

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144539.1 + phase: 0
(163 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
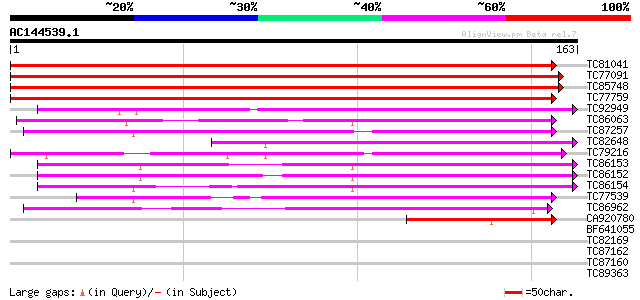
Score E
Sequences producing significant alignments: (bits) Value
TC81041 similar to GP|6714413|gb|AAF26101.1| unknown protein {Ar... 167 2e-42
TC77091 weakly similar to GP|11602751|emb|CAC18558. ENOD18 prote... 163 3e-41
TC85748 similar to GP|11692860|gb|AAG40033.1 AT3g53990 {Arabidop... 162 6e-41
TC77759 homologue to GP|11602747|emb|CAC18556. early nodulin ENO... 137 1e-33
TC92949 similar to GP|10998131|dbj|BAB03102. gene_id:MEC18.3~unk... 81 2e-16
TC86063 weakly similar to GP|16323382|gb|AAL15185.1 unknown prot... 80 5e-16
TC87257 similar to GP|16323382|gb|AAL15185.1 unknown protein {Ar... 78 1e-15
TC82648 similar to GP|10998131|dbj|BAB03102. gene_id:MEC18.3~unk... 73 6e-14
TC79216 similar to GP|5669654|gb|AAD46412.1| ER6 protein {Lycope... 67 3e-12
TC86153 similar to GP|15451080|gb|AAK96811.1 Unknown protein {Ar... 67 4e-12
TC86152 similar to GP|15451080|gb|AAK96811.1 Unknown protein {Ar... 65 9e-12
TC86154 similar to GP|15451080|gb|AAK96811.1 Unknown protein {Ar... 65 9e-12
TC77539 similar to GP|16118856|gb|AAL14629.1 growth-on protein G... 50 5e-07
TC86962 similar to GP|15320408|dbj|BAB63912. RD2 protein {Arabid... 42 1e-04
CA920780 similar to GP|21689829|gb| unknown protein {Arabidopsis... 40 4e-04
BF641055 similar to GP|20804872|dbj contains EST C19329(E10264)~... 39 0.001
TC82169 similar to GP|21703134|gb|AAM74507.1 AT4g27320/M4I22_130... 35 0.013
TC87162 homologue to PIR|S48698|S48698 3-dehydroquinate dehydrat... 32 0.15
TC87160 similar to PIR|S48698|S48698 3-dehydroquinate dehydratas... 31 0.19
TC89363 homologue to PIR|T01289|T01289 probable protein kinase A... 31 0.25
>TC81041 similar to GP|6714413|gb|AAF26101.1| unknown protein {Arabidopsis
thaliana}, partial (94%)
Length = 581
Score = 167 bits (423), Expect = 2e-42
Identities = 81/157 (51%), Positives = 108/157 (68%)
Frame = +2
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
MA A +G+AMDFSP S A +W VDN++ + D +I+I ++P +H +L+E TGSPL
Sbjct: 32 MAKAHIVGVAMDFSPTSKLALRWAVDNLINKNDQIIMINVQPPSADHTRKELFEDTGSPL 211
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PL E + K+Y I DPEV+ I TA + K V+ KVYWGD REKLC A+E + L
Sbjct: 212 VPLEELREINFTKQYGIAKDPEVIDILETASKIKGAKVVAKVYWGDPREKLCNAVEDLHL 391
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
D L +G+RGLGT++ ++GSVS +VV NASCPVTVVK
Sbjct: 392 DSLVIGSRGLGTIKSVLLGSVSKHVVTNASCPVTVVK 502
>TC77091 weakly similar to GP|11602751|emb|CAC18558. ENOD18 protein {Vicia
faba}, partial (67%)
Length = 896
Score = 163 bits (412), Expect = 3e-41
Identities = 77/159 (48%), Positives = 107/159 (66%)
Frame = +3
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
MAS R++G+A+DFS S A +W +DN+++ GD L ++ + LW TGSPL
Sbjct: 105 MASGRQIGVALDFSKGSKIALKWAIDNLLRTGDTLYIVHVNHSHPTESRNLLWATTGSPL 284
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PL EF ++ +YE+ D EVL I TA QK+V V+ KVYWGDAREK+ +++ + L
Sbjct: 285 IPLSEFREKNVVHQYEVDPDAEVLDILDTASRQKQVTVVGKVYWGDAREKIVDSVGDLKL 464
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSS 159
D L MG+RGLG ++R ++GSVS YV +NASCPVT+VK S
Sbjct: 465 DALVMGSRGLGAIQRVLLGSVSTYVTSNASCPVTIVKES 581
>TC85748 similar to GP|11692860|gb|AAG40033.1 AT3g53990 {Arabidopsis
thaliana}, complete
Length = 823
Score = 162 bits (410), Expect = 6e-41
Identities = 77/159 (48%), Positives = 108/159 (67%)
Frame = +1
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
MA R +G+A+DFS S A +W ++N+ +GDN+ +I I + + QLW GSPL
Sbjct: 58 MAKDRTIGVALDFSKSSKNALKWALENLADKGDNIYIIHISHDSLDEARNQLWAKDGSPL 237
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PL EF ++ KKY ++ D EVL + T QK+V V+ KVYWGDAREKL +A+E + L
Sbjct: 238 IPLKEFREPEIMKKYGVQIDIEVLDLLDTFSRQKEVNVVTKVYWGDAREKLMDAVEDLKL 417
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSS 159
D L MG+RGL T++R ++GSVSN+V+ NA CPVT+VK +
Sbjct: 418 DSLVMGSRGLSTIQRILLGSVSNFVMTNAPCPVTIVKDN 534
>TC77759 homologue to GP|11602747|emb|CAC18556. early nodulin ENOD18 {Vicia
faba}, complete
Length = 848
Score = 137 bits (346), Expect = 1e-33
Identities = 67/157 (42%), Positives = 101/157 (63%)
Frame = +1
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
M R++G+A+DFS S A +W + N+ +GD LI I + + + TGSPL
Sbjct: 70 MVEDRKIGVAIDFSKNSKNALKWAIVNMADKGDTFYLIHINSNSSDESRNKQFAKTGSPL 249
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
L E ++ KY ++TD EVL + T QK+V V+ K+YWGDAR+KL ++IE + L
Sbjct: 250 ISLEELKEVEVMSKYGVQTDVEVLDMLDTLATQKEVSVVAKLYWGDARQKLMDSIEDLKL 429
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
D L +G+RGL T++R ++GSVSN+V+ ++ CPVT+VK
Sbjct: 430 DALVLGSRGLSTIKRILLGSVSNFVMVHSPCPVTIVK 540
>TC92949 similar to GP|10998131|dbj|BAB03102. gene_id:MEC18.3~unknown
protein {Arabidopsis thaliana}, partial (45%)
Length = 727
Score = 80.9 bits (198), Expect = 2e-16
Identities = 52/164 (31%), Positives = 84/164 (50%), Gaps = 9/164 (5%)
Frame = +3
Query: 9 IAMDFSPCSIKAFQWTVDNIVK-----EGDNL----ILIIIRPEEYEHGEMQLWEVTGSP 59
+A+D S S A +W +DN+ EG + ++ ++ E H + G+
Sbjct: 102 VAIDESDGSFYALKWALDNLFNVMTTMEGTSSENEGMVFLVHVEPIFHNYVHPIGPGGAA 281
Query: 60 LTPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVP 119
P ++S KK + + +L A + K+V + GD RE +C+A EQ+
Sbjct: 282 FYPASVVVDS--VKKGQQERSAVILSRALQMCKDKQVKAESVILNGDPREMICQASEQMQ 455
Query: 120 LDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSSGQHH 163
+D L MG+RGLGTL+RA +GSVS+Y ++A P+ +VK HH
Sbjct: 456 VDLLIMGSRGLGTLKRAFLGSVSDYCAHHAKAPILIVKPPEDHH 587
>TC86063 weakly similar to GP|16323382|gb|AAL15185.1 unknown protein
{Arabidopsis thaliana}, partial (46%)
Length = 984
Score = 79.7 bits (195), Expect = 5e-16
Identities = 50/163 (30%), Positives = 85/163 (51%), Gaps = 8/163 (4%)
Frame = +2
Query: 3 SARRLGIAMDFSPCSIKAFQWTVDNIVKEG--DNLILIIIRPEEYEHGEMQLWEVTGSPL 60
+ RR+ +A+D SI A W + N+V + D+LIL+ ++P V S
Sbjct: 272 NGRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPR----------VVYSAF 421
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVV------VLVKVYWGDAREKLCEA 114
G +SD+ E + ++A +E+ K+V V ++ GD R+ +C+A
Sbjct: 422 DGTGYLFSSDITATMEKYSQ----QVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQA 589
Query: 115 IEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
++++ +D L MG+ G G ++RA +GSVSN+ N CPV +VK
Sbjct: 590 VQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 718
>TC87257 similar to GP|16323382|gb|AAL15185.1 unknown protein {Arabidopsis
thaliana}, partial (64%)
Length = 747
Score = 78.2 bits (191), Expect = 1e-15
Identities = 47/156 (30%), Positives = 81/156 (51%), Gaps = 3/156 (1%)
Frame = +3
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDN---LILIIIRPEEYEHGEMQLWEVTGSPLT 61
R++ +A+D S S+ A W++ N++ + +N L+L+ ++P + + +
Sbjct: 168 RKIMVAIDESEESMYALSWSISNLIADTNNNNKLVLLYVKPPSAVYSLDSAGYIFSNDTI 347
Query: 62 PLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLD 121
E +S L K + + T I +KVV GDA+ +C A +++ D
Sbjct: 348 DTLENYSSQLAKSVMKRAEAIYRNFDDTDINIEKVVGT-----GDAKNVICNAAKKLGAD 512
Query: 122 GLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
L MG+ G G ++RA++GSVS+Y V NA CPV +VK
Sbjct: 513 TLVMGSHGYGFIKRALLGSVSDYCVKNAKCPVVIVK 620
>TC82648 similar to GP|10998131|dbj|BAB03102. gene_id:MEC18.3~unknown
protein {Arabidopsis thaliana}, partial (48%)
Length = 814
Score = 72.8 bits (177), Expect = 6e-14
Identities = 40/111 (36%), Positives = 60/111 (54%), Gaps = 6/111 (5%)
Frame = +2
Query: 59 PLTPLGEFINSDLP------KKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLC 112
P+ P G +P KK + + + A + K+V + GDARE +C
Sbjct: 290 PMGPAGVAFYPSIPEVGDSLKKAQQEKSAAIASQALQMCKDKQVKAESVILNGDAREMIC 469
Query: 113 EAIEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSSGQHH 163
EA E++ +D L MG+RGLG L+RA +GSVS+Y ++A P+ +VK HH
Sbjct: 470 EATEKMQVDLLIMGSRGLGKLKRAFLGSVSDYCAHHAKAPILIVKPPEDHH 622
>TC79216 similar to GP|5669654|gb|AAD46412.1| ER6 protein {Lycopersicon
esculentum}, partial (86%)
Length = 1094
Score = 67.0 bits (162), Expect = 3e-12
Identities = 50/182 (27%), Positives = 82/182 (44%), Gaps = 22/182 (12%)
Frame = +3
Query: 1 MASARRLGI---AMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTG 57
M+ + LG+ A+D S S+ A +W ++N+ P+ + G + V
Sbjct: 75 MSESGNLGVVVVAVDGSEESMNALRWALENLKLRSP-------APDSTDAGSFIILHVQS 233
Query: 58 SPLT---------PLGEFINSDLP----------KKYEIKTDPEVLKIATTAIEQKKVVV 98
P P G + ++P K+ L I +T + KV
Sbjct: 234 PPSIATGLNPGSIPFGGPSDLEVPAFAAAIEAHQKRITDSIFDHALGICSTFNVKTKVRT 413
Query: 99 LVKVYWGDAREKLCEAIEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKS 158
V V GD +EK+CE ++ + D L MG+R G ++R +GSVSNY +++ CPVT++K
Sbjct: 414 HVVV--GDPKEKICETVQDLHADVLVMGSRAFGPIKRMFLGSVSNYCAHHSECPVTIIKG 587
Query: 159 SG 160
G
Sbjct: 588 KG 593
>TC86153 similar to GP|15451080|gb|AAK96811.1 Unknown protein {Arabidopsis
thaliana}, partial (74%)
Length = 784
Score = 66.6 bits (161), Expect = 4e-12
Identities = 44/164 (26%), Positives = 77/164 (46%), Gaps = 9/164 (5%)
Frame = +1
Query: 9 IAMDFSPCSIKAFQWTVDNIVKEGDNLI-LIIIRPEEYEHGEMQLWEVTGSPLTPLGEFI 67
+A+D S S A +WT+D+ +++ L+++ + L + + P+
Sbjct: 115 VAIDESDHSAYALKWTLDHFFSTNNSVFKLVLVHARPAATSSVGLAGPGAAEVLPI---- 282
Query: 68 NSDLPKKYEIKTDPEVLKIATTAIEQKKVV--------VLVKVYWGDAREKLCEAIEQVP 119
D ++ KIA E K + V+V+V GDAR LC+ +E+
Sbjct: 283 -----------VDSDLRKIAARVAENAKQLCIKKSVNDVIVEVVEGDARNVLCDTVEKYR 429
Query: 120 LDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSSGQHH 163
L +G+ G G ++RA++GSVS+Y ++A C V +VK H
Sbjct: 430 ASILVVGSHGYGAIKRAVLGSVSDYCAHHAHCTVMIVKKPKTKH 561
>TC86152 similar to GP|15451080|gb|AAK96811.1 Unknown protein {Arabidopsis
thaliana}, partial (51%)
Length = 834
Score = 65.5 bits (158), Expect = 9e-12
Identities = 45/157 (28%), Positives = 78/157 (49%), Gaps = 2/157 (1%)
Frame = +3
Query: 9 IAMDFSPCSIKAFQWTVDNIVKEGDNLI-LIIIRPEEYEHGEMQLWEVTGSPLTPLGEFI 67
+A+D S S A +WT+D+ +++ L+++ + L + + +
Sbjct: 99 VAIDESDHSAYALKWTLDHFFSTNNSVFKLVLVHARPAATSSVGLAGPVYAGAAEVLPIV 278
Query: 68 NSDLPKKYEIKTDPEVLKIATTAIEQKKVV-VLVKVYWGDAREKLCEAIEQVPLDGLTMG 126
+SDL K V + A +K V V+V+V GDAR LC+ +E+ L +G
Sbjct: 279 DSDLRK-----IAARVAENAKQLCIKKSVNDVIVEVVEGDARNVLCDTVEKYRASILVVG 443
Query: 127 NRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSSGQHH 163
+ G G ++RA++GSVS+Y ++A C V +VK H
Sbjct: 444 SHGYGAIKRAVLGSVSDYCAHHAHCTVMIVKKPKTKH 554
>TC86154 similar to GP|15451080|gb|AAK96811.1 Unknown protein {Arabidopsis
thaliana}, partial (75%)
Length = 1086
Score = 65.5 bits (158), Expect = 9e-12
Identities = 46/167 (27%), Positives = 82/167 (48%), Gaps = 12/167 (7%)
Frame = +1
Query: 9 IAMDFSPCSIKAFQWTVDNIVKEGDN----LILIIIRPEEYEHGEMQLWEVTGSPLTPLG 64
+ +D S S A +WT+D++V N L+L+ +P SP T +G
Sbjct: 331 VGVDDSEFSTYALEWTLDHLVTTLPNPIFKLVLVFAKP---------------SPSTNVG 465
Query: 65 EFINSDLPKKYEIKTDPEVLKIATTAIEQKKVV--------VLVKVYWGDAREKLCEAIE 116
F+ + + ++ + AT IE+ + + V+++V GDA LC+A++
Sbjct: 466 -FVGPAGAAEILPIVEADLKRTATIVIERAQEICTKRSVKDVIMEVVDGDATNVLCDAVD 642
Query: 117 QVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSSGQHH 163
+ L +G+ G G ++RA++GSVS+Y ++A C V +VK H
Sbjct: 643 KHHASILVVGSHGYGAIKRAVLGSVSDYCAHHAHCTVMIVKKPKPKH 783
>TC77539 similar to GP|16118856|gb|AAL14629.1 growth-on protein GRO10
{Euphorbia esula}, partial (44%)
Length = 2020
Score = 49.7 bits (117), Expect = 5e-07
Identities = 36/142 (25%), Positives = 65/142 (45%), Gaps = 4/142 (2%)
Frame = +1
Query: 20 AFQWTVDNIVKEGDN----LILIIIRPEEYEHGEMQLWEVTGSPLTPLGEFINSDLPKKY 75
AF WTV I++ + L L + P+E + ++ +G +F N K+
Sbjct: 1222 AFDWTVSKIIRNNVSAFHLLFLHVQVPDEDGYDDVDSIYASGE------DFKNM---KQQ 1374
Query: 76 EIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLTMGNRGLGTLRR 135
E +L+ + V + GD +E + +++V D L +G+RGLG ++
Sbjct: 1375 EKARGTHLLEYFVNRCNEIGVTCEAWIKQGDPKEVILNEVKRVRPDLLVVGSRGLGPFQK 1554
Query: 136 AIMGSVSNYVVNNASCPVTVVK 157
+G+VS + +A CPV +K
Sbjct: 1555 VFVGTVSEFCWKHAECPVMTIK 1620
>TC86962 similar to GP|15320408|dbj|BAB63912. RD2 protein {Arabidopsis
thaliana}, partial (88%)
Length = 913
Score = 42.0 bits (97), Expect = 1e-04
Identities = 34/153 (22%), Positives = 62/153 (40%), Gaps = 1/153 (0%)
Frame = +2
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLG 64
R + +A+D P S AF W + ++ + D + L+ H + T LT
Sbjct: 251 RDIVLAIDHGPNSKHAFDWALIHLCRLADTIHLV--------HAVSDVKNQTVYDLT--- 397
Query: 65 EFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLT 124
+ K+A A + V + ++ GDA + +C+ E++ +
Sbjct: 398 ---------------QGLMEKLAVEAFQVSMVKTVARIVQGDAGKVICKEAERIKPAAVV 532
Query: 125 MGNRGLGTLRRAIMGSVSNYVVNNA-SCPVTVV 156
+G RG + I GSV Y ++ + PV +V
Sbjct: 533 LGTRGRSLFQSVIQGSVGEYCFHHCKAAPVVIV 631
>CA920780 similar to GP|21689829|gb| unknown protein {Arabidopsis thaliana},
partial (27%)
Length = 780
Score = 40.0 bits (92), Expect = 4e-04
Identities = 19/46 (41%), Positives = 30/46 (64%), Gaps = 3/46 (6%)
Frame = -1
Query: 115 IEQVPLDGLTMGNRGLGTLRRAI---MGSVSNYVVNNASCPVTVVK 157
+E++ L + MG+RG G +R+ +GSVS+Y V + CPV VV+
Sbjct: 717 VERLGLSAVIMGSRGFGATKRSSNGKLGSVSDYCVRHCVCPVVVVR 580
>BF641055 similar to GP|20804872|dbj contains EST C19329(E10264)~similar to
Arabidopsis thaliana chromosome 1 T23J18.3~unknown
protein, partial (31%)
Length = 492
Score = 38.5 bits (88), Expect = 0.001
Identities = 22/63 (34%), Positives = 33/63 (51%)
Frame = +2
Query: 3 SARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTP 62
S R++ IA+D S S A +W V N ++ GD +IL+ +RP +G W SP
Sbjct: 179 SNRKVAIAVDLSDESAYAVRWAVQNYLRPGDTVILLHVRPTYVLYGAD--WGSVTSPTAD 352
Query: 63 LGE 65
G+
Sbjct: 353 GGD 361
>TC82169 similar to GP|21703134|gb|AAM74507.1 AT4g27320/M4I22_130
{Arabidopsis thaliana}, partial (50%)
Length = 600
Score = 35.0 bits (79), Expect = 0.013
Identities = 14/38 (36%), Positives = 25/38 (64%)
Frame = +3
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRP 42
R++G+A+D S S A +W V + ++ GD +IL+ + P
Sbjct: 186 RKIGVAVDLSDESAYAVRWAVQHYIRPGDAVILLHVSP 299
>TC87162 homologue to PIR|S48698|S48698 3-dehydroquinate dehydratase (EC
4.2.1.10) / shikimate 5-dehydrogenase (EC 1.1.1.25)
precursor -, partial (83%)
Length = 1054
Score = 31.6 bits (70), Expect = 0.15
Identities = 19/52 (36%), Positives = 27/52 (51%)
Frame = +1
Query: 79 TDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLTMGNRGL 130
T +++KIATTA+E V + ++ QVP GL MG+RGL
Sbjct: 454 TGADIVKIATTAVEITDVARMFEIM----------VHSQVPFIGLVMGDRGL 579
>TC87160 similar to PIR|S48698|S48698 3-dehydroquinate dehydratase (EC
4.2.1.10) / shikimate 5-dehydrogenase (EC 1.1.1.25)
precursor -, partial (56%)
Length = 832
Score = 31.2 bits (69), Expect = 0.19
Identities = 17/52 (32%), Positives = 28/52 (53%)
Frame = +2
Query: 79 TDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLTMGNRGL 130
T +++KIATTA+E V + ++ + + VP GL MG+RG+
Sbjct: 668 TGADIVKIATTAVEITDVARMFEI-------MVHSQVRHVPFIGLVMGDRGI 802
>TC89363 homologue to PIR|T01289|T01289 probable protein kinase At2g19410
[imported] - Arabidopsis thaliana, partial (1%)
Length = 911
Score = 30.8 bits (68), Expect = 0.25
Identities = 25/98 (25%), Positives = 41/98 (41%)
Frame = +3
Query: 17 SIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLGEFINSDLPKKYE 76
S +A QW ++N+V + D LIL+ + P ++T P TP G +I
Sbjct: 99 SRRALQWAMENVVPQADRLILVHVIP-----------KITSIP-TPAGGYIPIS------ 224
Query: 77 IKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEA 114
+ D ++QK + V +KLCE+
Sbjct: 225 -EADAHAFAAYVQDVKQKSEEIFVSF------KKLCES 317
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.135 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,431,910
Number of Sequences: 36976
Number of extensions: 47282
Number of successful extensions: 227
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 226
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 226
length of query: 163
length of database: 9,014,727
effective HSP length: 89
effective length of query: 74
effective length of database: 5,723,863
effective search space: 423565862
effective search space used: 423565862
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC144539.1


