

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144591.1 - phase: 0
(167 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
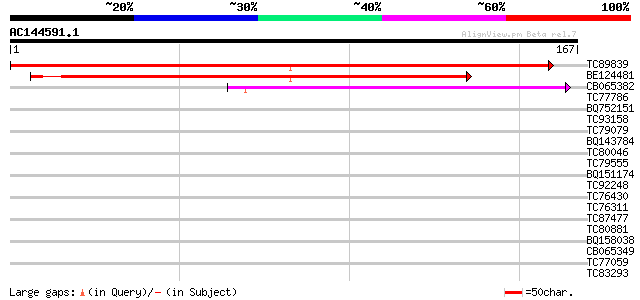
Sequences producing significant alignments: (bits) Value
TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC ... 191 2e-49
BE124481 151 1e-37
CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fra... 86 7e-18
TC77786 35 0.014
BQ752151 similar to GP|1945155|emb MN1 {Homo sapiens}, partial (1%) 33 0.041
TC93158 similar to PIR|T01123|T01123 hypothetical protein At2g32... 33 0.041
TC79079 similar to GP|19698893|gb|AAL91182.1 splicing factor pu... 32 0.091
BQ143784 weakly similar to SP|P33485|VNUA Probable nuclear antig... 32 0.12
TC80046 similar to SP|P22699|GUN6_DICDI Endoglucanase precursor ... 32 0.16
TC79555 homologue to GP|10567859|gb|AAG18593.1 L8019.5 {Leishman... 32 0.16
BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xy... 30 0.45
TC92248 weakly similar to GP|22535591|dbj|BAC10766. putative DEI... 30 0.45
TC76430 similar to PIR|T07623|T07623 extensin homolog HRGP2 - so... 29 0.77
TC76311 homologue to GP|3204132|emb|CAA07235.1 extensin {Cicer a... 29 0.77
TC87477 homologue to SP|O43992|RS2_LEIAM 40S ribosomal protein S... 29 1.0
TC80881 similar to PIR|T11622|T11622 extensin class 1 precursor ... 28 1.3
BQ158038 similar to GP|4775353|emb hyperpolarization-activated c... 28 1.7
CB065349 similar to SP|P10713|CONX_N Conidiation-specific protei... 28 1.7
TC77059 similar to GP|18958499|gb|AAL82617.1 elongation factor 1... 28 1.7
TC83293 similar to GP|6648203|gb|AAF21201.1| unknown protein {Ar... 28 1.7
>TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC 4.6.1.1)
(ATP pyrophosphate-lyase) (Adenylyl cyclase). [Yeast],
partial (0%)
Length = 665
Score = 191 bits (484), Expect = 2e-49
Identities = 100/181 (55%), Positives = 118/181 (64%), Gaps = 21/181 (11%)
Frame = +3
Query: 1 MSSFNLSAHMQTLAQTHKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQ 60
MSS +HMQTL QT KMDQLEQE+HELR E+T LRA+VE+LT+ VSSLM T D P Q
Sbjct: 63 MSSSKFGSHMQTLVQTQKMDQLEQEVHELRGEVTTLRAEVEKLTSLVSSLMATNDPPLVQ 242
Query: 61 QRPQPPRQPSFTQQPRQQAPR---------------------TRFDPIPMKYAEFLPSLL 99
QRPQ P QP Q+PRQQAPR ++ DPIP+KYA+ LP LL
Sbjct: 243 QRPQSPYQPMCPQKPRQQAPRQSIPQNQVPQKFIPQNQVQKASQCDPIPVKYADLLPILL 422
Query: 100 KGNYVQTRPPPPRPEKLPAGFRADLSCVFHQGAPGHDVERCYALRNAVQDLVEAKILSFT 159
K N +QT P P P LP +R DL+CVFHQGAPGHD E+CY L+ VQ L+E + SF
Sbjct: 423 KKNLIQTLPLPRVPNSLPPWYRPDLNCVFHQGAPGHDTEQCYPLKEEVQKLIENNVWSFD 602
Query: 160 D 160
D
Sbjct: 603 D 605
>BE124481
Length = 538
Score = 151 bits (381), Expect = 1e-37
Identities = 83/151 (54%), Positives = 97/151 (63%), Gaps = 21/151 (13%)
Frame = +3
Query: 7 SAHMQTLAQTHKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQPP 66
SAH+ T KMDQLEQE+HELR E+T LRA+VE+LT+ VSSLM T D P QQRPQ P
Sbjct: 99 SAHI-----TQKMDQLEQEVHELRGEVTTLRAEVEKLTSLVSSLMATNDPPLVQQRPQSP 263
Query: 67 RQPSFTQQPRQQAPR---------------------TRFDPIPMKYAEFLPSLLKGNYVQ 105
QP Q+PRQQAPR ++ DPIP+KYA+ LP LLK N +Q
Sbjct: 264 YQPMCPQKPRQQAPRQSIPQNQVPQKFIPQNQVQKASQCDPIPVKYADLLPILLKKNLIQ 443
Query: 106 TRPPPPRPEKLPAGFRADLSCVFHQGAPGHD 136
T P P P LP +R DL+CVFHQGAPGHD
Sbjct: 444 TLPLPRVPNSLPPWYRPDLNCVFHQGAPGHD 536
>CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fragment),
partial (9%)
Length = 624
Score = 85.9 bits (211), Expect = 7e-18
Identities = 48/103 (46%), Positives = 61/103 (58%), Gaps = 2/103 (1%)
Frame = -2
Query: 65 PPRQ--PSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEKLPAGFRA 122
PPR P QQ + Q R F IPM YAE LP+LL + R P P+ LP FR+
Sbjct: 539 PPRNNHPPVHQQRQ*QQARPTFPLIPMLYAE*LPTLLLRGHCTIRQGKPPPDPLPPRFRS 360
Query: 123 DLSCVFHQGAPGHDVERCYALRNAVQDLVEAKILSFTDLNPNL 165
DL C FHQGA GHDVE CYAL++ V+ L+ L+F + P++
Sbjct: 359 DLKCDFHQGALGHDVEGCYALKHIVKKLINQGKLTFENNVPHV 231
>TC77786
Length = 1094
Score = 35.0 bits (79), Expect = 0.014
Identities = 19/52 (36%), Positives = 30/52 (57%), Gaps = 3/52 (5%)
Frame = +3
Query: 112 RPEKLPAGFRADLS--CVFHQGAPGHDVERCYALRNAVQDLV-EAKILSFTD 160
+P+ L R D + C +H+ GHD E C+ LR A++DL+ + K+ F D
Sbjct: 777 KPQLLK*SARTDKAK*CCYHRSR-GHDTEECFQLREAIEDLIKKVKLGRFID 929
>BQ752151 similar to GP|1945155|emb MN1 {Homo sapiens}, partial (1%)
Length = 650
Score = 33.5 bits (75), Expect = 0.041
Identities = 24/69 (34%), Positives = 36/69 (51%), Gaps = 8/69 (11%)
Frame = +3
Query: 57 PQFQQRPQ--PPRQPSFTQQPRQQAPRTRFD-PIPMKYAEFLPSLLKGNYVQ-----TRP 108
PQ QQ+P PP P +QP+QQAP + P P A+ +P G ++Q + P
Sbjct: 297 PQSQQQPSAWPPSHPPPPKQPQQQAPPPQQAWPSPHVSAQKIP---PGGHLQQQTWASAP 467
Query: 109 PPPRPEKLP 117
P +P++ P
Sbjct: 468 PQQQPQQAP 494
>TC93158 similar to PIR|T01123|T01123 hypothetical protein At2g32840
[imported] - Arabidopsis thaliana, partial (9%)
Length = 673
Score = 33.5 bits (75), Expect = 0.041
Identities = 21/61 (34%), Positives = 24/61 (38%)
Frame = +2
Query: 57 PQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEKL 116
P +P P QQP+QQA R P P Y PS N+ PPP P
Sbjct: 98 PNPNPNSKPIHYPQQQQQPQQQALHVR-SPNPFLYPFASPSRASANHAVGGYPPPPPPSQ 274
Query: 117 P 117
P
Sbjct: 275 P 277
>TC79079 similar to GP|19698893|gb|AAL91182.1 splicing factor putative
{Arabidopsis thaliana}, partial (17%)
Length = 1114
Score = 32.3 bits (72), Expect = 0.091
Identities = 20/64 (31%), Positives = 27/64 (41%), Gaps = 11/64 (17%)
Frame = +1
Query: 65 PPRQPSFTQQ----PRQQAPRTRFDPIPMKYAE-------FLPSLLKGNYVQTRPPPPRP 113
PP PS + P P ++F P+P YA +P + Q PPPP P
Sbjct: 187 PPMHPSISMNNQGIPIPPPPGSQFTPVPRPYAPHPHPSSGMMPMMHPPPPPQGVPPPPPP 366
Query: 114 EKLP 117
E+ P
Sbjct: 367 EEAP 378
>BQ143784 weakly similar to SP|P33485|VNUA Probable nuclear antigen. [strain
Kaplan PRV] {Pseudorabies virus}, partial (2%)
Length = 1022
Score = 32.0 bits (71), Expect = 0.12
Identities = 20/54 (37%), Positives = 23/54 (42%)
Frame = +3
Query: 60 QQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRP 113
+Q PP P+ +P Q P R P P A G QT PPPPRP
Sbjct: 57 KQNEGPPPHPTRPPRPPQDPPPARNPPQPTGTATRRKGGA-GRGAQTDPPPPRP 215
>TC80046 similar to SP|P22699|GUN6_DICDI Endoglucanase precursor (EC
3.2.1.4) (Endo-1 4-beta-glucanase) (Spore germination
protein 270-6), partial (11%)
Length = 643
Score = 31.6 bits (70), Expect = 0.16
Identities = 23/74 (31%), Positives = 34/74 (45%), Gaps = 9/74 (12%)
Frame = +1
Query: 53 TKDLPQFQQRPQ---------PPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNY 103
T+ LP Q+ PQ PP++P Q+P Q+ P+ +P + + LP
Sbjct: 256 TRSLPLPQRPPQRPPQRPPQKPPQRPP--QKPPQRPPQRPPQKLPQRPPQKLPQ------ 411
Query: 104 VQTRPPPPRPEKLP 117
RPP P+KLP
Sbjct: 412 ---RPPQKLPQKLP 444
Score = 28.1 bits (61), Expect = 1.7
Identities = 20/59 (33%), Positives = 30/59 (49%), Gaps = 2/59 (3%)
Frame = +1
Query: 61 QRP--QPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEKLP 117
QRP +PP++P Q+P Q+ P+ +P + + LP L Q P P P+K P
Sbjct: 325 QRPPQKPPQRPP--QRPPQKLPQRPPQKLPQRPPQKLPQKLPQKPPQR--PLPHPQKPP 489
>TC79555 homologue to GP|10567859|gb|AAG18593.1 L8019.5 {Leishmania major},
partial (1%)
Length = 1410
Score = 31.6 bits (70), Expect = 0.16
Identities = 25/79 (31%), Positives = 32/79 (39%), Gaps = 13/79 (16%)
Frame = +2
Query: 57 PQFQQRPQPPRQPSFTQQPRQ-----QAPRTRFDPI-------PMKYAEFLPSLLKGNYV 104
P RP P +PS TQ P + + P T P P F P+ G
Sbjct: 536 PATTTRPPPTSKPSPTQTPSRSPSSARTPSTSSTPTRAPLLSSPPPPPPFFPARRPGTRS 715
Query: 105 QTRPPP-PRPEKLPAGFRA 122
+ PP PRP +LP+ RA
Sbjct: 716 LPKSPPRPRPARLPSARRA 772
>BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xylostella
granulovirus}, partial (42%)
Length = 909
Score = 30.0 bits (66), Expect = 0.45
Identities = 20/62 (32%), Positives = 27/62 (43%), Gaps = 2/62 (3%)
Frame = +1
Query: 60 QQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLL-KGNYVQTRP-PPPRPEKLP 117
Q PQPP+ P ++PR P P P + P+ +G RP PPP P +
Sbjct: 262 QPPPQPPKPPHHEKRPRTPEPPGGRPPPPTTPKQARPTTQERGGARGARPRPPPAPPQGT 441
Query: 118 AG 119
G
Sbjct: 442 GG 447
>TC92248 weakly similar to GP|22535591|dbj|BAC10766. putative DEIH-box
RNA/DNA helicase {Oryza sativa (japonica
cultivar-group)}, partial (4%)
Length = 605
Score = 30.0 bits (66), Expect = 0.45
Identities = 18/44 (40%), Positives = 24/44 (53%), Gaps = 2/44 (4%)
Frame = +3
Query: 40 VEELTNRVSSLMVTKDLPQFQQRPQP--PRQPSFTQQPRQQAPR 81
V+ LT V++ + K PQ Q +PQP QP QQP+ Q R
Sbjct: 210 VDNLTTMVNATNLGKPQPQPQPQPQPQPQPQPQPQQQPQPQPQR 341
>TC76430 similar to PIR|T07623|T07623 extensin homolog HRGP2 - soybean
(fragment), partial (61%)
Length = 912
Score = 29.3 bits (64), Expect = 0.77
Identities = 20/63 (31%), Positives = 22/63 (34%), Gaps = 2/63 (3%)
Frame = +3
Query: 57 PQFQQRPQPPRQ-PSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGN-YVQTRPPPPRPE 114
P + Q P PP P + P T P P Y P Y T PPPP P
Sbjct: 57 PYYYQSPPPPSPTPHTPYHYKSPPPPTASPPPPYHYVSPPPPTSSPPPYHYTSPPPPSPA 236
Query: 115 KLP 117
P
Sbjct: 237 PAP 245
>TC76311 homologue to GP|3204132|emb|CAA07235.1 extensin {Cicer arietinum},
partial (77%)
Length = 973
Score = 29.3 bits (64), Expect = 0.77
Identities = 20/63 (31%), Positives = 22/63 (34%), Gaps = 2/63 (3%)
Frame = +2
Query: 57 PQFQQRPQPPRQ-PSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGN-YVQTRPPPPRPE 114
P + Q P PP P + P T P P Y P Y T PPPP P
Sbjct: 170 PYYYQSPPPPSPTPHTPYHYKSPPPPTASPPPPYHYVSPPPPTSSPPPYHYTSPPPPSPA 349
Query: 115 KLP 117
P
Sbjct: 350 PAP 358
>TC87477 homologue to SP|O43992|RS2_LEIAM 40S ribosomal protein S2.
{Leishmania amazonensis}, partial (6%)
Length = 836
Score = 28.9 bits (63), Expect = 1.0
Identities = 18/50 (36%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Frame = -3
Query: 69 PSFTQQPRQQAPRTRFDPIPMKYAE-FLPSLLKGNYVQTRPPPPRPEKLP 117
P F Q P P+ + P+P + FLPS ++RPP PRP + P
Sbjct: 234 PKFPQSPM---PQFQSPPLPQSLSSCFLPS-------RSRPPRPRPPRPP 115
Score = 26.9 bits (58), Expect = 3.8
Identities = 16/53 (30%), Positives = 23/53 (43%), Gaps = 2/53 (3%)
Frame = -3
Query: 63 PQPPRQPSFTQQPRQQAPRTRF--DPIPMKYAEFLPSLLKGNYVQTRPPPPRP 113
P PP P+ P +F P+P + LP L ++ +R PPRP
Sbjct: 291 PPPPYSPTS*YGCLNPLPGPKFPQSPMPQFQSPPLPQSLSSCFLPSRSRPPRP 133
>TC80881 similar to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (34%)
Length = 597
Score = 28.5 bits (62), Expect = 1.3
Identities = 25/97 (25%), Positives = 34/97 (34%), Gaps = 5/97 (5%)
Frame = +3
Query: 26 LHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQPPRQPSFTQQP---RQQAPRT 82
LH L + + +T + + P + + PP PS P + P +
Sbjct: 246 LHLLHRHHLHMSTNTHLITTNLHLHHLPSPPPPYVYKSPPPPSPS-PPPPYVYKSPPPPS 422
Query: 83 RFDPIPMKYAEFLPSL--LKGNYVQTRPPPPRPEKLP 117
P P Y P L NYV PPPP P P
Sbjct: 423 PSPPPPYVYKSPPPPSPSLPPNYVYKSPPPPSPSPPP 533
>BQ158038 similar to GP|4775353|emb hyperpolarization-activated cyclic
nucleotide gated cation channel hHCN4 {Homo sapiens},
partial (1%)
Length = 1060
Score = 28.1 bits (61), Expect = 1.7
Identities = 29/87 (33%), Positives = 34/87 (38%), Gaps = 4/87 (4%)
Frame = +3
Query: 62 RPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEKLPAGFR 121
+PQ P Q S P + P TR K + P L GN PP +P P R
Sbjct: 138 QPQAPCQLSIPHVPTFR*P-TRVQKKRSKEKKKAP--LYGNPGVPNPPASQPPGTPPERR 308
Query: 122 ----ADLSCVFHQGAPGHDVERCYALR 144
F APG D+ERC LR
Sbjct: 309 HRWPPSKPNEFPPRAPGGDIERCSDLR 389
>CB065349 similar to SP|P10713|CONX_N Conidiation-specific protein 10.
{Neurospora crassa}, partial (27%)
Length = 597
Score = 28.1 bits (61), Expect = 1.7
Identities = 15/33 (45%), Positives = 18/33 (54%), Gaps = 2/33 (6%)
Frame = +2
Query: 60 QQRPQPPRQPSFTQQPRQ--QAPRTRFDPIPMK 90
Q+R QP RQP Q+P Q Q PR F P +
Sbjct: 170 QERQQPHRQPGQQQEPGQLRQRPRESFRSRPQR 268
>TC77059 similar to GP|18958499|gb|AAL82617.1 elongation factor 1-gamma
{Glycine max}, partial (96%)
Length = 1582
Score = 28.1 bits (61), Expect = 1.7
Identities = 13/33 (39%), Positives = 19/33 (57%)
Frame = +2
Query: 54 KDLPQFQQRPQPPRQPSFTQQPRQQAPRTRFDP 86
K L Q +Q P PS QQP++ P+T+ +P
Sbjct: 683 KILGQVKQTEAMPPIPSAKQQPKESKPKTKDEP 781
>TC83293 similar to GP|6648203|gb|AAF21201.1| unknown protein {Arabidopsis
thaliana}, complete
Length = 421
Score = 28.1 bits (61), Expect = 1.7
Identities = 23/79 (29%), Positives = 29/79 (36%), Gaps = 3/79 (3%)
Frame = -2
Query: 47 VSSLMVTKDLPQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIP---MKYAEFLPSLLKGNY 103
+ SL V D PQ + P P ++P PR P P A ++P
Sbjct: 258 IKSLWVQYDFPQCIKLPAPKKRPKIAALPRVVCPICLIFKNPGTLYLIASYVPYT----- 94
Query: 104 VQTRPPPPRPEKLPAGFRA 122
T PP P PA RA
Sbjct: 93 APTAPPTAPPTAAPAVTRA 37
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.132 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,923,439
Number of Sequences: 36976
Number of extensions: 76361
Number of successful extensions: 769
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 693
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 744
length of query: 167
length of database: 9,014,727
effective HSP length: 89
effective length of query: 78
effective length of database: 5,723,863
effective search space: 446461314
effective search space used: 446461314
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC144591.1


