

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144538.13 + phase: 0 /pseudo
(1574 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
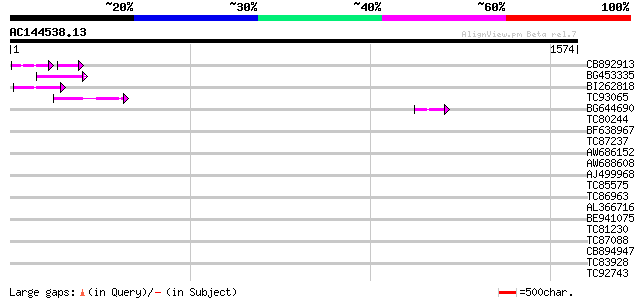
Score E
Sequences producing significant alignments: (bits) Value
CB892913 similar to GP|1769897|emb lectin receptor kinase {Arabi... 59 2e-15
BG453335 weakly similar to GP|18071369|g putative gag-pol polypr... 77 4e-14
BI262818 weakly similar to GP|6642775|gb| gag-pol polyprotein {V... 76 1e-13
TC93065 61 3e-09
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 50 7e-06
TC80244 weakly similar to GP|8777419|dbj|BAA97009.1 mitotic chec... 35 0.18
BF638967 similar to GP|8777419|dbj mitotic checkpoint protein-li... 35 0.23
TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-inf... 34 0.51
AW686152 weakly similar to PIR|T47645|T47 centromere protein-lik... 34 0.51
AW688608 34 0.51
AJ499968 weakly similar to GP|23495961|gb| hypothetical protein ... 33 0.67
TC85575 similar to PIR|D86410|D86410 protein F3M18.16 [imported]... 33 0.67
TC86963 similar to GP|15148918|gb|AAK84886.1 homeodomain leucine... 33 0.87
AL366716 similar to GP|17380890|gb| putative membrane transporte... 33 1.1
BE941075 33 1.1
TC81230 32 1.5
TC87088 similar to PIR|F86399|F86399 protein F17L21.22 [imported... 32 1.9
CB894947 32 1.9
TC83928 similar to GP|17134882|dbj|BAB77438. ORF_ID:all8519~hypo... 31 3.3
TC92743 similar to GP|9759091|dbj|BAB09660.1 contains similarity... 31 4.3
>CB892913 similar to GP|1769897|emb lectin receptor kinase {Arabidopsis
thaliana}, partial (35%)
Length = 837
Score = 58.5 bits (140), Expect(2) = 2e-15
Identities = 38/117 (32%), Positives = 59/117 (49%)
Frame = +3
Query: 4 GGSSNRPPLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLNEKKKEDWSEE 63
G +S + PL +G+ Y KM L + D +W VVE G DED K + D ++
Sbjct: 108 GAASFQVPLLKGSTYDNCFIKMMALLGAHD--VWEVVEKGYKESQDEDSLTKAQRDTLKD 281
Query: 64 EGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDI 120
KR + KA + AL +E++++ +AKE W+ LK +G VK+ R+ I
Sbjct: 282 SRKR---DKKALFLIYQALDEDEFEKISNATSAKEAWEKLKTSCQGEDKVKKVRLQI 443
Score = 43.1 bits (100), Expect(2) = 2e-15
Identities = 25/70 (35%), Positives = 37/70 (52%)
Frame = +1
Query: 134 ETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKKLRLE 193
E + + R I N+L+ + + KILR L + HIV I E KDL+ + +E
Sbjct: 472 ERVRVYFSRVISISNQLKRNSEKLEDVRIMEKILRSLDPKFEHIVEKIEETKDLETMTIE 651
Query: 194 DLIGSLKAHE 203
L GSL+A+E
Sbjct: 652 KLQGSLQAYE 681
>BG453335 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (11%)
Length = 660
Score = 77.4 bits (189), Expect = 4e-14
Identities = 44/142 (30%), Positives = 86/142 (59%), Gaps = 1/142 (0%)
Frame = +1
Query: 75 KLFLTM-ALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEMKET 133
+LFL +L + R+ + +KE + L+ +HEG V++ ++ RKF+L +M++
Sbjct: 139 ELFLIQ*SLDEGNFKRISKSTRSKEASNILEKYHEGDDKVEQIKLQSLRRKFKLMQMEDE 318
Query: 134 ETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKKLRLE 193
+ I + + ++N++++ + T + V KI+R L + IV AI E+KD+K L++E
Sbjct: 319 KKIADYISKLINVVNQIKAYGEVVTDQQIVEKIMRTLSPRFDFIVVAIHESKDVKTLKIE 498
Query: 194 DLIGSLKAHEVLLQEDKSSNKS 215
+L SL+AH++++ E +SS +S
Sbjct: 499 ELKSSLEAHKLMVYE-RSSERS 561
>BI262818 weakly similar to GP|6642775|gb| gag-pol polyprotein {Vitis
vinifera}, partial (24%)
Length = 615
Score = 75.9 bits (185), Expect = 1e-13
Identities = 43/144 (29%), Positives = 80/144 (54%), Gaps = 1/144 (0%)
Frame = +1
Query: 11 PLFEGTDYYYWKGKMELFLQS*DNHMWTVVENG-NYIPYDEDLNEKKKEDWSEEEGKRML 69
P+F+G +Y+ W ++E +L++ N +W VE +P ++ + ++ E + ++
Sbjct: 178 PVFDGENYHIWAARIEAYLEA--NDLWEAVEEDYEVLPLSDNPTMAQIKNHKERKTRKS- 348
Query: 70 LNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFE 129
KA+ L A+S E + R+ K+A EIW+ LK +EG ++ + +R+FE+ +
Sbjct: 349 ---KARATLFAAVSEEIFTRIMTIKSAFEIWNFLKTEYEGDERIRGMQALNLIREFEMQK 519
Query: 130 MKETETIDEMYGRFTIIMNELRSL 153
MKE+ETI E + I N++R L
Sbjct: 520 MKESETIKEYANKLISIANKVRLL 591
>TC93065
Length = 783
Score = 61.2 bits (147), Expect = 3e-09
Identities = 50/212 (23%), Positives = 87/212 (40%), Gaps = 5/212 (2%)
Frame = +2
Query: 123 RKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAIT 182
R+FE +MKETET+ E R + ++ ++R L +D + V KIL CLP + ++++
Sbjct: 221 REFEALKMKETETVREFSDRISKVVTQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLE 400
Query: 183 EAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQEL 242
E K+ ++ + +L+ +L+A E Q++ L
Sbjct: 401 ENKNFSEITVAELVNALQASE----------------------------------QRRSL 478
Query: 243 DEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDK-----SQVTCYGCYKLG 297
E++ + L K + + K+ P T LDK V C C +LG
Sbjct: 479 RMEENVEGAFLANNKGKNQSFKSFGKKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLG 658
Query: 298 HYKNECPLNKRKSSNLQTNQKSFMTTWDEPEE 329
H + C + +N Q + + E EE
Sbjct: 659 HVEKVC----KNKTNQQEQEARVVEHHQEDEE 742
Score = 32.7 bits (73), Expect = 1.1
Identities = 37/177 (20%), Positives = 73/177 (40%), Gaps = 21/177 (11%)
Frame = +2
Query: 384 NENLELRKEKEILEKENKDLIKNIQHSESKEVSENK--NNSQKENIILKENVLKLKNDIS 441
NE+ +LR+E E L+ + + ++ SK V++ + + ++++ ++ L
Sbjct: 200 NESAKLRREFEALKMKETETVREFSDRISKVVTQIRLLGEDLSDQRVVEKILVCLPEMFE 379
Query: 442 NFVKSTETFQKIMGSQVSVFDKACIGFKTNQKQKLYEN----FFIPKKNKNTCERY---- 493
+ S E + V+ A + + ++ EN F K KN +
Sbjct: 380 AKISSLEENKNFSEITVAELVNALQASEQRRSLRMEENVEGAFLANNKGKNQSFKSFGKK 559
Query: 494 ---KCTYCEKYGHLEPFCFRK---RKRLASQ-----KFKNNYTNFHKKETLNNTNHQ 539
C +C+K HL+ FC+ + + R +Q K N TN ++E +HQ
Sbjct: 560 KFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQEARVVEHHQ 730
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 50.1 bits (118), Expect = 7e-06
Identities = 42/100 (42%), Positives = 53/100 (53%), Gaps = 3/100 (3%)
Frame = -3
Query: 1125 PDW*LKVTINKKALTTMKPMLLWLDLKP*EF---FLHMHLISASNFFKWT*KVLS*MVF* 1181
P W K TI KK T M+ L + K EF LH S+ KW *+V M
Sbjct: 408 PSWWCKDTIKKKE*TMMRLFHLLPEWKLLEF**LLLHSW---GSSCTKWM*RVHLLMEIS 238
Query: 1182 MKRFMFINLLVLKI*KNQITCSN*PKLYMD*NKLQEPGMK 1221
+R + NLL LK+ + QI CS+* + YM *+KLQE GMK
Sbjct: 237 KRRCLSSNLLDLKMQRYQIMCSD*IRHYMV*SKLQEHGMK 118
>TC80244 weakly similar to GP|8777419|dbj|BAA97009.1 mitotic checkpoint
protein-like {Arabidopsis thaliana}, partial (35%)
Length = 1054
Score = 35.4 bits (80), Expect = 0.18
Identities = 58/251 (23%), Positives = 99/251 (39%), Gaps = 21/251 (8%)
Frame = +1
Query: 209 DKSSNKSKMIALKTNQE---SLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRR 265
D KS L T QE S +E + ++ D +D + I L +LQ +Q
Sbjct: 271 DTERKKSVEQLLYTEQELAASKGREKALQDQLLKEVTDSQDRLRKYIQLNTELQAKLQ-- 444
Query: 266 DQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWD 325
N+ N + E+ T + + G KLG+ S +Q +K
Sbjct: 445 --NETNLRIQAESHATSTQEKATSLEG--KLGNL----------SDTIQREKKRL----- 567
Query: 326 EPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNE 385
D+ + L K D+ ++ E+ E +N ++ML+++ +RL+++
Sbjct: 568 -------HDDHSQL----KKDSNFSISRITANLEQMECRANNAEREAEMLKEQVDRLKDQ 714
Query: 386 NLELRKEKEILEKEN-----------------KDLIKNIQHSESKEVSENKNNSQKENI- 427
E +K EK+ K L + +QH ES+ K S ENI
Sbjct: 715 FNECLHQKTEAEKKLATFSSQEVSSTWSDVLVKQLQQELQHYESEVREARKLRSNHENIE 894
Query: 428 ILKENVLKLKN 438
+LKE +L+ K+
Sbjct: 895 LLKEKLLEEKS 927
Score = 33.5 bits (75), Expect = 0.67
Identities = 37/165 (22%), Positives = 76/165 (45%), Gaps = 6/165 (3%)
Frame = +1
Query: 295 KLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNE-----E 349
+L + + E +K + LQ +T + +++ + Q + L AK NE +
Sbjct: 298 QLLYTEQELAASKGREKALQDQLLKEVT---DSQDRLRKYIQLNTELQAKLQNETNLRIQ 468
Query: 350 VNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQH 409
S+ EK L L S +++E +RL +++ +L+K+ + N++
Sbjct: 469 AESHATSTQEKATSLEGKLGNLSDTIQREKKRLHDDHSQLKKDSNFSISR---ITANLEQ 639
Query: 410 SESKEVSENKNNSQKENIILKENVLKLKNDISNFV-KSTETFQKI 453
E + NN+++E +LKE V +LK+ + + + TE +K+
Sbjct: 640 MECRA-----NNAEREAEMLKEQVDRLKDQFNECLHQKTEAEKKL 759
>BF638967 similar to GP|8777419|dbj mitotic checkpoint protein-like
{Arabidopsis thaliana}, partial (26%)
Length = 675
Score = 35.0 bits (79), Expect = 0.23
Identities = 19/78 (24%), Positives = 38/78 (48%)
Frame = +3
Query: 388 ELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKENIILKENVLKLKNDISNFVKST 447
E ++ E+L+ E + + ++ E + + EN I + +L+LK I+ K
Sbjct: 381 EAKQTIEVLQNELQKTKEKLKAVEELKSQSGETGKLVENYI-SDKILQLKEQIATLEKRE 557
Query: 448 ETFQKIMGSQVSVFDKAC 465
E ++ + ++SVF AC
Sbjct: 558 ERYKTVFADRISVFRXAC 611
>TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-infected
erythrocyte surface antigen {Plasmodium falciparum},
partial (2%)
Length = 2007
Score = 33.9 bits (76), Expect = 0.51
Identities = 54/260 (20%), Positives = 105/260 (39%), Gaps = 5/260 (1%)
Frame = +1
Query: 208 EDKS--SNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRR 265
E+KS N+ ++++E + E E N D ++ + + E+ +D +++ Q
Sbjct: 55 EEKSHLENEESKETGESSEEKSHLENEENKDEEKSKQENEEIKDG--------EKIQQEN 210
Query: 266 DQNKRNFPARKENAKT-ELDKSQVTCYGCYKLGHYKNECPLNK--RKSSNLQTNQKSFMT 322
++NK +++EN + + +KSQ +NE N+ K + T +KS
Sbjct: 211 EENKDEEKSQQENEENKDEEKSQ-----------QENELKKNEGGEKETGEITEEKS--- 348
Query: 323 TWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERL 382
+ E+T+E D +NEE N + + E+ + + E+
Sbjct: 349 --KQENEETSETNSKD------KENEESNQNGSDAKEQ--------------VGENHEQD 462
Query: 383 RNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKENIILKENVLKLKNDISN 442
+ E E EKE D IK S+++ KNN +E +N D +
Sbjct: 463 SKQGTEETNGTEGGEKEEHDKIKEDTSSDNQVQDGEKNNEAREENYSGDNASSAVVDNKS 642
Query: 443 FVKSTETFQKIMGSQVSVFD 462
S +T ++ + + F+
Sbjct: 643 QESSNKTEEQFDKKEKNEFE 702
>AW686152 weakly similar to PIR|T47645|T47 centromere protein-like -
Arabidopsis thaliana, partial (14%)
Length = 661
Score = 33.9 bits (76), Expect = 0.51
Identities = 43/185 (23%), Positives = 79/185 (42%), Gaps = 1/185 (0%)
Frame = +1
Query: 224 QESLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTEL 283
+E ++ LE++ + Q D ++++ + +L+ IQR ++ K + E EL
Sbjct: 97 EEKISYALEVSTHIRSQIADRASAKEELNRVKTELEIRIQRLEKEKNEMQSALEK---EL 267
Query: 284 DKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMA 343
D+ +KL Y++E + + L S E+ E + +M
Sbjct: 268 DRRSSDW--SFKLEKYQSEEQRLRERIRELAEQNVSLQREVSSFSERETESKS----VMT 429
Query: 344 KSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLE-LRKEKEILEKENKD 402
+D + L S EK + L + L+ C ++ EN + LR+ E EKE KD
Sbjct: 430 HTDQQLKVLT--SKAEKMKGEILGLQQNLSELQDRC-KIAEENRDCLRRNFEEKEKECKD 600
Query: 403 LIKNI 407
L K++
Sbjct: 601 LHKSV 615
>AW688608
Length = 678
Score = 33.9 bits (76), Expect = 0.51
Identities = 24/100 (24%), Positives = 48/100 (48%)
Frame = +3
Query: 366 DNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKE 425
+ L + +EQE E +R +N+ L+K+K L +EN+ L + + KE N S+
Sbjct: 42 EKLKLEKKTVEQEFENMREQNVILQKDKVELLEENRQLRIEVVNGVEKE-----NRSKST 206
Query: 426 NIILKENVLKLKNDISNFVKSTETFQKIMGSQVSVFDKAC 465
L+ +++L+ ++ + FQ+ G + + C
Sbjct: 207 LAALQAEMIELR-------QTNQVFQEENGKMLDEKNSLC 305
>AJ499968 weakly similar to GP|23495961|gb| hypothetical protein {Plasmodium
falciparum 3D7}, partial (19%)
Length = 342
Score = 33.5 bits (75), Expect = 0.67
Identities = 25/73 (34%), Positives = 37/73 (50%), Gaps = 3/73 (4%)
Frame = -3
Query: 378 ECERLRNENLEL-RKEKEILEKENKDLIKNIQH--SESKEVSENKNNSQKENIILKENVL 434
E ER+ E E RKEKE E+E ++ ++ + E KE E Q+E +LKE V+
Sbjct: 229 ELERIAKEKEERERKEKE--EREERERLEAVVRIEKERKEAEEKALKEQEEKALLKEGVI 56
Query: 435 KLKNDISNFVKST 447
K + K+T
Sbjct: 55 DKKEEEEPKSKAT 17
>TC85575 similar to PIR|D86410|D86410 protein F3M18.16 [imported] -
Arabidopsis thaliana, partial (15%)
Length = 1225
Score = 33.5 bits (75), Expect = 0.67
Identities = 49/218 (22%), Positives = 75/218 (33%), Gaps = 16/218 (7%)
Frame = +3
Query: 291 YGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEV 350
YG Y GH P ++N+ N +E N SDN+
Sbjct: 252 YGLY--GHDTGLHPPTTTNTNNVDPNTYQSYEKTSSHDEINNNKVNNYFNYQQDSDNKYP 425
Query: 351 N------LDPCSSCEKTEHLFDNLFYSS--------QMLEQECERLRNENLELRKEKEIL 396
N L P S + + +DN YSS + E+ N+N K
Sbjct: 426 NEVSDTKLTPTSYSSSSNNNYDNNKYSSSSDDFSNKKFPEEGYNSNENQNNNNNDNKYFY 605
Query: 397 EKENKDLIKNIQHSESKEVS-ENKNNSQKENIILKENVLKLKNDISNFVKSTETFQKIMG 455
NKD N + +E S EN+NN+ + N K+ SN E + +
Sbjct: 606 NNNNKDSSSNTKFTEEGYNSMENRNNNNNKYFYNNNN----KDSSSNTKFPEEGYNSMQN 773
Query: 456 SQVSVFDKACIGFKTNQKQKLYENF-FIPKKNKNTCER 492
+ ++K + N K + + + F N N ER
Sbjct: 774 QNNNNYEK----YNYNNKVAVNDKYSFKSNNNYNNGER 875
>TC86963 similar to GP|15148918|gb|AAK84886.1 homeodomain leucine zipper
protein HDZ2 {Phaseolus vulgaris}, partial (87%)
Length = 1519
Score = 33.1 bits (74), Expect = 0.87
Identities = 22/84 (26%), Positives = 39/84 (46%), Gaps = 2/84 (2%)
Frame = +1
Query: 365 FDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHS--ESKEVSENKNNS 422
+DNL + L++E L+N+ + KEK E ++ N H+ + ++ N+
Sbjct: 958 YDNLLQENDKLKEEVNSLKNKLIPRDKEKVNSEDKSSPEAINSPHNNIDPMDIISITNSE 1137
Query: 423 QKENIILKENVLKLKNDISNFVKS 446
+ L VLK K + +N KS
Sbjct: 1138 NGSKMSLPNMVLKCKQEDANSAKS 1209
>AL366716 similar to GP|17380890|gb| putative membrane transporter protein
{Arabidopsis thaliana}, partial (16%)
Length = 474
Score = 32.7 bits (73), Expect = 1.1
Identities = 36/120 (30%), Positives = 50/120 (41%), Gaps = 8/120 (6%)
Frame = +3
Query: 387 LELRKEKEILEKENKDLIKNIQHS-ESKEVSENKNNSQKENIILK-ENVLKLKNDISNFV 444
+ +RKEK +E E L K +H+ ES + +N S K I K E L L IS F
Sbjct: 24 ISIRKEKTFVEFELGHL-KICRHAIESNQCPKNHFASHKSKFIEKQEQFLVLIGLISTFK 200
Query: 445 KSTE----TFQKIMGSQVSVFDKACIGF--KTNQKQKLYENFFIPKKNKNTCERYKCTYC 498
K T + Q+ GF + N KQK Y ++ K C C++C
Sbjct: 201 KGFSARILTPSSLPPQQLHFIKCQPFGFWNQKNHKQKRYNSY---NSKKQECRTSPCSFC 371
>BE941075
Length = 651
Score = 32.7 bits (73), Expect = 1.1
Identities = 16/47 (34%), Positives = 27/47 (57%)
Frame = +2
Query: 375 LEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNN 421
L E + + +N +LRK+ +L KEN+DL+ +Q S++ NN
Sbjct: 185 LYMENQTIIEQNEKLRKQAMLLHKENQDLMSQLQMKLSEQNINTNNN 325
>TC81230
Length = 958
Score = 32.3 bits (72), Expect = 1.5
Identities = 48/215 (22%), Positives = 83/215 (38%), Gaps = 19/215 (8%)
Frame = +1
Query: 12 LFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYD-----EDLNEKKKEDWSEEEGK 66
+ G++Y +W M FL+ +W V P D K E+W + +
Sbjct: 235 ILNGSNYNHWAESMCGFLKG--RRLWRYVTGDKKCPTKGKDDTADAFADKLEEW-DSKNH 405
Query: 67 RMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHH--EGTSHVKETRIDIGVRK 124
+++ F+ ++ + ++ NAKE+WD LK + SH + D+ K
Sbjct: 406 QIITWFRNTSIPSIHMQFGRFE------NAKEVWDHLKQRYTISDLSHQYQLLKDLSNLK 567
Query: 125 FELFEMKETETIDEMYGRFTIIMNELRSLE---------KDFTIH-ERVRKI--LRCLPR 172
+ + + E + +I N+L S E K + H RVR I L L
Sbjct: 568 -----QQSGQPVYEFLAQMEVIWNQLTSCEPSLKDATDMKTYETHRNRVRLIQFLMALTD 732
Query: 173 SWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQ 207
+ + + L LE+ + LK+ E LQ
Sbjct: 733 EYEPVRASSLHQNPLP--TLENALPCLKSEETRLQ 831
>TC87088 similar to PIR|F86399|F86399 protein F17L21.22 [imported] -
Arabidopsis thaliana, partial (4%)
Length = 2997
Score = 32.0 bits (71), Expect = 1.9
Identities = 24/122 (19%), Positives = 53/122 (42%), Gaps = 1/122 (0%)
Frame = +3
Query: 302 ECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDP-CSSCEK 360
+ P + S NQ SF TW + E N + + + D + P ++ +
Sbjct: 156 DAPFTDQSSMGGMVNQSSFRDTWTD-EYGINRHFNPNQRVGSLEDQFLSRIGPNFNNFDV 332
Query: 361 TEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKN 420
+HL ++ +Q+ ERL+ + E R +++ E+ + + +Q +++ + + N
Sbjct: 333 ADHLMLQKLQKERLQQQQAERLQQQQAE-RLQQQQAERLQQQQAERLQQQQAERLQQQTN 509
Query: 421 NS 422
S
Sbjct: 510 IS 515
>CB894947
Length = 658
Score = 32.0 bits (71), Expect = 1.9
Identities = 27/122 (22%), Positives = 55/122 (44%), Gaps = 3/122 (2%)
Frame = +3
Query: 334 DEQADLCLMAKSDNE--EVNLDPCSSCEKTEHLFDNLFYSS-QMLEQECERLRNENLELR 390
+ AD C++A ++++ C K+++L + SS + E L ++ +
Sbjct: 96 ESSADDCMIADEHIHAPSMSIEDCKPVLKSKYLINQCSPSSLEDFELSIGSLEEDSEDAN 275
Query: 391 KEKEILEKENKDLIKNIQHSESKEVSENKNNSQKENIILKENVLKLKNDISNFVKSTETF 450
+ KE ++ + Q ESK S K S N+ + E L+ ++DIS ++ E
Sbjct: 276 RSKEARIYGDRVCLSETQ-GESKHFSMRKKVSSFNNVTVVEQDLQFRSDISAHAEAEEVV 452
Query: 451 QK 452
++
Sbjct: 453 EE 458
>TC83928 similar to GP|17134882|dbj|BAB77438. ORF_ID:all8519~hypothetical
protein {Nostoc sp. PCC 7120}, partial (2%)
Length = 588
Score = 31.2 bits (69), Expect = 3.3
Identities = 22/57 (38%), Positives = 31/57 (53%), Gaps = 2/57 (3%)
Frame = +2
Query: 213 NKSKMIALKTNQESLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQ--RRDQ 267
NKS I+ KTN QE+E +QE+++E QD+ K+Q I+ RRDQ
Sbjct: 8 NKSLHISQKTNNSETEQEVEQEV---EQEVEQEVEQDEGESNCEKIQVDIELARRDQ 169
>TC92743 similar to GP|9759091|dbj|BAB09660.1 contains similarity to unknown
protein~gb|AAF47871.1~gene_id:MJJ3.6 {Arabidopsis
thaliana}, partial (14%)
Length = 662
Score = 30.8 bits (68), Expect = 4.3
Identities = 21/79 (26%), Positives = 34/79 (42%)
Frame = -1
Query: 498 CEKYGHLEPFCFRKRKRLASQKFKNNYTNFHKKETLNNTNHQGPKKIWVPKALLTSTAGM 557
C + GH P C R ++ F+ + TNFH + +N +H WV +
Sbjct: 635 CGQDGH-GPGCQRSKQ*CPXVIFQRSTTNFHHRNEVNPKDHH-----WVDSPIY------ 492
Query: 558 SHNSQEKALVLGQWMLKTY 576
H ++ K L L W ++ Y
Sbjct: 491 -HRNKSKFLGLNFWDIQLY 438
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.355 0.157 0.539
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,886,696
Number of Sequences: 36976
Number of extensions: 697543
Number of successful extensions: 8961
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 2877
Number of HSP's successfully gapped in prelim test: 525
Number of HSP's that attempted gapping in prelim test: 5840
Number of HSP's gapped (non-prelim): 3893
length of query: 1574
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1465
effective length of database: 4,984,343
effective search space: 7302062495
effective search space used: 7302062495
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 65 (29.6 bits)
Medicago: description of AC144538.13


