

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
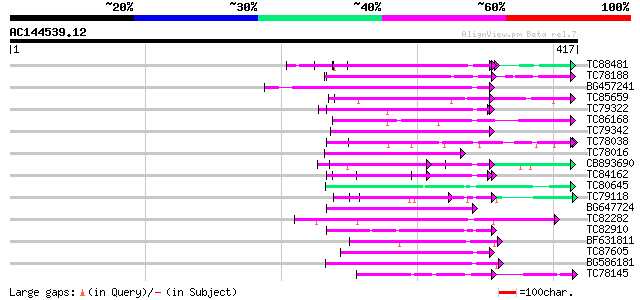
Query= AC144539.12 + phase: 0
(417 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC88481 similar to PIR|T00806|T02445 probable U4/U6 small nuclea... 79 3e-15
TC78188 similar to PIR|C84870|C84870 probable splicing factor [i... 67 1e-11
BG457241 homologue to PIR|F84600|F8 coatomer alpha subunit [impo... 64 1e-10
TC85659 SP|O24076|GBLP_MEDSA Guanine nucleotide-binding protein ... 63 2e-10
TC79322 similar to GP|13605815|gb|AAK32893.1 At2g32700/F24L7.16 ... 63 2e-10
TC86168 homologue to GP|13936812|gb|AAK49947.1 TGF-beta receptor... 60 2e-09
TC79342 similar to GP|10177096|dbj|BAB10430. Notchless protein h... 59 3e-09
TC78038 weakly similar to GP|21592898|gb|AAM64848.1 unknown {Ara... 59 5e-09
TC78016 similar to GP|21537191|gb|AAM61532.1 PRL1 protein {Arabi... 59 5e-09
CB893690 weakly similar to GP|9759025|dbj gene_id:K6A12.9~simila... 45 6e-09
TC84162 similar to GP|15810485|gb|AAL07130.1 putative WD40-repea... 57 1e-08
TC80645 similar to PIR|T50983|T50983 probable pleiotropic regula... 57 2e-08
TC79118 similar to GP|16323188|gb|AAL15328.1 AT5g67320/K8K14_4 {... 56 2e-08
BG647724 similar to GP|11141605|gb LEUNIG {Arabidopsis thaliana}... 56 2e-08
TC82282 similar to PIR|T08544|T08544 hypothetical protein F27B13... 55 5e-08
TC82910 similar to GP|19347844|gb|AAL86002.1 unknown protein {Ar... 53 2e-07
BF631811 similar to GP|9759350|dbj WD-repeat protein-like {Arabi... 52 6e-07
TC87605 homologue to PIR|G96563|G96563 probable coatomer complex... 51 7e-07
BG586181 similar to GP|4689316|gb| peroxisomal targeting signal ... 50 1e-06
TC78145 similar to GP|15294240|gb|AAK95297.1 AT3g18860/MCB22_3 {... 49 4e-06
>TC88481 similar to PIR|T00806|T02445 probable U4/U6 small nuclear
ribonucleoprotein [imported] - Arabidopsis thaliana,
partial (60%)
Length = 1836
Score = 79.0 bits (193), Expect = 3e-15
Identities = 42/121 (34%), Positives = 60/121 (48%)
Frame = +1
Query: 238 RGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNAL 297
+GH + F SG+Y+ T S D+ ++W +ET L GH + L + +L
Sbjct: 1192 KGHLERLARIAFHPSGKYLGTASYDKTWRLWDVETEEELLLQEGHSRSVYGLDFHHDGSL 1371
Query: 298 VASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDA 357
AS D + RVW L G + L GH ++ I+FSP Y L + +D TCRIWD
Sbjct: 1372 AASCGLDALARVWDLRTGRSVLALEGHVKSILGISFSPNG---YHLATGGEDNTCRIWDL 1542
Query: 358 R 358
R
Sbjct: 1543 R 1545
Score = 63.9 bits (154), Expect = 1e-10
Identities = 32/120 (26%), Positives = 57/120 (46%)
Frame = +1
Query: 239 GHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALV 298
GH +VY F G + D L ++W + T S+ + GHV I ++ S N +
Sbjct: 1321 GHSRSVYGLDFHHDGSLAASCGLDALARVWDLRTGRSVLALEGHVKSILGISFSPNGYHL 1500
Query: 299 ASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDAR 358
A+ D R+W L + + H+ ++ + F P+ Y L+++S D T ++W +R
Sbjct: 1501 ATGGEDNTCRIWDLRKKKSLYTIPAHSNLISQVKFEPQEG--YFLVTASYDMTAKVWSSR 1674
Score = 51.2 bits (121), Expect = 7e-07
Identities = 37/159 (23%), Positives = 68/159 (42%), Gaps = 2/159 (1%)
Frame = +1
Query: 204 GLPRHHRAPSIRAACYAIAKPSTM-VQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDD 262
GL HH S+ A+C A ++ +++ + GH ++ F +G ++ TG +D
Sbjct: 1342 GLDFHHDG-SLAASCGLDALARVWDLRTGRSVLALEGHVKSILGISFSPNGYHLATGGED 1518
Query: 263 RLVKIWSMETAYSLASCRGHVGDITDLAVSSNNA-LVASSSNDYIIRVWRLPDGLPISVL 321
+IW + SL + H I+ + + ++S D +VW D P+ L
Sbjct: 1519 NTCRIWDLRKKKSLYTIPAHSNLISQVKFEPQEGYFLVTASYDMTAKVWSSRDFKPVKTL 1698
Query: 322 RGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYT 360
GH VT++ +++ S D T ++W T
Sbjct: 1699 SGHEAKVTSLDVLGDGG---YIVTVSHDRTIKLWSXXTT 1806
Score = 49.3 bits (116), Expect = 3e-06
Identities = 33/109 (30%), Positives = 52/109 (47%), Gaps = 1/109 (0%)
Frame = +1
Query: 249 FDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS-NNALVASSSNDYII 307
F R G+ + T S K+WSM +++ +GH TD+A S + +A++S D
Sbjct: 973 FSRDGKGLATCSFTGATKLWSMPNVKKVSTLKGHTQRATDVAYSPVHKNHLATASADRTA 1152
Query: 308 RVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
+ W G + +GH + IAF P L ++S D T R+WD
Sbjct: 1153 KYWN-DQGALLGTFKGHLERLARIAFHPSGK---YLGTASYDKTWRLWD 1287
Score = 47.0 bits (110), Expect = 1e-05
Identities = 46/196 (23%), Positives = 79/196 (39%), Gaps = 4/196 (2%)
Frame = +1
Query: 225 STMVQKMQNIKRI---RGHCN-AVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCR 280
+T + M N+K++ +GH A A ++ T S DR K W+ + A L + +
Sbjct: 1018 ATKLWSMPNVKKVSTLKGHTQRATDVAYSPVHKNHLATASADRTAKYWNDQGAL-LGTFK 1194
Query: 281 GHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAV 340
GH+ + +A + + ++S D R+W + + + GH+ +V + F +
Sbjct: 1195 GHLERLARIAFHPSGKYLGTASYDKTWRLWDVETEEELLLQEGHSRSVYGLDFHHDGSLA 1374
Query: 341 YQLLSSSDDGTCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNAN 400
S D R+WD R TGRS + I +F+ N
Sbjct: 1375 ---ASCGLDALARVWDLR----------------TGRSVLALEGHV---KSILGISFSPN 1488
Query: 401 GTVFVTGSSDNLARVY 416
G TG DN R++
Sbjct: 1489 GYHLATGGEDNTCRIW 1536
Score = 40.8 bits (94), Expect = 0.001
Identities = 25/92 (27%), Positives = 47/92 (50%), Gaps = 3/92 (3%)
Frame = +1
Query: 268 WSMETAYSLASCRGHVGD---ITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGH 324
W+++ A +L +GD +T + S + +A+ S ++W +P+ +S L+GH
Sbjct: 895 WTLKQAANLNLEFSEIGDDRPLTGCSFSRDGKGLATCSFTGATKLWSMPNVKKVSTLKGH 1074
Query: 325 TGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
T T +A+SP L ++S D T + W+
Sbjct: 1075TQRATDVAYSPVHK--NHLATASADRTAKYWN 1164
>TC78188 similar to PIR|C84870|C84870 probable splicing factor [imported] -
Arabidopsis thaliana, partial (95%)
Length = 1436
Score = 67.4 bits (163), Expect = 1e-11
Identities = 47/184 (25%), Positives = 78/184 (41%), Gaps = 1/184 (0%)
Frame = +1
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSME-TAYSLASCRGHVGDITDLAVS 292
I + GH +AVY F+ +G V +GS D+ + +W++ + +GH + DL +
Sbjct: 292 IMLLTGHQSAVYTMKFNPTGSVVASGSHDKEIFLWNVHGDCKNFMVLKGHKNAVLDLHWT 471
Query: 293 SNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
S+ + S+S D +R+W G I + H V + P ++S SDDGT
Sbjct: 472 SDGTQIISASPDKTLRLWDTETGKQIKKMVEHLSYVNSCC--PTRRGPPLVVSGSDDGTA 645
Query: 353 RIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSDNL 412
++WD R S T P +QI +F+ TG DN
Sbjct: 646 KLWDMRQRGSI--------------------QTFPDKYQITAVSFSDASDKIYTGGIDND 765
Query: 413 ARVY 416
+++
Sbjct: 766 VKIW 777
Score = 50.1 bits (118), Expect = 2e-06
Identities = 46/186 (24%), Positives = 79/186 (41%), Gaps = 1/186 (0%)
Frame = +1
Query: 232 QNIKRIRGHCNAVY-CAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLA 290
+ IK++ H + V C R V++GSDD K+W M S+ + IT ++
Sbjct: 541 KQIKKMVEHLSYVNSCCPTRRGPPLVVSGSDDGTAKLWDMRQRGSIQTFPDKY-QITAVS 717
Query: 291 VSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDG 350
S + + + D +++W L G L+GH +T++ S P+ Y L + D
Sbjct: 718 FSDASDKIYTGGIDNDVKIWDLRKGEVTMTLQGHQDMITSMQLS--PDGSYLLTNGMDCK 891
Query: 351 TCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSD 410
C IWD R Y P+ + G N + C ++ +G+ GSSD
Sbjct: 892 LC-IWDM-------RPYAPQ-NRCVKILEGHQHNF---EKNLLKCGWSPDGSKVTAGSSD 1035
Query: 411 NLARVY 416
+ ++
Sbjct: 1036RMVYIW 1053
Score = 47.8 bits (112), Expect = 8e-06
Identities = 31/128 (24%), Positives = 56/128 (43%), Gaps = 1/128 (0%)
Frame = +1
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAV 291
+N ++GH NAV + G +I+ S D+ +++W ET + H+ +
Sbjct: 415 KNFMVLKGHKNAVLDLHWTSDGTQIISASPDKTLRLWDTETGKQIKKMVEHLSYVNSCCP 594
Query: 292 SSNN-ALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDG 350
+ LV S S+D ++W + I +TA++FS + +Y + D
Sbjct: 595 TRRGPPLVVSGSDDGTAKLWDMRQRGSIQTFPDKY-QITAVSFSDASDKIY---TGGIDN 762
Query: 351 TCRIWDAR 358
+IWD R
Sbjct: 763 DVKIWDLR 786
Score = 37.0 bits (84), Expect = 0.014
Identities = 23/76 (30%), Positives = 33/76 (43%), Gaps = 5/76 (6%)
Frame = +1
Query: 234 IKRIRGHC-----NAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITD 288
+K + GH N + C + G V GS DR+V IW + L GH G + +
Sbjct: 937 VKILEGHQHNFEKNLLKCG-WSPDGSKVTAGSSDRMVYIWDTTSRRILYKLPGHNGSVNE 1113
Query: 289 LAVSSNNALVASSSND 304
N +V S S+D
Sbjct: 1114 CVFHPNEPIVGSCSSD 1161
Score = 36.2 bits (82), Expect = 0.024
Identities = 27/116 (23%), Positives = 48/116 (41%), Gaps = 8/116 (6%)
Frame = +1
Query: 249 FDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIR 308
F + + TG D VKIW + + +GH IT + +S + + + ++ D +
Sbjct: 718 FSDASDKIYTGGIDNDVKIWDLRKGEVTMTLQGHQDMITSMQLSPDGSYLLTNGMDCKLC 897
Query: 309 VWRL----PDGLPISVLRGH----TGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
+W + P + +L GH + +SP + V + S D IWD
Sbjct: 898 IWDMRPYAPQNRCVKILEGHQHNFEKNLLKCGWSPDGSKV---TAGSSDRMVYIWD 1056
>BG457241 homologue to PIR|F84600|F8 coatomer alpha subunit [imported] -
Arabidopsis thaliana, partial (15%)
Length = 654
Score = 63.5 bits (153), Expect = 1e-10
Identities = 47/169 (27%), Positives = 74/169 (42%)
Frame = +1
Query: 188 TKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAKPSTMVQKMQNIKRIRGHCNAVYCA 247
TK+N+V GL H + P I A+ ++ + I + H V
Sbjct: 91 TKSNRVKGLSF---------HPKRPWILASLHSGVIQLWDYRMGTLIDKFDEHDGPVRGV 243
Query: 248 IFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYII 307
F S ++G DD +K+W+ + L + GH+ I + + + S+S+D I
Sbjct: 244 HFHNSQPLFVSGGDDYKIKVWNYKLHRCLFTLLGHLDYIRTVQFHHESPWIVSASDDQTI 423
Query: 308 RVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
R+W ISVL GH V F P+ + V +S+S D T R+WD
Sbjct: 424 RIWNWQSRTCISVLTGHNHYVMCALFHPKDDLV---VSASLDQTVRVWD 561
Score = 30.8 bits (68), Expect = 1.0
Identities = 18/63 (28%), Positives = 29/63 (45%)
Frame = +1
Query: 208 HHRAPSIRAACYAIAKPSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKI 267
HH +P I +A Q I + GH + V CA+F V++ S D+ V++
Sbjct: 376 HHESPWIVSASDDQTIRIWNWQSRTCISVLTGHNHYVMCALFHPKDDLVVSASLDQTVRV 555
Query: 268 WSM 270
W +
Sbjct: 556 WDI 564
>TC85659 SP|O24076|GBLP_MEDSA Guanine nucleotide-binding protein beta
subunit-like protein. [Alfalfa] {Medicago sativa},
complete
Length = 1261
Score = 63.2 bits (152), Expect = 2e-10
Identities = 47/181 (25%), Positives = 84/181 (45%), Gaps = 4/181 (2%)
Frame = +2
Query: 240 HCNAVYCAIFDRSGRY--VITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNAL 297
H + V C F S +++ S DR VK+W++ + GH G + +AVS + +L
Sbjct: 503 HSDWVSCVRFSPSTLQPTIVSASWDRTVKVWNLTNCKLRNTLAGHSGYVNTVAVSPDGSL 682
Query: 298 VASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDA 357
AS D +I +W L +G + L + + A+ FSP L ++ + + +IWD
Sbjct: 683 CASGGKDGVILLWDLAEGKRLYSLDAGS-IIHALCFSPN----RYWLCAATESSIKIWDL 847
Query: 358 RYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFN--ANGTVFVTGSSDNLARV 415
L V +++ G +T + I+C + N A+G+ +G +D + RV
Sbjct: 848 ESKSIVEDLKVDLKTEADAAIGG---DTTTKKKVIYCTSLNWSADGSTLFSGYTDGVVRV 1018
Query: 416 Y 416
+
Sbjct: 1019W 1021
Score = 59.3 bits (142), Expect = 3e-09
Identities = 32/124 (25%), Positives = 62/124 (49%), Gaps = 2/124 (1%)
Frame = +2
Query: 235 KRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSN 294
+R+ GH + V + G++ ++GS D +++W + S GH D+ +A S +
Sbjct: 230 RRLTGHSHFVQDVVLSSDGQFALSGSWDGELRLWDLNAGTSARRFVGHTKDVLSVAFSID 409
Query: 295 NALVASSSNDYIIRVWRLPDGLPISVLRG--HTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
N + S+S D I++W ++ G H+ V+ + FSP ++S+S D T
Sbjct: 410 NRQIVSASRDRTIKLWNTLGECKYTIQDGDAHSDWVSCVRFSP-STLQPTIVSASWDRTV 586
Query: 353 RIWD 356
++W+
Sbjct: 587 KVWN 598
>TC79322 similar to GP|13605815|gb|AAK32893.1 At2g32700/F24L7.16
{Arabidopsis thaliana}, partial (59%)
Length = 1824
Score = 63.2 bits (152), Expect = 2e-10
Identities = 35/124 (28%), Positives = 60/124 (48%), Gaps = 1/124 (0%)
Frame = +2
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
+ IR V C F G+ + + D+ V +W+MET + ++ H ITD+
Sbjct: 779 VSSIRKSNGKVVCCHFSSDGKLLASAGHDKKVVLWNMETLKTQSTPEEHTVIITDVRFRP 958
Query: 294 NNALVASSSNDYIIRVWRLPD-GLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
N+ +A+SS D +R+W D + + GHT V ++ F P+ N ++ S D+
Sbjct: 959 NSTQLATSSFDTTVRLWDAADPSVSLQAYSGHTSHVASLDFHPKKNDLF--CSCDDNNEI 1132
Query: 353 RIWD 356
R W+
Sbjct: 1133RFWN 1144
Score = 42.0 bits (97), Expect = 4e-04
Identities = 31/130 (23%), Positives = 63/130 (47%), Gaps = 2/130 (1%)
Frame = +2
Query: 228 VQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSL--ASCRGHVGD 285
V+ + + ++GH V+C +D +G Y+ + S + VK+WS+ + + + G++
Sbjct: 1262 VETDRQMHSLQGHSGEVHCVCWDTNGDYLASVSQES-VKVWSLASGDCIHELNSSGNMFH 1438
Query: 286 ITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLS 345
S +N LV + +W + + +++ H G ++A+A SP V S
Sbjct: 1439 SCVFHPSYSNLLVIGGYQS--LELWHMAENKCMTI-PAHEGVISALAQSPVTGMV---AS 1600
Query: 346 SSDDGTCRIW 355
+S D + +IW
Sbjct: 1601 ASHDKSVKIW 1630
>TC86168 homologue to GP|13936812|gb|AAK49947.1 TGF-beta
receptor-interacting protein 1 {Phaseolus vulgaris},
complete
Length = 1322
Score = 60.1 bits (144), Expect = 2e-09
Identities = 50/191 (26%), Positives = 82/191 (42%), Gaps = 12/191 (6%)
Frame = +2
Query: 238 RGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSL---------ASCRGHVGDITD 288
RGH AV+C R +ITGS D+ K+W+++T L S VGD
Sbjct: 314 RGHNGAVWCCDVSRDSGRLITGSADQTAKLWNVQTGQQLFTFNFDSPARSVDFSVGD--K 487
Query: 289 LAVSSNNALVASSSNDYIIRVWRLPD---GLPISVLRGHTGAVTAIAFSPRPNAVYQLLS 345
LAV + + + SS ++ R+ + P + V++G G + + P + +S
Sbjct: 488 LAVITTDPFMELSSAIHVKRIAKDPSDQTAESLLVIKGPQGRINRAIWGPLNKTI---IS 658
Query: 346 SSDDGTCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFV 405
+ +D RIWD+ TG+ S + I A +A+G+ F
Sbjct: 659 AGEDAVIRIWDS----------------ETGKLLKESDKELGHKKTITSLAKSADGSHFC 790
Query: 406 TGSSDNLARVY 416
TGS D A+++
Sbjct: 791 TGSLDKSAKIW 823
Score = 37.4 bits (85), Expect = 0.011
Identities = 22/79 (27%), Positives = 38/79 (47%)
Frame = +2
Query: 280 RGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNA 339
+GH +T L + + L+ S + D+ VW +G + RGH GAV S
Sbjct: 188 KGHERPLTFLKYNRDGDLLFSCAKDHNPTVWFADNGERLGTYRGHNGAVWCCDVSRDSG- 364
Query: 340 VYQLLSSSDDGTCRIWDAR 358
+L++ S D T ++W+ +
Sbjct: 365 --RLITGSADQTAKLWNVQ 415
>TC79342 similar to GP|10177096|dbj|BAB10430. Notchless protein homolog
{Arabidopsis thaliana}, partial (47%)
Length = 873
Score = 59.3 bits (142), Expect = 3e-09
Identities = 39/123 (31%), Positives = 58/123 (46%), Gaps = 3/123 (2%)
Frame = +3
Query: 237 IRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNA 296
I GH AV F GR + +GS D V+ W + T + +C GH + +A S +
Sbjct: 483 ISGHGEAVLSVAFSPDGRQLASGSGDTTVRFWDLGTQTPMYTCTGHKNWVLCIAWSPDGK 662
Query: 297 LVASSSNDYIIRVWRLPDGLPI-SVLRGHTGAVTAIAFSP-RPNA-VYQLLSSSDDGTCR 353
+ S S + W G + + L GH +T I++ P NA + +S+S DG R
Sbjct: 663 YLVSGSMSGELICWDPQTGKQLGNALTGHKKWITGISWEPVHLNAPCRRFVSASKDGDAR 842
Query: 354 IWD 356
IWD
Sbjct: 843 IWD 851
Score = 40.0 bits (92), Expect = 0.002
Identities = 26/80 (32%), Positives = 35/80 (43%)
Frame = +3
Query: 277 ASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPR 336
A+ GH + +A S + +AS S D +R W L P+ GH V IA+SP
Sbjct: 477 ATISGHGEAVLSVAFSPDGRQLASGSGDTTVRFWDLGTQTPMYTCTGHKNWVLCIAWSPD 656
Query: 337 PNAVYQLLSSSDDGTCRIWD 356
L+S S G WD
Sbjct: 657 GK---YLVSGSMSGELICWD 707
>TC78038 weakly similar to GP|21592898|gb|AAM64848.1 unknown {Arabidopsis
thaliana}, partial (60%)
Length = 1986
Score = 58.5 bits (140), Expect = 5e-09
Identities = 48/196 (24%), Positives = 84/196 (42%), Gaps = 13/196 (6%)
Frame = +3
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLA-----------SCRGH 282
+K+ GH AV C +F +I+G + V++WS+ + S H
Sbjct: 516 LKKWHGHNTAVTCLVFSEDDSLLISGFKNGSVRVWSLLMIFDDVRRREASKIYEYSFSDH 695
Query: 283 VGDITDLAVSSN--NALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAV 340
+ D+ + NA++AS+S+D +VW L +G+ + + AIA P + +
Sbjct: 696 TLCVNDVVIGYGGCNAIIASASDDRTCKVWTLSNGM-LQRNIVFPSKIKAIALDPAEHVL 872
Query: 341 YQLLSSSDDGTCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNAN 400
Y + S+DG + T T +D +G SN S + C A++A
Sbjct: 873 Y---AGSEDGKIFVAALNTTRIT-------TNDQVSYITGSFSN---HSKAVTCLAYSAA 1013
Query: 401 GTVFVTGSSDNLARVY 416
++GS D + RV+
Sbjct: 1014ENFLISGSDDGIVRVW 1061
Score = 47.8 bits (112), Expect = 8e-06
Identities = 45/190 (23%), Positives = 80/190 (41%), Gaps = 22/190 (11%)
Frame = +3
Query: 250 DRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRV 309
+ SG Y++ G + +W +ET L GH +T L S +++L+ S + +RV
Sbjct: 438 NHSGTYLVGGGLSGEIYLWEVETGRLLKKWHGHNTAVTCLVFSEDDSLLISGFKNGSVRV 617
Query: 310 WRLPDGLPI--SVLRGHTGAVTAIAFSPRPNAVYQLL-----------SSSDDGTCRIWD 356
W L L I V R + +FS V ++ S+SDD TC++W
Sbjct: 618 WSL---LMIFDDVRRREASKIYEYSFSDHTLCVNDVVIGYGGCNAIIASASDDRTCKVW- 785
Query: 357 ARYTHSTPRLYVPKPSDSTGRSSGPSSNTM---PQSHQIFCCAFNA------NGTVFVTG 407
++ + + PS + P+ + + + +IF A N + ++TG
Sbjct: 786 -TLSNGMLQRNIVFPSKIKAIALDPAEHVLYAGSEDGKIFVAALNTTRITTNDQVSYITG 962
Query: 408 SSDNLARVYT 417
S N ++ T
Sbjct: 963 SFSNHSKAVT 992
Score = 37.4 bits (85), Expect = 0.011
Identities = 20/77 (25%), Positives = 37/77 (47%), Gaps = 4/77 (5%)
Frame = +3
Query: 219 YAIAKPSTMVQKMQNIKRIRG----HCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAY 274
+ A +T + + I G H AV C + + ++I+GSDD +V++W+ T
Sbjct: 900 FVAALNTTRITTNDQVSYITGSFSNHSKAVTCLAYSAAENFLISGSDDGIVRVWNASTHN 1079
Query: 275 SLASCRGHVGDITDLAV 291
+ + G IT++ V
Sbjct: 1080IIRVFKHAKGPITNILV 1130
Score = 31.6 bits (70), Expect = 0.59
Identities = 19/70 (27%), Positives = 31/70 (44%)
Frame = +3
Query: 286 ITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLS 345
IT LA + + + I +W + G + GH AVT + FS + L+S
Sbjct: 420 ITPLASNHSGTYLVGGGLSGEIYLWEVETGRLLKKWHGHNTAVTCLVFSEDDSL---LIS 590
Query: 346 SSDDGTCRIW 355
+G+ R+W
Sbjct: 591 GFKNGSVRVW 620
>TC78016 similar to GP|21537191|gb|AAM61532.1 PRL1 protein {Arabidopsis
thaliana}, partial (81%)
Length = 1759
Score = 58.5 bits (140), Expect = 5e-09
Identities = 30/104 (28%), Positives = 47/104 (44%)
Frame = +1
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAV 291
+N + I GH V D S + TGS DR +KIW + + + GH+ + LA+
Sbjct: 559 KNYRVISGHLGWVRSVAVDPSNTWFATGSADRTIKIWDLASGVLKLTLTGHIEQVRGLAI 738
Query: 292 SSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSP 335
S + + S+ +D ++ W L I GH V +A P
Sbjct: 739 SHKHTYMFSAGDDKQVKCWDLEQNKVIRSYHGHLSGVYCLAIHP 870
Score = 40.8 bits (94), Expect = 0.001
Identities = 28/87 (32%), Positives = 45/87 (51%), Gaps = 1/87 (1%)
Frame = +3
Query: 230 KMQNIKRIRGHCNAVYCAIFDR-SGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITD 288
KMQ I + GH N V C++F R + V+TGS D +K+W + ++++ H +
Sbjct: 936 KMQ-IHALSGHENTV-CSVFTRPTDPQVVTGSHDSTIKMWDLRYGKTMSTLTNHKKSVRA 1109
Query: 289 LAVSSNNALVASSSNDYIIRVWRLPDG 315
+A AS+S D I+ + LP G
Sbjct: 1110 MAQHPKEQAFASASADN-IKKFTLPKG 1187
Score = 37.7 bits (86), Expect = 0.008
Identities = 18/73 (24%), Positives = 34/73 (45%)
Frame = +3
Query: 267 IWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTG 326
+W + + + + GH + + + V + S+D I++W L G +S L H
Sbjct: 918 VWDIRSKMQIHALSGHENTVCSVFTRPTDPQVVTGSHDSTIKMWDLRYGKTMSTLTNHKK 1097
Query: 327 AVTAIAFSPRPNA 339
+V A+A P+ A
Sbjct: 1098SVRAMAQHPKEQA 1136
>CB893690 weakly similar to GP|9759025|dbj gene_id:K6A12.9~similar to unknown
protein~sp|O15736 {Arabidopsis thaliana}, partial (59%)
Length = 916
Score = 45.1 bits (105), Expect(2) = 6e-09
Identities = 21/76 (27%), Positives = 41/76 (53%), Gaps = 1/76 (1%)
Frame = +2
Query: 236 RIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNN 295
R+ H +F+ + +ITG D VK+W T +S G +G + DL ++++N
Sbjct: 260 RLNAHEGGCASLLFEYNSSKLITGGQDHSVKVWDTNTGSLSSSLTGCLGSVLDLTITNDN 439
Query: 296 -ALVASSSNDYIIRVW 310
+++A+SS++ + W
Sbjct: 440 RSVIAASSSNNLYPYW 487
Score = 41.2 bits (95), Expect = 7e-04
Identities = 34/132 (25%), Positives = 59/132 (43%), Gaps = 2/132 (1%)
Frame = +3
Query: 227 MVQKMQNIKRIRGHCNAVYCA--IFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVG 284
++Q Q + GH + V CA I S R V++ + D +K+W + Y +
Sbjct: 450 LLQAAQTTYTLTGHTDKV-CAVDISKVSSRLVVSAAYDGTIKVWDLMKGYCTNTIM-FPS 623
Query: 285 DITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLL 344
L S++ + S D +R+W + G +S + H+ AVT+I+ S N L
Sbjct: 624 ICNALCFSTDGQTIFSGHVDGNLRLWDIQSGRLLSEVATHSHAVTSISLSRNGNIA---L 794
Query: 345 SSSDDGTCRIWD 356
+S D ++D
Sbjct: 795 TSGGDNLHNLFD 830
Score = 32.7 bits (73), Expect(2) = 6e-09
Identities = 26/118 (22%), Positives = 47/118 (39%), Gaps = 22/118 (18%)
Frame = +3
Query: 321 LRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTHSTPRLYVPKPSDS------ 374
L GHT V A+ S + + ++S++ DGT ++WD + T + P ++
Sbjct: 480 LTGHTDKVCAVDISKVSSRL--VVSAAYDGTIKVWDLMKGYCTNTIMFPSICNALCFSTD 653
Query: 375 -----TGRSSGP-----------SSNTMPQSHQIFCCAFNANGTVFVTGSSDNLARVY 416
+G G S SH + + + NG + +T DNL ++
Sbjct: 654 GQTIFSGHVDGNLRLWDIQSGRLLSEVATHSHAVTSISLSRNGNIALTSGGDNLHNLF 827
>TC84162 similar to GP|15810485|gb|AAL07130.1 putative WD40-repeat protein
{Arabidopsis thaliana}, partial (29%)
Length = 628
Score = 57.0 bits (136), Expect = 1e-08
Identities = 30/103 (29%), Positives = 48/103 (46%)
Frame = +1
Query: 256 VITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDG 315
V +GS DR +W + S+ +GH I + S + V ++S D IR+W + DG
Sbjct: 31 VCSGSQDRTACVWRLPDLVSVVVFKGHKRGIWSVEFSPVDQCVVTASGDKTIRIWAISDG 210
Query: 316 LPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDAR 358
+ GHT +V F R Q++S DG ++W +
Sbjct: 211 SCLKTFEGHTSSVLRALFVTRGT---QIVSCGADGLVKLWTVK 330
Score = 46.2 bits (108), Expect = 2e-05
Identities = 24/60 (40%), Positives = 34/60 (56%)
Frame = +1
Query: 296 ALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIW 355
+LV S S D VWRLPD + + V +GH + ++ FSP V +++S D T RIW
Sbjct: 25 SLVCSGSQDRTACVWRLPDLVSVVVFKGHKRGIWSVEFSPVDQCV---VTASGDKTIRIW 195
Score = 45.8 bits (107), Expect = 3e-05
Identities = 22/118 (18%), Positives = 47/118 (39%)
Frame = +1
Query: 238 RGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNAL 297
+GH ++ F + V+T S D+ ++IW++ L + GH + +
Sbjct: 103 KGHKRGIWSVEFSPVDQCVVTASGDKTIRIWAISDGSCLKTFEGHTSSVLRALFVTRGTQ 282
Query: 298 VASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIW 355
+ S D ++++W + ++ H V A+A + L + D +W
Sbjct: 283 IVSCGADGLVKLWTVKSNECVATYDNHEDKVWALAVGRKTET---LATGGSDAVVNLW 447
Score = 45.4 bits (106), Expect = 4e-05
Identities = 20/77 (25%), Positives = 41/77 (52%)
Frame = +1
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
+K GH ++V A+F G +++ D LVK+W++++ +A+ H + LAV
Sbjct: 217 LKTFEGHTSSVLRALFVTRGTQIVSCGADGLVKLWTVKSNECVATYDNHEDKVWALAVGR 396
Query: 294 NNALVASSSNDYIIRVW 310
+A+ +D ++ +W
Sbjct: 397 KTETLATGGSDAVVNLW 447
>TC80645 similar to PIR|T50983|T50983 probable pleiotropic regulator 1
(PLRG1) [imported] - Neurospora crassa, partial (41%)
Length = 1141
Score = 56.6 bits (135), Expect = 2e-08
Identities = 43/184 (23%), Positives = 72/184 (38%)
Frame = +2
Query: 233 NIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVS 292
N+ + GH V + VITGS D V++W + S+ H + LA
Sbjct: 56 NVHVLSGHTGTVSDLTCQEADPQVITGSLDATVRLWDLAAGKSMGVLTHHKKGVRALATH 235
Query: 293 SNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTC 352
AS S+ I+ W+ P+G + GH + ++ + R N L S D+G+
Sbjct: 236 PEEFTFASGSSG-SIKQWKCPEGAFMQNFEGHNAIINTLSVN-RDNV---LFSGGDNGSM 400
Query: 353 RIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSDNL 412
WD + H L S +G S+T F+ +G + G +D
Sbjct: 401 NFWDWKTGHRFQSLETTAQPGSLDAEAGIMSST-----------FDNSGLRLICGEADKT 547
Query: 413 ARVY 416
+++
Sbjct: 548 IKIW 559
>TC79118 similar to GP|16323188|gb|AAL15328.1 AT5g67320/K8K14_4 {Arabidopsis
thaliana}, partial (44%)
Length = 1098
Score = 56.2 bits (134), Expect = 2e-08
Identities = 43/129 (33%), Positives = 63/129 (48%), Gaps = 9/129 (6%)
Frame = +2
Query: 239 GHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVS-----S 293
GH V C +D +G + + SDD KIWS++ L R H +I + S +
Sbjct: 311 GHQGEVNCVKWDPTGTLLASCSDDITAKIWSVKQDKYLHDFREHSKEIYTIRWSPTGPGT 490
Query: 294 NNA----LVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDD 349
NN L+AS+S D +++W + G I L GH V ++AFS PN Y + S S D
Sbjct: 491 NNPNKKLLLASASFDSTVKLWDIELGKLIHSLNGHRHPVYSVAFS--PNGEY-IASGSLD 661
Query: 350 GTCRIWDAR 358
+ IW +
Sbjct: 662 KSLHIWSLK 688
Score = 48.5 bits (114), Expect = 5e-06
Identities = 36/176 (20%), Positives = 67/176 (37%), Gaps = 9/176 (5%)
Frame = +2
Query: 251 RSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVW 310
R+ + S D ++ + + + + GH G++ + L+AS S+D ++W
Sbjct: 221 RTNTSFASSSTDNMIYVCKIGENHPTQTFAGHQGEVNCVKWDPTGTLLASCSDDITAKIW 400
Query: 311 RLPDGLPISVLRGHTGAVTAIAFSP------RPNAVYQLLSSSDDGTCRIWD---ARYTH 361
+ + R H+ + I +SP PN L S+S D T ++WD + H
Sbjct: 401 SVKQDKYLHDFREHSKEIYTIRWSPTGPGTNNPNKKLLLASASFDSTVKLWDIELGKLIH 580
Query: 362 STPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSDNLARVYT 417
S H ++ AF+ NG +GS D +++
Sbjct: 581 S----------------------LNGHRHPVYSVAFSPNGEYIASGSLDKSLHIWS 682
Score = 42.4 bits (98), Expect = 3e-04
Identities = 22/69 (31%), Positives = 32/69 (45%)
Frame = +2
Query: 258 TGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLP 317
+ S D VK+W +E + S GH + +A S N +AS S D + +W L +G
Sbjct: 521 SASFDSTVKLWDIELGKLIHSLNGHRHPVYSVAFSPNGEYIASGSLDKSLHIWSLKEGKI 700
Query: 318 ISVLRGHTG 326
I G G
Sbjct: 701 IRTYNGSGG 727
Score = 36.2 bits (82), Expect = 0.024
Identities = 16/51 (31%), Positives = 27/51 (52%)
Frame = +2
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVG 284
I + GH + VY F +G Y+ +GS D+ + IWS++ + + G G
Sbjct: 575 IHSLNGHRHPVYSVAFSPNGEYIASGSLDKSLHIWSLKEGKIIRTYNGSGG 727
>BG647724 similar to GP|11141605|gb LEUNIG {Arabidopsis thaliana}, partial
(25%)
Length = 757
Score = 56.2 bits (134), Expect = 2e-08
Identities = 31/112 (27%), Positives = 56/112 (49%), Gaps = 1/112 (0%)
Frame = +2
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
+ +R N V C+ F G+ + +G D+ V +W ++ A+ H ITD+ S
Sbjct: 410 LNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSP 589
Query: 294 NNALVASSSNDYIIRVWRLPD-GLPISVLRGHTGAVTAIAFSPRPNAVYQLL 344
+ +A+SS D +RVW + + G + GH+ AV ++ F P + + L+
Sbjct: 590 SMPRLATSSYDKTVRVWDVDNPGYSLRTFTGHSAAVMSLDFHPNKDDLSALV 745
>TC82282 similar to PIR|T08544|T08544 hypothetical protein F27B13.70 -
Arabidopsis thaliana, partial (84%)
Length = 904
Score = 55.1 bits (131), Expect = 5e-08
Identities = 50/211 (23%), Positives = 90/211 (41%), Gaps = 16/211 (7%)
Frame = +2
Query: 210 RAPSIRAACYAIAKP---STMVQKMQNIKRI-RGHCNAVYCAIFDRSGR----YVITGSD 261
+A S+ + C K +T+ K+ IK I H ++V+ + + ++TGS
Sbjct: 11 KAASLHSLCSREQKSKEFTTVAMKLAGIKSIDNAHDDSVWAVTWAPATATRPPLLLTGSL 190
Query: 262 DRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVL 321
D V++W + + GH + +A ++ ASSS D +RV+ + I+ L
Sbjct: 191 DETVRLWKSDDLVLERTNTGHCLGVASVAAHPLGSIAASSSLDSFVRVFDVDSNATIATL 370
Query: 322 RGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRI-------WDARYTHSTPRLYVPKPSDS 374
V + F P+ +L+ + G+ + W+ T S PR+ PKPSD
Sbjct: 371 EAPPSEVWQMRFDPKG----AILAVAGGGSASVNLWDTSTWELVVTLSIPRVEGPKPSDK 538
Query: 375 TGRSSGPSSNT-MPQSHQIFCCAFNANGTVF 404
+G S P ++ C + + +VF
Sbjct: 539 SGSKKFVLSVAWSPDGKRLACGSMDGTISVF 631
Score = 35.4 bits (80), Expect = 0.041
Identities = 31/131 (23%), Positives = 54/131 (40%)
Frame = +2
Query: 286 ITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLS 345
+ +A S + +A S D I V+ + + L GH V ++ +SP + L S
Sbjct: 557 VLSVAWSPDGKRLACGSMDGTISVFDVQRAKFLHHLEGHFMPVRSLVYSPYDPRL--LFS 730
Query: 346 SSDDGTCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFV 405
+SDDG ++DA G++ + + + + C + NG
Sbjct: 731 ASDDGNVHMYDAE-----------------GKALXGTMSX--HASWVLCVDVSPNGAAIA 853
Query: 406 TGSSDNLARVY 416
TGSSD R++
Sbjct: 854 TGSSDKTVRLW 886
>TC82910 similar to GP|19347844|gb|AAL86002.1 unknown protein {Arabidopsis
thaliana}, partial (19%)
Length = 821
Score = 52.8 bits (125), Expect = 2e-07
Identities = 33/125 (26%), Positives = 62/125 (49%)
Frame = +1
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSS 293
+ + H +++ C F SG + G D++ IW +E + + + C G +
Sbjct: 148 VSTLPSHSSSIECVGFAPSGSWAAIGGMDKMT-IWDVEHSLARSICEHEYG--VTCSTWL 318
Query: 294 NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCR 353
+ VA+ SND +R+W G + RGH+ ++ +++ S N Y L+S+S D T R
Sbjct: 319 GTSYVATGSNDGAVRLWDSRSGECVRTFRGHSESIQSLSLS--ANQEY-LVSASLDHTAR 489
Query: 354 IWDAR 358
++D +
Sbjct: 490 VFDVK 504
>BF631811 similar to GP|9759350|dbj WD-repeat protein-like {Arabidopsis
thaliana}, partial (21%)
Length = 575
Score = 51.6 bits (122), Expect = 6e-07
Identities = 33/121 (27%), Positives = 58/121 (47%), Gaps = 9/121 (7%)
Frame = +3
Query: 251 RSGRYVITGSDDRLVKIWSMETAYSL-ASCRGHVGD---ITDLAVSSNNALVASSSNDYI 306
+ ++++ ++ + +W++E L +GH I A +AS S D
Sbjct: 63 KDNKFLLVNLLNQEINLWNIEGNPKLIGKYKGHRRSRFIIRSCFGGLEQAFIASGSEDSQ 242
Query: 307 IRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIW-----DARYTH 361
+ +W G PI L GH+GAV ++++P + L S+SDD T RIW + +Y H
Sbjct: 243 VYIWHRSSGEPIEALPGHSGAVNCVSWNPA--NPHMLASASDDRTIRIWGLNSLNKKYQH 416
Query: 362 S 362
+
Sbjct: 417 A 419
>TC87605 homologue to PIR|G96563|G96563 probable coatomer complex subunit
33791-27676 [imported] - Arabidopsis thaliana, partial
(23%)
Length = 765
Score = 51.2 bits (121), Expect = 7e-07
Identities = 31/114 (27%), Positives = 56/114 (48%), Gaps = 1/114 (0%)
Frame = +3
Query: 244 VYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSN 303
V A F ++V+ G+DD +++++ T + H I +AV V SSS+
Sbjct: 303 VRSAKFVARKQWVVAGADDMFIRVYNYNTMDKVKVFEAHTDYIRCVAVHPTLPYVLSSSD 482
Query: 304 DYIIRVWRLPDG-LPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWD 356
D +I++W G + + GH+ V + F+P+ + S+S D T +IW+
Sbjct: 483 DMLIKLWDWEKGWICTQIFEGHSHYVMQVTFNPKDTNTF--ASASLDRTIKIWN 638
Score = 40.0 bits (92), Expect = 0.002
Identities = 22/84 (26%), Positives = 38/84 (45%), Gaps = 2/84 (2%)
Frame = +3
Query: 231 MQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASC-RGHVGDITDL 289
M +K H + + C + YV++ SDD L+K+W E + GH + +
Sbjct: 390 MDKVKVFEAHTDYIRCVAVHPTLPYVLSSSDDMLIKLWDWEKGWICTQIFEGHSHYVMQV 569
Query: 290 AVSSNNA-LVASSSNDYIIRVWRL 312
+ + AS+S D I++W L
Sbjct: 570 TFNPKDTNTFASASLDRTIKIWNL 641
>BG586181 similar to GP|4689316|gb| peroxisomal targeting signal type 2
receptor {Arabidopsis thaliana}, partial (56%)
Length = 804
Score = 50.4 bits (119), Expect = 1e-06
Identities = 38/142 (26%), Positives = 66/142 (45%), Gaps = 11/142 (7%)
Frame = +1
Query: 233 NIKRIRGHCNAVYCAIFD-RSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAV 291
+++ + H VY ++++ R + S D V+IW + S H +I
Sbjct: 13 SVRTFKEHAYCVYSSVWNPRHADVFASASGDCTVRIWDVREPGSTMILPAHEHEILSCDW 192
Query: 292 SS-NNALVASSSNDYIIRVWRLPD-GLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDD 349
+ + ++A+ S D ++VW + +P++VL GH AV + FSP + L+S S D
Sbjct: 193 NKYDECIIATGSVDKSVKVWDVRSYRVPVAVLNGHGYAVRKVKFSPHVRNL--LVSCSYD 366
Query: 350 GTCRIWD--------ARYTHST 363
T +WD +RY H T
Sbjct: 367 MTVCVWDFMVEDALVSRYDHHT 432
>TC78145 similar to GP|15294240|gb|AAK95297.1 AT3g18860/MCB22_3 {Arabidopsis
thaliana}, partial (93%)
Length = 2633
Score = 48.9 bits (115), Expect = 4e-06
Identities = 43/162 (26%), Positives = 74/162 (45%)
Frame = +3
Query: 256 VITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDG 315
++TGS D +K W +T L + +GH + LAV S+ ++ S+S+D +R+W + G
Sbjct: 573 LVTGSSDTTLKTWKGKTC--LHTFQGHADTVRGLAVMSDLGIL-SASHDGSLRLWAV-SG 740
Query: 316 LPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTHSTPRLYVPKPSDST 375
+ + GHT AI +S +A ++S S+D +IW
Sbjct: 741 EGLMEMVGHT----AIVYSVDSHASGLIVSGSEDRFAKIW-------------------- 848
Query: 376 GRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSDNLARVYT 417
G ++ ++ F NG + VT SD +AR++T
Sbjct: 849 --KDGVCVQSIEHPGCVWDTKFMENGDI-VTACSDGIARIWT 965
Score = 48.1 bits (113), Expect = 6e-06
Identities = 29/103 (28%), Positives = 49/103 (47%)
Frame = +3
Query: 256 VITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDG 315
V++G D LV +W + + + S +GH +T + V + + SSS D +R WR +G
Sbjct: 336 VVSGGMDTLVLVWDLNSGEKVQSLKGHQLQVTGIVVDDGD--LVSSSMDCTLRRWR--NG 503
Query: 316 LPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDAR 358
I H G + A+ P +L++ S D T + W +
Sbjct: 504 QCIETWEAHKGPIQAVIKLP----TGELVTGSSDTTLKTWKGK 620
Score = 34.3 bits (77), Expect = 0.091
Identities = 23/79 (29%), Positives = 34/79 (42%)
Frame = +3
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAV 291
+ + + GH VY SG +++GS+DR KIW H G + D
Sbjct: 741 EGLMEMVGHTAIVYSVDSHASG-LIVSGSEDRFAKIWKDGVCVQSIE---HPGCVWDTKF 908
Query: 292 SSNNALVASSSNDYIIRVW 310
N +V + S D I R+W
Sbjct: 909 MENGDIVTACS-DGIARIW 962
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.134 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,408,101
Number of Sequences: 36976
Number of extensions: 253346
Number of successful extensions: 1590
Number of sequences better than 10.0: 141
Number of HSP's better than 10.0 without gapping: 1358
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1517
length of query: 417
length of database: 9,014,727
effective HSP length: 99
effective length of query: 318
effective length of database: 5,354,103
effective search space: 1702604754
effective search space used: 1702604754
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144539.12


