

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
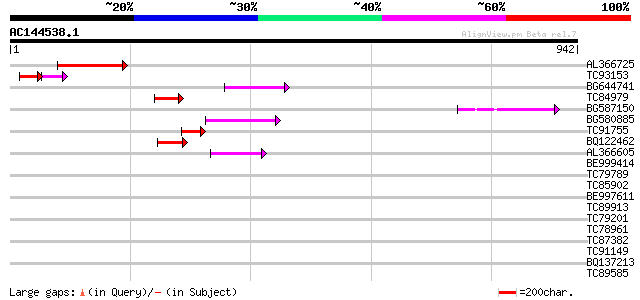
Query= AC144538.1 - phase: 0 /pseudo
(942 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AL366725 96 5e-20
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 73 4e-15
BG644741 76 7e-14
TC84979 74 2e-13
BG587150 similar to GP|13592175|gb ppg3 {Leishmania major}, part... 60 4e-09
BG580885 60 5e-09
TC91755 weakly similar to PIR|G75077|G75077 hypothetical protein... 56 7e-08
BQ122462 55 1e-07
AL366605 48 2e-05
BE999414 38 0.021
TC79789 similar to PIR|D96519|D96519 mysoin-like protein 11013-... 36 0.061
TC85902 similar to SP|Q9XF97|RL4_PRUAR 60S ribosomal protein L4 ... 35 0.18
BE997611 33 0.67
TC89913 similar to PIR|H96569|H96569 unknown protein 54928-5675... 31 2.5
TC79201 weakly similar to GP|10800070|dbj|BAB16490. contains EST... 30 3.3
TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 30 3.3
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 30 4.3
TC91149 similar to GP|15081723|gb|AAK82516.1 T10F18_70/T10F18_70... 29 7.4
BQ137213 29 9.7
TC89585 similar to GP|21553818|gb|AAM62911.1 unknown {Arabidopsi... 29 9.7
>AL366725
Length = 485
Score = 96.3 bits (238), Expect = 5e-20
Identities = 65/117 (55%), Positives = 75/117 (63%)
Frame = +3
Query: 80 SSIRTILPRLLSFQSVSNLITVCVLRLREQLVTRRSVSFLSW*AVAEFMRRIPRLITRL* 139
SSIRT+L RLLS ++ S+ V R Q T S L * AE RRI RL+TRL*
Sbjct: 3 SSIRTMLLRLLSSRNASSSRMV*GPTSRGQSDTNSSEFSLI**IPAESTRRILRLMTRL* 182
Query: 140 MSEKPKVSRVVLSLTVLLLIRVNREWLMRGALERKMLLLRLSAILVVRKATRVMLVL 196
MS +P+ +RV LSL V LLIR NREW M G L R+MLL RLS + RKATRVM VL
Sbjct: 183 MSGRPRANRVALSLIVPLLIRANREWSMIGVLRRRMLLRRLSVSTMARKATRVMFVL 353
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 72.8 bits (177), Expect(2) = 4e-15
Identities = 31/38 (81%), Positives = 34/38 (88%)
Frame = +2
Query: 17 WWVSLLPTLEQDGAVVTWALFRREFLDRYFSEDVRGKK 54
WW+SLLP LEQD AVVTWA+FR+EFL RYF EDVRGKK
Sbjct: 275 WWISLLPVLEQDDAVVTWAMFRKEFLGRYFPEDVRGKK 388
Score = 27.3 bits (59), Expect(2) = 4e-15
Identities = 17/43 (39%), Positives = 24/43 (55%)
Frame = +3
Query: 53 KKRSSSLT*SKETCQ*QSMLPILWSWQSSIRTILPRLLSFQSV 95
+KR + +*++ TC QSML LW+ S + R LS SV
Sbjct: 384 RKRLNFWS*NRVTCLSQSMLQSLWNLPHSTLITVRRQLSSPSV 512
>BG644741
Length = 735
Score = 75.9 bits (185), Expect = 7e-14
Identities = 39/109 (35%), Positives = 56/109 (50%)
Frame = -2
Query: 357 LHLFPFSLRDRASAWFHSLEVGSITSWDDMRRAFLARFSPPSKTAKLRDQITRFNQKDGE 416
L +FP SL A WF L SI +W+ +R FLAR+ P SK D++ F GE
Sbjct: 587 LRVFPLSLMGEADIWFTELPYNSIFTWNQLRDVFLARYYPVSKKLNHNDRVNNFVALPGE 408
Query: 417 SLYEA*ERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAAGGA 465
S+ + +RF LR P+H ++ + FY G K+ +D AGG+
Sbjct: 407 SVSSSWDRFTSFLRSVPNHRIDDDSLKEYFYRGQDDNNKVVLDTIAGGS 261
>TC84979
Length = 641
Score = 74.3 bits (181), Expect = 2e-13
Identities = 41/49 (83%), Positives = 42/49 (85%)
Frame = +3
Query: 241 **VI*YLFLYVI*FSFK*ILFYFILVLFPF*VSLLQIALYYISGTNIGW 289
**VI YLFLYVI*F+F *I YFILVLFPF* SLLQIA YYISG NIGW
Sbjct: 495 **VIYYLFLYVI*FNFN*I*VYFILVLFPF*DSLLQIAFYYISGMNIGW 641
>BG587150 similar to GP|13592175|gb ppg3 {Leishmania major}, partial (1%)
Length = 754
Score = 60.1 bits (144), Expect = 4e-09
Identities = 50/170 (29%), Positives = 78/170 (45%)
Frame = +2
Query: 744 EAQFAKFLNILKRICIKIPFAEALSRMPLYAKFLREIFSKKKAIDHKETIALTRESSAIV 803
E + F + I ++ PF EA L+ F RE ++ I A + I
Sbjct: 149 EKEMESFQKRVFMIPLEKPFEEAYFTHRLWM-FFRETRETEEDIRRMFCEAREKMKKRIT 325
Query: 804 KKQPQKLRDPGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFKRMGIGELKPTEMILKL 863
K K D G FAI + ALC+ GASV +LP + +G+ ++ P++
Sbjct: 326 LK---KKSDSGKFAISWTMKGIEFPHALCNTGASVIILPRDMADHLGL-KVDPSKESFTF 493
Query: 864 ADRSTIPLAGYIEDIPVKIEGIYIPTDFVVVDIEEDHDVPIILGRPFLAT 913
D S G I D+ V+I +P DF V+DI+ + + ++L R FL+T
Sbjct: 494 VDCSQRSSGGIIRDLEVQIGNALVPVDFHVLDIKINWNSSLLLRRAFLST 643
>BG580885
Length = 609
Score = 59.7 bits (143), Expect = 5e-09
Identities = 35/127 (27%), Positives = 66/127 (51%), Gaps = 2/127 (1%)
Frame = +3
Query: 326 FSGSPTEDPNLHISSFLRLSGTIKENQEAVRLHLFPFSLRDRASAWFH-SLEVGSIT-SW 383
F G+P E P H+S F ++ + ++ +FP +L + ++ W+ ++E I+ SW
Sbjct: 54 FRGTPNESPITHLSRFNKVCRANNASSVEMQKKIFPVTLEEESALWYDLNIEPYYISLSW 233
Query: 384 DDMRRAFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLIV 443
D+++ +FL + +LR ++ +Q + E + R + +L+ P HGLE +I
Sbjct: 234 DEIKLSFLQAYYEIEPVEELRSELMGIHQGEKERVRSYFLRLQWILKRWPEHGLEDDVIK 413
Query: 444 HTFYNGL 450
F NGL
Sbjct: 414 GVFVNGL 434
>TC91755 weakly similar to PIR|G75077|G75077 hypothetical protein PAB1697 -
Pyrococcus abyssi (strain Orsay), partial (4%)
Length = 746
Score = 55.8 bits (133), Expect = 7e-08
Identities = 27/40 (67%), Positives = 29/40 (72%)
Frame = -3
Query: 286 NIGWIESWSKRKGLGADLVRKYEDSETKIRLPKVRSLARP 325
NIGW ESWSKRKG GAD +K+ED ETK K SLARP
Sbjct: 333 NIGWRESWSKRKGFGADWTQKHEDWETKYISHKSESLARP 214
>BQ122462
Length = 757
Score = 55.1 bits (131), Expect = 1e-07
Identities = 29/50 (58%), Positives = 35/50 (70%)
Frame = +1
Query: 246 YLFLYVI*FSFK*ILFYFILVLFPF*VSLLQIALYYISGTNIGWIESWSK 295
YLF I*+ F *IL FI ++FPF*+ LLQ+ + Y NIGWIESWSK
Sbjct: 610 YLFYP*I*YCFI*ILVIFIFIIFPF*IILLQLHILYFK-NNIGWIESWSK 756
>AL366605
Length = 422
Score = 47.8 bits (112), Expect = 2e-05
Identities = 26/93 (27%), Positives = 50/93 (52%)
Frame = -3
Query: 334 PNLHISSFLRLSGTIKENQEAVRLHLFPFSLRDRASAWFHSLEVGSITSWDDMRRAFLAR 393
P HI ++R G K+N +++ +H F SL + A+ W+ SL I ++D++ AF +
Sbjct: 291 PQNHIIKYVRKMGNYKDN-DSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSH 115
Query: 394 FSPPSKTAKLRDQITRFNQKDGESLYEA*ERFK 426
+ ++ R+ + +QK ES E +R++
Sbjct: 114 YGFNTRLKPNREFLRSLSQKKEESFREYAQRWR 16
>BE999414
Length = 613
Score = 37.7 bits (86), Expect = 0.021
Identities = 22/65 (33%), Positives = 34/65 (51%)
Frame = +3
Query: 331 TEDPNLHISSFLRLSGTIKENQEAVRLHLFPFSLRDRASAWFHSLEVGSITSWDDMRRAF 390
T DP+ H+ + + + T + + AV+ LF +LR A WF +L SI SW D+ F
Sbjct: 27 T*DPDEHMEN-IEVVLTYRSVRGAVKCKLFVTTLRRGAMTWFKNLRRNSIGSWGDLCHEF 203
Query: 391 LARFS 395
F+
Sbjct: 204 TTHFT 218
>TC79789 similar to PIR|D96519|D96519 mysoin-like protein 11013-7318
[imported] - Arabidopsis thaliana, partial (21%)
Length = 1097
Score = 36.2 bits (82), Expect = 0.061
Identities = 39/160 (24%), Positives = 72/160 (44%), Gaps = 5/160 (3%)
Frame = +3
Query: 577 RPNQNFYKNPQGSYGQTAPPGYTNNQRVAQKSSLEILLENCMMNQNKQLQELKNQTGSLN 636
+PNQ+ YK P +Y Q + Y++ LE+ + ++ Q L+++ LN
Sbjct: 312 QPNQDVYKKP--NYVQISVESYSHLTG----------LEDQVKTYEEKAQTLEDEINELN 455
Query: 637 DSLSKLNTKVDS----IATHTKMLETQISQVAQQVAISSQTPGVFPGQTETNPKAHVNAI 692
+ LS NT++++ + H K+ E +S + A + T + A A
Sbjct: 456 EKLSAANTEINTKEALVKQHAKVAEEAVSGWEKAEAEALALKNSLESVTLSKLTAEDQAS 635
Query: 693 SLGGNKLEETITKAKSVKGEIVKLLGEKNAIKT-PLDKNK 731
L G L+E + + +++K E + + + KT LDK K
Sbjct: 636 QLDG-ALKECMRQIRNLKEEHELKIQDISLAKTKQLDKIK 752
>TC85902 similar to SP|Q9XF97|RL4_PRUAR 60S ribosomal protein L4 (L1).
[Apricot] {Prunus armeniaca}, complete
Length = 1571
Score = 34.7 bits (78), Expect = 0.18
Identities = 28/102 (27%), Positives = 51/102 (49%), Gaps = 2/102 (1%)
Frame = +1
Query: 703 ITKAKSVKGEIVKLLG--EKNAIKTPLDKNKTLNPLRLTKLNLEAQFAKFLNILKRICIK 760
+ +AK V ++ +++ E ++ P+ K+ + + L+ K LN++ R+
Sbjct: 931 LPRAKMVNSDLTRIINSDEVQSVVRPIKKD-------VKRATLKKNPLKNLNVMLRLN-- 1083
Query: 761 IPFAEALSRMPLYAKFLREIFSKKKAIDHKETIALTRESSAI 802
P+A+ RM L A+ R + SKK+ D K I E+SAI
Sbjct: 1084 -PYAKTAKRMALLAEAER-VKSKKEKFDKKRKIVSKEEASAI 1203
>BE997611
Length = 547
Score = 32.7 bits (73), Expect = 0.67
Identities = 20/78 (25%), Positives = 39/78 (49%)
Frame = -2
Query: 721 NAIKTPLDKNKTLNPLRLTKLNLEAQFAKFLNILKRICIKIPFAEALSRMPLYAKFLREI 780
NA++ DK +T + + + +F +ILK + +P E + ++P KFL+E+
Sbjct: 495 NAVEVAEDKPQTEDA*E--EKEVPPKFNALHHILKGYKVNMPITETVEQIPCCIKFLQEL 322
Query: 781 FSKKKAIDHKETIALTRE 798
+ +E I+L+ E
Sbjct: 321 LKTNANLSEEEFISLSSE 268
>TC89913 similar to PIR|H96569|H96569 unknown protein 54928-56750
[imported] - Arabidopsis thaliana, partial (37%)
Length = 761
Score = 30.8 bits (68), Expect = 2.5
Identities = 21/88 (23%), Positives = 42/88 (46%), Gaps = 3/88 (3%)
Frame = +2
Query: 731 KTLNPL---RLTKLNLEAQFAKFLNILKRICIKIPFAEALSRMPLYAKFLREIFSKKKAI 787
K +NPL K NL +++ IC +PF++ + +PL++ +KK+
Sbjct: 44 KHINPLINFARKKQNLSLNSLMAMSVASSICTLLPFSKPHNSVPLFSS-----QTKKRCR 208
Query: 788 DHKETIALTRESSAIVKKQPQKLRDPGS 815
++ R+++++V K + L D S
Sbjct: 209 IIISSVTEDRQAASVVSKDKKVLEDSSS 292
>TC79201 weakly similar to GP|10800070|dbj|BAB16490. contains ESTs
C61063(AU063369) C61063(AU093310)~unknown protein,
partial (18%)
Length = 951
Score = 30.4 bits (67), Expect = 3.3
Identities = 34/138 (24%), Positives = 56/138 (39%)
Frame = +3
Query: 708 SVKGEIVKLLGEKNAIKTPLDKNKTLNPLRLTKLNLEAQFAKFLNILKRICIKIPFAEAL 767
S KGE+VK N K D NKTL L+ + + + L ++ +P + +
Sbjct: 189 SEKGELVKSPPIINGEKKICDANKTLVALKNHREAERKRRNRINGHLAKLRALVPSSPKM 368
Query: 768 SRMPLYAKFLREIFSKKKAIDHKETIALTRESSAIVKKQPQKLRDPGSFAIPCVIGKETV 827
+ L A+ +R++ KK D VK +P + + GSF I +
Sbjct: 369 DKATLLAEVIRQVKHLKKNADEASKGYSIPTDDDEVKVEPYE--NGGSFLYKASISCDYR 542
Query: 828 DKALCHLGASVSLLPLSL 845
+ L L ++ L L L
Sbjct: 543 PELLSDLRQTLDKLQLQL 596
>TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (71%)
Length = 974
Score = 30.4 bits (67), Expect = 3.3
Identities = 13/28 (46%), Positives = 14/28 (49%)
Frame = +2
Query: 536 PSPASPTCEVCGITGHTGVDCQLGSAAN 563
P P S C CGI GH DC+ G N
Sbjct: 338 PPPGSGRCFNCGIDGHWARDCKAGDWKN 421
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 30.0 bits (66), Expect = 4.3
Identities = 21/81 (25%), Positives = 34/81 (41%), Gaps = 13/81 (16%)
Frame = +2
Query: 330 PTEDPNLHISSFLRLSGTI------KENQEAVRLHLFPFSLRDRASAWFHSLE------- 376
P + NL + FL TI KE E ++ + L+ A W+ +L+
Sbjct: 629 PDFEGNLQLDDFLDWLQTIERVFEYKEVPEEQKVKIVAAKLKKHALIWWENLKRRRKREG 808
Query: 377 VGSITSWDDMRRAFLARFSPP 397
I +WD MR+ ++ PP
Sbjct: 809 KSKIKTWDKMRQKLTRKYLPP 871
>TC91149 similar to GP|15081723|gb|AAK82516.1 T10F18_70/T10F18_70
{Arabidopsis thaliana}, partial (14%)
Length = 895
Score = 29.3 bits (64), Expect = 7.4
Identities = 20/53 (37%), Positives = 28/53 (52%)
Frame = +1
Query: 672 QTPGVFPGQTETNPKAHVNAISLGGNKLEETITKAKSVKGEIVKLLGEKNAIK 724
+ PG+ E NPK + AIS+ T TK++S KGEI K + N+ K
Sbjct: 274 ENPGISIDIVELNPKPNSKAISISFPDF-VTNTKSESGKGEIFKGKSKLNSKK 429
>BQ137213
Length = 1006
Score = 28.9 bits (63), Expect = 9.7
Identities = 19/53 (35%), Positives = 25/53 (46%), Gaps = 11/53 (20%)
Frame = +2
Query: 259 ILFYFILVLFPF*VSLLQIALYYISG-----------TNIGWIESWSKRKGLG 300
I FY I++ F VSLL Y +SG T+ W E+W R+G G
Sbjct: 50 IFFYSIIICF---VSLLFTFAYSVSGCARRFSDRSPFTSYVWSETWVPRRGAG 199
>TC89585 similar to GP|21553818|gb|AAM62911.1 unknown {Arabidopsis
thaliana}, partial (80%)
Length = 1445
Score = 28.9 bits (63), Expect = 9.7
Identities = 34/119 (28%), Positives = 54/119 (44%), Gaps = 9/119 (7%)
Frame = +1
Query: 628 LKNQTGSLNDS-------LSKLNTKVDSIATHTKMLETQISQVAQQVAISSQTPGVFPGQ 680
LK+QT S N S ++ K + +TK + ++Q I + TP F +
Sbjct: 115 LKHQTLSFNFSSHPPSHFINLFKVKSNRPIKYTKQKLKLYASLSQSEQIET-TPTTFRIK 291
Query: 681 TETNPKAHVNAISLGGNK-LEETITKAKSVKG-EIVKLLGEKNAIKTPLDKNKTLNPLR 737
NPK VN + +G + + + K +G + + EK+ IK +DK TLN LR
Sbjct: 292 ---NPK-DVNVLVVGSTGYIGKFVVKELIQRGFNVTAIAREKSGIKGSIDKETTLNELR 456
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.332 0.142 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,112,178
Number of Sequences: 36976
Number of extensions: 373943
Number of successful extensions: 2252
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 1701
Number of HSP's successfully gapped in prelim test: 67
Number of HSP's that attempted gapping in prelim test: 509
Number of HSP's gapped (non-prelim): 1817
length of query: 942
length of database: 9,014,727
effective HSP length: 105
effective length of query: 837
effective length of database: 5,132,247
effective search space: 4295690739
effective search space used: 4295690739
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC144538.1


