

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144541.14 - phase: 0
(63 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
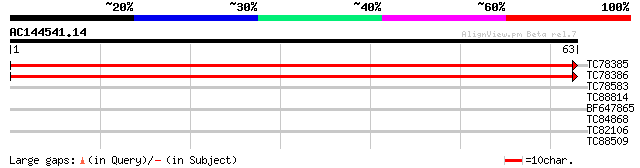
TC78385 weakly similar to PIR|T21861|T21861 hypothetical protein... 125 4e-30
TC78386 similar to SP|Q9NZW4|DSPP_HUMAN Dentin sialophosphoprote... 125 4e-30
TC78583 similar to GP|4097585|gb|AAD09518.1| NTGP4 {Nicotiana ta... 28 0.54
TC88814 homologue to PIR|T06447|T06447 GTP-binding protein - gar... 28 0.70
BF647865 27 1.2
TC84868 homologue to GP|18376071|emb|CAD21099. hypothetical prot... 25 4.5
TC82106 similar to PIR|T49191|T49191 hypothetical protein MAA21.... 25 5.9
TC88509 similar to PIR|T01826|T01826 microfibril-associated prot... 25 7.7
>TC78385 weakly similar to PIR|T21861|T21861 hypothetical protein F36F2.3 -
Caenorhabditis elegans, partial (1%)
Length = 1417
Score = 125 bits (313), Expect = 4e-30
Identities = 63/63 (100%), Positives = 63/63 (100%)
Frame = +1
Query: 1 PNGERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNA 60
PNGERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNA
Sbjct: 907 PNGERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNA 1086
Query: 61 SFD 63
SFD
Sbjct: 1087SFD 1095
>TC78386 similar to SP|Q9NZW4|DSPP_HUMAN Dentin sialophosphoprotein
precursor [Contains: Dentin phosphoprotein (Dentin
phosphophoryn) (DPP);, partial (2%)
Length = 726
Score = 125 bits (313), Expect = 4e-30
Identities = 63/63 (100%), Positives = 63/63 (100%)
Frame = +3
Query: 1 PNGERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNA 60
PNGERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNA
Sbjct: 423 PNGERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNA 602
Query: 61 SFD 63
SFD
Sbjct: 603 SFD 611
>TC78583 similar to GP|4097585|gb|AAD09518.1| NTGP4 {Nicotiana tabacum},
partial (86%)
Length = 1269
Score = 28.5 bits (62), Expect = 0.54
Identities = 14/49 (28%), Positives = 31/49 (62%), Gaps = 1/49 (2%)
Frame = +3
Query: 12 LSKFLGRNKDDGVRRS-AVSGKKILLKLDKTKEDKEAESKRNELLNFLN 59
L +LGR + +++ ++ G + +L +KTK++K+ + +LL+F+N
Sbjct: 546 LDDYLGRECPESLKQILSLCGNRCVLFDNKTKDEKKRSGQVQQLLSFVN 692
>TC88814 homologue to PIR|T06447|T06447 GTP-binding protein - garden pea,
complete
Length = 1183
Score = 28.1 bits (61), Expect = 0.70
Identities = 14/24 (58%), Positives = 15/24 (62%)
Frame = -3
Query: 36 LKLDKTKEDKEAESKRNELLNFLN 59
LKLDK KE +KRNE FLN
Sbjct: 1175 LKLDK*IPSKETRTKRNESARFLN 1104
>BF647865
Length = 638
Score = 27.3 bits (59), Expect = 1.2
Identities = 13/28 (46%), Positives = 17/28 (60%)
Frame = +2
Query: 25 RRSAVSGKKILLKLDKTKEDKEAESKRN 52
+RS V + LD++ EDKE E KRN
Sbjct: 218 KRSVVGACSPEVSLDESMEDKELEGKRN 301
>TC84868 homologue to GP|18376071|emb|CAD21099. hypothetical protein
{Neurospora crassa}, partial (0%)
Length = 288
Score = 25.4 bits (54), Expect = 4.5
Identities = 11/26 (42%), Positives = 17/26 (65%)
Frame = -3
Query: 20 KDDGVRRSAVSGKKILLKLDKTKEDK 45
++D VRR+ GKK+ L DK + D+
Sbjct: 202 REDMVRRAIQEGKKLGLHFDKGRADR 125
>TC82106 similar to PIR|T49191|T49191 hypothetical protein MAA21.130 -
Arabidopsis thaliana, partial (5%)
Length = 990
Score = 25.0 bits (53), Expect = 5.9
Identities = 16/51 (31%), Positives = 26/51 (50%)
Frame = +2
Query: 3 GERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNE 53
GE PV+ ++ KD+ +G +I +T EDK+A+ K+NE
Sbjct: 203 GELEPEPVRETELKPAPKDEA------AGSEI----QQTSEDKQAQKKKNE 325
>TC88509 similar to PIR|T01826|T01826 microfibril-associated protein homolog
T15F16.8 - Arabidopsis thaliana, partial (83%)
Length = 1306
Score = 24.6 bits (52), Expect = 7.7
Identities = 13/47 (27%), Positives = 26/47 (54%), Gaps = 2/47 (4%)
Frame = +2
Query: 14 KFLGRNKDDGVRRSAVS--GKKILLKLDKTKEDKEAESKRNELLNFL 58
+ L R ++DG+RRS V +K L+ + K K+ +R+ ++ +
Sbjct: 425 RMLWRREEDGLRRSCVREIRRKRFLRKRRRKRRKKRRRRRSLIMRLI 565
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.307 0.129 0.336
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,351,539
Number of Sequences: 36976
Number of extensions: 10658
Number of successful extensions: 55
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 55
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 55
length of query: 63
length of database: 9,014,727
effective HSP length: 39
effective length of query: 24
effective length of database: 7,572,663
effective search space: 181743912
effective search space used: 181743912
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 51 (24.3 bits)
Medicago: description of AC144541.14


