25
Identities = 11/27 (40%), Positives = 22/27 (80%)
Frame = +3
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
++++E EEDEEE ++ +DN+ +++DE
Sbjct: 657 DDEEEGGEEDEEEGVDEEDNEEEEEDE 737
Score = 29.6 bits (65), Expect = 2.7
Identities = 8/27 (29%), Positives = 22/27 (80%)
Frame = +3
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
+E E+E++++ ++ + DD++++DDD+
Sbjct: 582 DENGEEEDDEDGDDQDEDDDEDEDDDD 662
Score = 29.3 bits (64), Expect = 3.5
Identities = 9/37 (24%), Positives = 24/37 (64%)
Frame = +3
Query: 23 AAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
A N ++E+ ++ +++DE+++ + DD+D ++ E
Sbjct: 570 AGGADENGEEEDDEDGDDQDEDDDEDEDDDDEEEGGE 680
Score = 28.9 bits (63), Expect = 4.6
Identities = 9/27 (33%), Positives = 20/27 (73%)
Frame = +3
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
E+ +Q+E+D+E+E + D+ + ++DE
Sbjct: 609 EDGDDQDEDDDEDEDDDDEEEGGEEDE 689
>TC92853 similar to GP|20268748|gb|AAM14077.1 unknown protein {Arabidopsis
thaliana}, partial (21%)
Length = 761
Score = 34.7 bits (78), Expect = 0.084
Identities = 13/31 (41%), Positives = 25/31 (79%)
Frame = +3
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDD 58
+++ KE ++E+E+EDEEEE SDD+ ++ ++
Sbjct: 369 DDEGKENEEEEEDEDEEEEDTSDDDGSESEE 461
Score = 33.5 bits (75), Expect = 0.19
Identities = 13/33 (39%), Positives = 22/33 (66%)
Frame = +3
Query: 27 TNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
++N +E +E EEE+E+E+ E +D +DD E
Sbjct: 354 SDNDNDDEGKENEEEEEDEDEEEEDTSDDDGSE 452
Score = 33.1 bits (74), Expect = 0.25
Identities = 12/40 (30%), Positives = 26/40 (65%)
Frame = +3
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNAS 67
N+ + +E +E+EE+++EEE ++ D+D + +E + S
Sbjct: 366 NDDEGKENEEEEEDEDEEEEDTSDDDGSESEEFNADEEDS 485
Score = 32.3 bits (72), Expect = 0.42
Identities = 13/48 (27%), Positives = 28/48 (58%)
Frame = +3
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSN 75
+N + ++ E+EEEDE+EE E +D+ + E ++++ T+ +
Sbjct: 363 DNDDEGKENEEEEEDEDEEEEDTSDDDGSESEEFNADEEDSD*TRADS 506
>TC86034 similar to GP|6815067|dbj|BAA90354.1 ESTs AU082210(C53655)
AU056546(S20671) AU056545(S20671) correspond to a
region of the predicted, partial (42%)
Length = 2412
Score = 34.7 bits (78), Expect = 0.084
Identities = 23/65 (35%), Positives = 30/65 (45%), Gaps = 5/65 (7%)
Frame = +3
Query: 17 VSEFNPAAQVTNNKKKEEQQEQ-----EEEDEEEELESDDNDNDDDDEIYSSNNASRQFT 71
V+E P Q N EE Q Q EEE EEE E +++D +DE N +
Sbjct: 177 VAEEEPTTQEEENTPIEEDQPQNEGEGEEEPEEEVAEEEEDDQQGEDETL-ENQQQQDGG 353
Query: 72 KGSNN 76
+ SNN
Sbjct: 354 EASNN 368
Score = 28.9 bits (63), Expect = 4.6
Identities = 20/73 (27%), Positives = 35/73 (47%), Gaps = 8/73 (10%)
Frame = +3
Query: 17 VSEFNPAAQVTNNKKKEEQQEQEEEDEEE-ELESDDNDNDDDDEIYSSNNASRQFT---- 71
+ E P + ++ EE+ +EEED+++ E E+ +N D S+N A + T
Sbjct: 222 IEEDQPQNEGEGEEEPEEEVAEEEEDDQQGEDETLENQQQQDGGEASNNAAVSEQTVTVE 401
Query: 72 ---KGSNNNHQNQ 81
G NNN + +
Sbjct: 402 ANGGGENNNGEEE 440
>TC78476 homologue to SP|O22437|CHLD_PEA Magnesium-chelatase subunit chlD
chloroplast precursor (Mg- protoporphyrin IX chelatase),
partial (68%)
Length = 1596
Score = 34.7 bits (78), Expect = 0.084
Identities = 15/38 (39%), Positives = 24/38 (62%)
Frame = +2
Query: 22 PAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
P N + EEQ E+E+++EEEE E DD D + +++
Sbjct: 506 PPPPPQNQESNEEQNEEEDKEEEEE-EEDDKDEEKEEQ 616
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.129 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,509,654
Number of Sequences: 36976
Number of extensions: 176354
Number of successful extensions: 2686
Number of sequences better than 10.0: 166
Number of HSP's better than 10.0 without gapping: 1525
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2106
length of query: 486
length of database: 9,014,727
effective HSP length: 100
effective length of query: 386
effective length of database: 5,317,127
effective search space: 2052411022
effective search space used: 2052411022
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144592.1


