

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144517.1 + phase: 0
(202 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
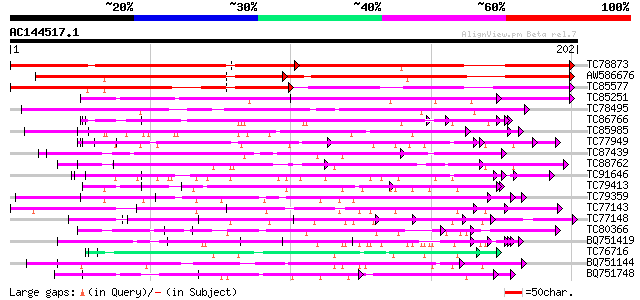
Score E
Sequences producing significant alignments: (bits) Value
TC78873 similar to GP|21740530|emb|CAD41509. OSJNBa0029H02.25 {O... 126 5e-30
AW586676 similar to GP|806720|gb|AA arabinogalactan-protein prec... 118 2e-27
TC85577 homologue to GP|6715512|gb|AAF26445.1| vacuolar H+-ATPas... 101 2e-22
TC85251 homologue to PIR|JQ2128|JQ2128 metallothionein - soybean... 89 1e-18
TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 ... 80 7e-16
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 78 2e-15
TC85985 similar to GP|5139695|dbj|BAA81686.1 expressed in cucumb... 77 3e-15
TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {A... 77 5e-15
TC87439 weakly similar to GP|5139695|dbj|BAA81686.1 expressed in... 76 8e-15
TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Ar... 75 2e-14
TC91646 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproli... 74 4e-14
TC79413 similar to GP|21439800|emb|CAD35173. unnamed protein pro... 72 1e-13
TC79359 weakly similar to PIR|S49915|S49915 extensin-like protei... 72 2e-13
TC77143 similar to GP|559557|gb|AAA51649.1|| arabinogalactan-pro... 72 2e-13
TC77148 ENOD20 71 3e-13
TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like... 69 1e-12
BQ751419 weakly similar to GP|21629340|gb L509.2 {Leishmania maj... 68 2e-12
TC76716 similar to PIR|T10863|T10863 extensin precursor - kidney... 67 4e-12
BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like ... 67 5e-12
BQ751748 similar to PIR|T47216|T47 probable V-ATPase 20K chain ... 67 6e-12
>TC78873 similar to GP|21740530|emb|CAD41509. OSJNBa0029H02.25 {Oryza
sativa}, partial (7%)
Length = 623
Score = 126 bits (317), Expect = 5e-30
Identities = 72/103 (69%), Positives = 80/103 (76%)
Frame = +3
Query: 1 MASYMVVLMLVASLLVSSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPF 60
MASY VVLMLVA+LLVSSTFA+S SSSP + SPA SP A SPA S P P+ N+PSPSP
Sbjct: 114 MASYKVVLMLVAALLVSSTFAQSPSSSP--SKSPAISPSAHSPAASPPAPVKNSPSPSPS 287
Query: 61 AVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIA 103
A+N P SPPPAS GSPA AAPAVTP SISTPP++APS A
Sbjct: 288 AINSPPSPPPASSGSPA---AAPAVTPSSISTPPAEAPSNGAA 407
Score = 120 bits (300), Expect = 5e-28
Identities = 67/124 (54%), Positives = 81/124 (65%), Gaps = 2/124 (1%)
Frame = +3
Query: 80 VAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPP- 138
+ A + + + PS +PS + A S SA+SP AS P PV +SPSPSPSAINSPPSPPP
Sbjct: 141 LVAALLVSSTFAQSPSSSPSKSPAISPSAHSPAASPPAPVKNSPSPSPSAINSPPSPPPA 320
Query: 139 -HASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMNGFNVAGYAAAVVFIA 197
SPAAAPA+TPSSISTPP +APS NGA +N F VAG AA V+F A
Sbjct: 321 SSGSPAAAPAVTPSSISTPPAEAPS--------------NGAALNRFTVAGSAAVVIFAA 458
Query: 198 ALIL 201
AL++
Sbjct: 459 ALMM 470
>AW586676 similar to GP|806720|gb|AA arabinogalactan-protein precursor
{Nicotiana alata}, partial (22%)
Length = 590
Score = 118 bits (295), Expect = 2e-27
Identities = 67/126 (53%), Positives = 86/126 (68%), Gaps = 2/126 (1%)
Frame = +2
Query: 78 SPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPP 137
SP ++P ++P++ +PAI+PS A+SP AS+P+PV ++PSPSPSAINSPPSPP
Sbjct: 89 SPSSSPTISPVA---------TPAISPS--ADSPAASAPIPVKNAPSPSPSAINSPPSPP 235
Query: 138 PHA--SPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMNGFNVAGYAAAVVF 195
P + SPA +PA+TPSSISTPP++ PS NGA +N F VAG AA VVF
Sbjct: 236 PASSDSPAVSPALTPSSISTPPSEGPSE-------------NGAALNRFTVAGSAAVVVF 376
Query: 196 IAALIL 201
AALIL
Sbjct: 377 AAALIL 394
Score = 107 bits (266), Expect = 4e-24
Identities = 59/92 (64%), Positives = 69/92 (74%), Gaps = 2/92 (2%)
Frame = +2
Query: 10 LVASLLVSSTFARSLSSSPVATP--SPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTS 67
L SLLVSSTFA+S SSSP +P +PA SP A SPA S+P+P+ NAPSPSP A+N P S
Sbjct: 50 LCFSLLVSSTFAQSPSSSPTISPVATPAISPSADSPAASAPIPVKNAPSPSPSAINSPPS 229
Query: 68 PPPASFGSPASPVAAPAVTPLSISTPPSQAPS 99
PPPAS SPA +PA+TP SISTPPS+ PS
Sbjct: 230 PPPASSDSPA---VSPALTPSSISTPPSEGPS 316
>TC85577 homologue to GP|6715512|gb|AAF26445.1| vacuolar H+-ATPase B subunit
{Nicotiana tabacum}, partial (14%)
Length = 841
Score = 101 bits (252), Expect = 2e-22
Identities = 66/108 (61%), Positives = 73/108 (67%), Gaps = 7/108 (6%)
Frame = -1
Query: 1 MASYMVVLMLVASLLVSSTFARSLSS----SPVATP---SPAGSPLAISPAVSSPVPLMN 53
MASY VVLMLVA+LLVSST A+S SS SPVATP S A SP A+SP S PVP+ N
Sbjct: 547 MASYTVVLMLVATLLVSSTIAQSPSSSPTISPVATPPKSSQAPSPSAVSPVASPPVPVKN 368
Query: 54 APSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPA 101
APSPS PPAS SPA +PAVTP SIST PS+APSP+
Sbjct: 367 APSPS----------PPASSDSPA---VSPAVTPSSISTTPSEAPSPS 263
Score = 91.7 bits (226), Expect = 2e-19
Identities = 56/124 (45%), Positives = 74/124 (59%)
Frame = -1
Query: 78 SPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPP 137
SP ++P ++P++ SQAPSP SA SPVAS PVPV ++PSPSP A +
Sbjct: 481 SPSSSPTISPVATPPKSSQAPSP------SAVSPVASPPVPVKNAPSPSPPASSD----- 335
Query: 138 PHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMNGFNVAGYAAAVVFIA 197
SPA +PA+TPSSIST P++APS + + A N F VAG AA +VF A
Sbjct: 334 ---SPAVSPAVTPSSISTTPSEAPS----------PSDNSAAAFNRFTVAGSAAVMVFAA 194
Query: 198 ALIL 201
AL++
Sbjct: 193 ALMM 182
>TC85251 homologue to PIR|JQ2128|JQ2128 metallothionein - soybean, partial
(93%)
Length = 1554
Score = 89.0 bits (219), Expect = 1e-18
Identities = 59/179 (32%), Positives = 78/179 (42%), Gaps = 3/179 (1%)
Frame = -2
Query: 26 SSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPA- 84
S+PV +P+P +P A SP V P P ++P +P + P PPP +P APA
Sbjct: 719 SAPVTSPAPKIAP-ASSPKVPPPQPPKSSPVSTP-TLPPPLPPPPKISPTPVQTPPAPAP 546
Query: 85 --VTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASP 142
TP+ P QAP+PA A S +P + PV + S P P H +P
Sbjct: 545 VKATPVPAPAPAKQAPTPAPATSPPIPAPTPAIEAPVPAPESSKPKRRRHRPKHRRHQAP 366
Query: 143 AAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMNGFNVAGYAAAVVFIAALIL 201
A AP + S PPT + T P APS NGA N F + F + L
Sbjct: 365 APAPTVIHKSPPAPPTDTTADSDTAPAPAPSFNLNGAPSNHFQGGNIWTTIGFAITVFL 189
Score = 41.6 bits (96), Expect = 2e-04
Identities = 27/87 (31%), Positives = 40/87 (45%), Gaps = 12/87 (13%)
Frame = -2
Query: 101 AIAPSTSANSPVASSPVPVTSSPSPSPSAINSPP---SPPPHASPAAAPAITP------- 150
A AP+T+ ++P +T + P+ + SPP PP +P +AP +P
Sbjct: 857 AQAPATAPTKLPPTTPTAITPVTTQPPTVVASPPITSQPPVTVAPKSAPVTSPAPKIAPA 678
Query: 151 SSISTPPTKAPSS--ISTPPTKAPSTP 175
SS PP + P S +STP P P
Sbjct: 677 SSPKVPPPQPPKSSPVSTPTLPPPLPP 597
>TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 - alfalfa,
partial (97%)
Length = 1049
Score = 79.7 bits (195), Expect = 7e-16
Identities = 61/191 (31%), Positives = 90/191 (46%), Gaps = 10/191 (5%)
Frame = +3
Query: 5 MVVLMLVASLLVSSTFARSLSSSPVATPSPAGSPLAISPAV--SSPVPLMNAPSPSPFAV 62
+VV+ L+ + S ++ S+SP ++P+P P P +SP P+ ++P P
Sbjct: 186 LVVIGLICVVFSSVGAQQAPSTSPNSSPAPPTPPANTPPTTPQASPPPVQSSPPP----- 350
Query: 63 NYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSS 122
+SPPP +SP A + P S+PP SP P T PV S+P P S
Sbjct: 351 -VQSSPPPLQ----SSPPPAQSTPPPVQSSPPPVQQSPPPTPLTP--PPVQSTP-PPASP 506
Query: 123 PSPSPSAINSPPSPPPHASPAAA---PAITPSSISTPPTKAPSSI-----STPPTKAPST 174
P SP + PP+ PP A+P A PA+TP+ +S+PP P+ S P AP
Sbjct: 507 PPASPPPFSPPPATPPPATPPPATPPPALTPTPLSSPPATTPAPAPAKLKSKAPALAPVL 686
Query: 175 PTNGATMNGFN 185
+ A G +
Sbjct: 687 SPSDAPAPGLS 719
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 78.2 bits (191), Expect = 2e-15
Identities = 52/152 (34%), Positives = 64/152 (41%), Gaps = 4/152 (2%)
Frame = +2
Query: 28 PVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTP 87
P SP P P P P+ + P P P+ Y +SPPP SP P +P
Sbjct: 11 PYCVRSPPPPPPNSPPPPPPPAPVFSPPPPVPY---YYSSPPPPPAHSPPPPPXSP---- 169
Query: 88 LSISTPPSQAPSPAIAP--STSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAA 145
PP +P+P + P S PV S P PV S P PSP PP PPP
Sbjct: 170 ----PPPPHSPTPPVYPYLSPPPPPPVHSPPPPVYSPPPPSPPPCVEPPPPPPPPCVEPP 337
Query: 146 PAITPSSISTP--PTKAPSSISTPPTKAPSTP 175
P +P+ TP P +PS +P PS P
Sbjct: 338 PPSSPAPHQTPYHPPPSPSPPPSPVYAYPSPP 433
Score = 70.5 bits (171), Expect = 4e-13
Identities = 53/157 (33%), Positives = 71/157 (44%), Gaps = 4/157 (2%)
Frame = +2
Query: 26 SSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPV-AAPA 84
SSP P PA SP P SP P ++P+P + Y + PPP SP PV + P
Sbjct: 116 SSP--PPPPAHSP---PPPPXSPPPPPHSPTPPVYP--YLSPPPPPPVHSPPPPVYSPPP 274
Query: 85 VTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPP--HASP 142
+P PP P P + P ++ +P SPSP PS + + PSPPP + SP
Sbjct: 275 PSPPPCVEPPPPPPPPCVEPPPPSSPAPHQTPYHPPPSPSPPPSPVYAYPSPPPPVYTSP 454
Query: 143 AAAPAITPSSISTPPTKAPSSISTPPT-KAPSTPTNG 178
+P + +PP SS PP + P P G
Sbjct: 455 PPSPVY---AYPSPPPPVYSSPPPPPVYEGPIPPVFG 556
Score = 69.7 bits (169), Expect = 7e-13
Identities = 48/133 (36%), Positives = 58/133 (43%), Gaps = 2/133 (1%)
Frame = +2
Query: 27 SPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVA-APAV 85
SP P SP P S P P+ + P PSP P PPP P P + AP
Sbjct: 185 SPTPPVYPYLSPPPPPPVHSPPPPVYSPPPPSPPPCVEPPPPPPPPCVEPPPPSSPAPHQ 364
Query: 86 TPLSISTPPSQAPSPAIA-PSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAA 144
TP PS PSP A PS + P PV + PSP P +SPP PP + P
Sbjct: 365 TPYHPPPSPSPPPSPVYAYPSPPPPVYTSPPPSPVYAYPSPPPPVYSSPPPPPVYEGP-- 538
Query: 145 APAITPSSISTPP 157
P + S ++PP
Sbjct: 539 IPPVFGISYASPP 577
Score = 64.7 bits (156), Expect = 2e-11
Identities = 47/132 (35%), Positives = 54/132 (40%)
Frame = +2
Query: 48 PVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTS 107
P P SP P N P PPP PA + P P S+PP P PA +P
Sbjct: 2 PTPPYCVRSPPPPPPNSPPPPPP-----PAPVFSPPPPVPYYYSSPP---PPPAHSPP-- 151
Query: 108 ANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTP 167
P SP P SP+P SPP PPP SP P +P S PP P P
Sbjct: 152 ---PPPXSPPPPPHSPTPPVYPYLSPPPPPPVHSPPP-PVYSPPPPSPPPCVEPPPPPPP 319
Query: 168 PTKAPSTPTNGA 179
P P P++ A
Sbjct: 320 PCVEPPPPSSPA 355
Score = 56.6 bits (135), Expect = 6e-09
Identities = 43/124 (34%), Positives = 52/124 (41%), Gaps = 1/124 (0%)
Frame = +2
Query: 28 PVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAA-PAVT 86
PV +P P P + P P P + P PS A + PP S P SPV A P+
Sbjct: 254 PVYSPPPPSPPPCVEPPPPPPPPCVEPPPPSSPAPHQTPYHPPPSPSPPPSPVYAYPSPP 433
Query: 87 PLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAP 146
P ++PP PSP A S P PV SSP P P + P PP A+P
Sbjct: 434 PPVYTSPP---PSPVYA--------YPSPPPPVYSSPPPPP--VYEGPIPPVFGISYASP 574
Query: 147 AITP 150
P
Sbjct: 575 PPPP 586
>TC85985 similar to GP|5139695|dbj|BAA81686.1 expressed in cucumber
hypocotyls {Cucumis sativus}, partial (42%)
Length = 892
Score = 77.4 bits (189), Expect = 3e-15
Identities = 56/160 (35%), Positives = 76/160 (47%), Gaps = 5/160 (3%)
Frame = +3
Query: 29 VATPSPAGSPLAISPAVSSP--VPLMNAPSP-SPFAVNYPTSPPPASFGSPASPVAAPAV 85
V SP+ +P P V++P P+ P SP V P S PPAS SP + PAV
Sbjct: 111 VGGQSPSSAPTTSPPTVTTPSAAPVAAPTKPKSPAPVASPKSSPPAS--SPTAATVTPAV 284
Query: 86 TPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASP--A 143
+P + P A SPA + A P PV+S P+P P ++SPP+P P +SP A
Sbjct: 285 SP---AAPVPVAKSPAASSPVVAPVSTPPKPAPVSSPPAPVP--VSSPPTPVPVSSPPTA 449
Query: 144 AAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMNG 183
+ PA+TPS+ + S T K S P + G
Sbjct: 450 STPAVTPSAEVPAAAPSKSKKKTKKGKKHSAPAPSPALEG 569
Score = 73.6 bits (179), Expect = 5e-14
Identities = 55/155 (35%), Positives = 79/155 (50%), Gaps = 15/155 (9%)
Frame = +3
Query: 25 SSSPVATPS---PAGSPLA--ISPAVS--SPVPLMNAPSPS-PFAVNYPTSPPPASFGSP 76
S +PVA+P PA SP A ++PAVS +PVP+ +P+ S P T P PA SP
Sbjct: 207 SPAPVASPKSSPPASSPTAATVTPAVSPAAPVPVAKSPAASSPVVAPVSTPPKPAPVSSP 386
Query: 77 ASPV-AAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVT------SSPSPSPSA 129
+PV + TP+ +S+PP+ A +PA+ PS + S T S+P+PSP A
Sbjct: 387 PAPVPVSSPPTPVPVSSPPT-ASTPAVTPSAEVPAAAPSKSKKKTKKGKKHSAPAPSP-A 560
Query: 130 INSPPSPPPHASPAAAPAITPSSISTPPTKAPSSI 164
+ PP+PP A + A +P S +I
Sbjct: 561 LEGPPAPPVGAPGPSLDASSPGPASAADESGAETI 665
Score = 64.3 bits (155), Expect = 3e-11
Identities = 56/184 (30%), Positives = 84/184 (45%), Gaps = 10/184 (5%)
Frame = +3
Query: 6 VVLMLVASLLVSSTFARSLSSSPVATP----SPAGSPLAISPAVSSPVPLMNAPSPSPFA 61
V+ + + ++ + +S SS+P +P +P+ +P+A SP P+ + P SP
Sbjct: 72 VLSLALICIVFAGVGGQSPSSAPTTSPPTVTTPSAAPVAAPTKPKSPAPVAS-PKSSP-- 242
Query: 62 VNYPTSPPPASFGSPASPVAAP---AVTPLSIS---TPPSQAPSPAIAPSTSANSPVASS 115
P S P A+ +PA AAP A +P + S P S P PA S A PV+S
Sbjct: 243 ---PASSPTAATVTPAVSPAAPVPVAKSPAASSPVVAPVSTPPKPAPVSSPPAPVPVSSP 413
Query: 116 PVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTP 175
P PV P SP ++P P PAAAP+ + + K + +P + P P
Sbjct: 414 PTPV---PVSSPPTASTPAVTPSAEVPAAAPSKSKKK-TKKGKKHSAPAPSPALEGPPAP 581
Query: 176 TNGA 179
GA
Sbjct: 582 PVGA 593
>TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {Arabidopsis
thaliana}, partial (34%)
Length = 1297
Score = 77.0 bits (188), Expect = 5e-15
Identities = 60/183 (32%), Positives = 82/183 (44%), Gaps = 17/183 (9%)
Frame = +3
Query: 31 TPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVA-APAVTPLS 89
TP + SP A + SP P PSPS GSP SPVA PA +P+
Sbjct: 495 TPKSSPSPSAGGLSPPSPSPTTTTPSPS---------------GSPPSPVAIPPASSPVP 629
Query: 90 ISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSP---PPHASPAAAP 146
S P + +PSP ++ + A P+ASSP PV S+P P+ + S PSP PP SPA P
Sbjct: 630 TSGPTASSPSPVVS-TPPAGGPMASSPSPVVSTP-PAGGPMASSPSPAGGPPALSPAGGP 803
Query: 147 AITPSSISTPPTKAPSSISTPP-------------TKAPSTPTNGATMNGFNVAGYAAAV 193
+ ++ PP P +T P AP +P + +T N + A
Sbjct: 804 S---TAAGGPPAPGPGGAATSPGPGGAGASGPGGAAAAPGSPGSNSTAPSGNSGSFVAPS 974
Query: 194 VFI 196
F+
Sbjct: 975 NFL 983
Score = 67.8 bits (164), Expect = 3e-12
Identities = 51/156 (32%), Positives = 78/156 (49%), Gaps = 13/156 (8%)
Frame = +3
Query: 27 SPVATPSPAG---SPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAP 83
+P ++PSP+ SP + SP ++P P + PSP A+ +SP P S + +SP +P
Sbjct: 495 TPKSSPSPSAGGLSPPSPSPTTTTPSP--SGSPPSPVAIPPASSPVPTSGPTASSP--SP 662
Query: 84 AVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPS-----AINSPPSPPP 138
V+ P + +PSP ++ + A P+ASSP P P+ SP+ A PP+P P
Sbjct: 663 VVSTPPAGGPMASSPSPVVS-TPPAGGPMASSPSPAGGPPALSPAGGPSTAAGGPPAPGP 839
Query: 139 HAS-----PAAAPAITPSSISTPPTKAPSSISTPPT 169
+ P A A P + P +P S ST P+
Sbjct: 840 GGAATSPGPGGAGASGPGGAAAAP-GSPGSNSTAPS 944
Score = 63.9 bits (154), Expect = 4e-11
Identities = 52/156 (33%), Positives = 67/156 (42%), Gaps = 14/156 (8%)
Frame = +3
Query: 26 SSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAV 85
S TPSP+GSP SPV + A SP P + +SP P PA A +
Sbjct: 549 SPTTTTPSPSGSP-------PSPVAIPPASSPVPTSGPTASSPSPVVSTPPAGGPMASSP 707
Query: 86 TPLSISTPPSQAP--------------SPAIAPSTSANSPVASSPVPVTSSPSPSPSAIN 131
+P+ +STPP+ P SPA PST+A P A P +SP P + +
Sbjct: 708 SPV-VSTPPAGGPMASSPSPAGGPPALSPAGGPSTAAGGPPAPGPGGAATSPGPGGAGAS 884
Query: 132 SPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTP 167
P AAAP +P S ST P+ S P
Sbjct: 885 GP------GGAAAAPG-SPGSNSTAPSGNSGSFVAP 971
Score = 63.5 bits (153), Expect = 5e-11
Identities = 56/158 (35%), Positives = 71/158 (44%), Gaps = 8/158 (5%)
Frame = +3
Query: 39 LAISPAV--SSPVPL---MNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTP 93
+ ISP SSP P ++ PSPSP T+ P+ GSP SPVA P
Sbjct: 477 VVISPRTPKSSPSPSAGGLSPPSPSP------TTTTPSPSGSPPSPVAIP---------- 608
Query: 94 PSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAP-AITPSS 152
P+ +P P + P ASSP PV S+P P+ + S PSP PA P A +PS
Sbjct: 609 PASSPVPT-------SGPTASSPSPVVSTP-PAGGPMASSPSPVVSTPPAGGPMASSPSP 764
Query: 153 ISTPPTKAPSSISTPPTKAPSTPTNG--ATMNGFNVAG 188
PP +P+ + P P G AT G AG
Sbjct: 765 AGGPPALSPAGGPSTAAGGPPAPGPGGAATSPGPGGAG 878
Score = 47.0 bits (110), Expect = 5e-06
Identities = 29/91 (31%), Positives = 41/91 (44%)
Frame = +3
Query: 25 SSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPA 84
S SPV + PAG P+A SP+ + P ++ A P +P P + P A A
Sbjct: 702 SPSPVVSTPPAGGPMASSPSPAGGPPALSPAGGPSTAAGGPPAPGPGGAATSPGPGGAGA 881
Query: 85 VTPLSISTPPSQAPSPAIAPSTSANSPVASS 115
P + P S + APS ++ S VA S
Sbjct: 882 SGPGGAAAAPGSPGSNSTAPSGNSGSFVAPS 974
>TC87439 weakly similar to GP|5139695|dbj|BAA81686.1 expressed in cucumber
hypocotyls {Cucumis sativus}, partial (57%)
Length = 897
Score = 76.3 bits (186), Expect = 8e-15
Identities = 60/175 (34%), Positives = 85/175 (48%), Gaps = 11/175 (6%)
Frame = +3
Query: 14 LLVSSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASF 73
++V+ +S S++P +PS +P A +P VSSPV +P V P S PPA+
Sbjct: 78 IIVAGVGGQSPSAAPTTSPS---TPAATTP-VSSPVAAPPTTPTTPAPVASPKSSPPATS 245
Query: 74 GSPASPVAA-------PAVTPLSISTPPSQAPSPAIAPSTSA--NSPVASSPV--PVTSS 122
A+P A PAVTP +STPP AP P +P T A +SP A +PV P T+
Sbjct: 246 PKAAAPTATPPAASSPPAVTP--VSTPP-PAPVPVKSPPTPAPVSSPPAVTPVAAPTTTP 416
Query: 123 PSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTN 177
P+P+ + H +PA +PA+ PP AP + P+T N
Sbjct: 417 AVPAPAPSKGKKNKKKHGAPAPSPALL--GPPAPPAGAPGPSEDASSPGPATTAN 575
Score = 61.2 bits (147), Expect = 3e-10
Identities = 42/131 (32%), Positives = 57/131 (43%)
Frame = +3
Query: 11 VASLLVSSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPP 70
VAS S +++P ATP A SP A++P ++ P P+P V P +P P
Sbjct: 210 VASPKSSPPATSPKAAAPTATPPAASSPPAVTP--------VSTPPPAPVPVKSPPTPAP 365
Query: 71 ASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAI 130
S +PVAAP TP + PS+ A SP P P + +P PS
Sbjct: 366 VSSPPAVTPVAAPTTTPAVPAPAPSKGKKNKKKHGAPAPSPALLGP-PAPPAGAPGPSED 542
Query: 131 NSPPSPPPHAS 141
S P P A+
Sbjct: 543 ASSPGPATTAN 575
>TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Arabidopsis
thaliana}, partial (3%)
Length = 805
Score = 74.7 bits (182), Expect = 2e-14
Identities = 63/174 (36%), Positives = 89/174 (50%), Gaps = 12/174 (6%)
Frame = +3
Query: 38 PLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPA-SFGSPAS-----PVAAPAVTPLSIS 91
P + P + P N +PSP+ N PTSP + S SP S P + PA +P+S +
Sbjct: 3 PSSSLPPGVAQTPSGNGIAPSPYQFNNPTSPSSSPSASSPVSTTTSPPASTPASSPVSTT 182
Query: 92 TPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPS 151
T P PA ST A+SPV ++ P +P+ SP + NSP + P + PAAA TPS
Sbjct: 183 TSP-----PA---STPASSPVPTTTSPPAPTPASSPVSTNSPTASPAGSLPAAA---TPS 329
Query: 152 SISTPPTKAPSSISTPP-TKAPSTPTNG-----ATMNGFNVAGYAAAVVFIAAL 199
ST +PS STPP T++ + T G A+ + V Y+ ++ AAL
Sbjct: 330 PSST-TAGSPSPSSTPPGTRSSNETTPGRSIGVASASPSGVWAYSITILVGAAL 488
Score = 55.5 bits (132), Expect = 1e-08
Identities = 52/147 (35%), Positives = 71/147 (47%), Gaps = 2/147 (1%)
Frame = +3
Query: 25 SSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPA 84
+SSPV+T + SP A +PA SSPV T+ PPAS PA
Sbjct: 111 ASSPVSTTT---SPPASTPA-SSPVS--------------TTTSPPAS---------TPA 209
Query: 85 VTPLSISTPPSQAPSPAIAPSTSANSPVAS--SPVPVTSSPSPSPSAINSPPSPPPHASP 142
+P+ +T P AP+PA +P S NSP AS +P ++PSPS + SP SP
Sbjct: 210 SSPVPTTTSP-PAPTPASSP-VSTNSPTASPAGSLPAAATPSPSSTTAGSP-------SP 362
Query: 143 AAAPAITPSSISTPPTKAPSSISTPPT 169
++ P T SS T P ++ S P+
Sbjct: 363 SSTPPGTRSSNETTPGRSIGVASASPS 443
Score = 39.7 bits (91), Expect = 8e-04
Identities = 31/99 (31%), Positives = 41/99 (41%), Gaps = 2/99 (2%)
Frame = +3
Query: 18 STFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPA 77
ST +S+P ++P P S P P P+ SP + N PT+ P S + A
Sbjct: 174 STTTSPPASTPASSPVPT--------TTSPPAP---TPASSPVSTNSPTASPAGSLPAAA 320
Query: 78 --SPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVAS 114
SP + A +P STPP S P S AS
Sbjct: 321 TPSPSSTTAGSPSPSSTPPGTRSSNETTPGRSIGVASAS 437
>TC91646 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (15%)
Length = 984
Score = 73.9 bits (180), Expect = 4e-14
Identities = 59/167 (35%), Positives = 80/167 (47%), Gaps = 16/167 (9%)
Frame = +1
Query: 24 LSSSPVATPSPAGSPLAISPA-VSSPVPLMNAPS----PSPFAVNYPTSPP-PASFGSPA 77
L S + PSP SP +SP S +PL PS PSP A P SPP P P
Sbjct: 226 LPSPLMEAPSPRPSPSPLSPLRFPSRLPLTTPPSLTTSPSPTASPPPRSPPRPPPRSPPR 405
Query: 78 SPVAAPAVTPLSISTPP---------SQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPS 128
++ P P S S PP + P P ++PS S SP A++P PV S P P P+
Sbjct: 406 QLMSPPRPLPRSPSPPPRNPPELSRTTPTPRPPMSPSPSPTSPPAATP-PV-SRPPPRPA 579
Query: 129 AIN-SPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPST 174
PP P P P++ ++ ++ PP P++ S+PPT PS+
Sbjct: 580 LRRLLPPPPLPPTLPSSMASLFLAASVLPPASPPTT*SSPPTP*PSS 720
Score = 68.9 bits (167), Expect = 1e-12
Identities = 54/153 (35%), Positives = 67/153 (43%), Gaps = 19/153 (12%)
Frame = +1
Query: 48 PVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTS 107
P PLM APSP P SP SP SP+ P+ PL+ TPPS SP+ S
Sbjct: 229 PSPLMEAPSPRP-------SP------SPLSPLRFPSRLPLT--TPPSLTTSPSPTASPP 363
Query: 108 ANSP-----------VASSPVPVTSSPSPSP------SAINSPPSPP--PHASPAAAPAI 148
SP + S P P+ SPSP P S P PP P SP + PA
Sbjct: 364 PRSPPRPPPRSPPRQLMSPPRPLPRSPSPPPRNPPELSRTTPTPRPPMSPSPSPTSPPAA 543
Query: 149 TPSSISTPPTKAPSSISTPPTKAPSTPTNGATM 181
TP PP A + PP P+ P++ A++
Sbjct: 544 TPPVSRPPPRPALRRLLPPPPLPPTLPSSMASL 642
Score = 65.9 bits (159), Expect = 1e-11
Identities = 67/193 (34%), Positives = 90/193 (45%), Gaps = 21/193 (10%)
Frame = +1
Query: 23 SLSSSPVATPSPAGSPLAISPAVSSPVPLMNAP-----SPSPFAVNYP----TSPPPASF 73
SL++SP T SP P S P LM+ P SPSP N P T+P P
Sbjct: 325 SLTTSPSPTASPPPRSPPRPPPRSPPRQLMSPPRPLPRSPSPPPRNPPELSRTTPTPRPP 504
Query: 74 GSPA-SPVAAPAVTPLSISTPPSQA--------PSPAIAPSTSANSPVASSPVPVTSSPS 124
SP+ SP + PA TP PP A P P PS+ A+ +A+S +P P+
Sbjct: 505 MSPSPSPTSPPAATPPVSRPPPRPALRRLLPPPPLPPTLPSSMASLFLAASVLP----PA 672
Query: 125 PSPSAINSPPSPPPHASPAAAPAITPSSISTP---PTKAPSSISTPPTKAPSTPTNGATM 181
P+ +SPP+P P S AA P+S + P T +P + T T+ PS T+ AT+
Sbjct: 673 SPPTT*SSPPTP*P--SSAALLLCLPASATLPCSEQTASPVTSWTTSTRCPSARTS-ATL 843
Query: 182 NGFNVAGYAAAVV 194
+ AA VV
Sbjct: 844 SAPVTLPSAAVVV 882
Score = 60.1 bits (144), Expect = 6e-10
Identities = 56/164 (34%), Positives = 78/164 (47%), Gaps = 14/164 (8%)
Frame = +1
Query: 28 PVATPSPAG-SPLAISPAVSSPVPL-----MNAPSPSPFAVNYPTSPPPASFGSPASPVA 81
P +PSP+ SP A +P VS P P + P P P PT P + A+ V
Sbjct: 499 PPMSPSPSPTSPPAATPPVSRPPPRPALRRLLPPPPLP-----PTLPSSMASLFLAASVL 663
Query: 82 APAVTPLSISTPPSQAPSPAI------APSTSANSPVASSPVPVTSSPSPSPSAINSPPS 135
PA P + S+PP+ PS A A +T S +SPV ++ + PSA S
Sbjct: 664 PPASPPTT*SSPPTP*PSSAALLLCLPASATLPCSEQTASPVTSWTTSTRCPSARTSATL 843
Query: 136 PPPHASPAAAPAITP--SSISTPPTKAPSSISTPPTKAPSTPTN 177
P P+AA + P SS+S+ P + +S ST + APS PT+
Sbjct: 844 SAPVTLPSAAVVVLPLLSSVSSSP-RPSASPSTSASTAPSVPTS 972
Score = 28.5 bits (62), Expect = 1.8
Identities = 18/45 (40%), Positives = 25/45 (55%), Gaps = 1/45 (2%)
Frame = +1
Query: 26 SSPVATPSPAGSPLAISPAV-SSPVPLMNAPSPSPFAVNYPTSPP 69
S+PV PS A L + +V SSP P + + + A + PTSPP
Sbjct: 844 SAPVTLPSAAVVVLPLLSSVSSSPRPSASPSTSASTAPSVPTSPP 978
>TC79413 similar to GP|21439800|emb|CAD35173. unnamed protein product
{Glycine max}, partial (42%)
Length = 971
Score = 72.4 bits (176), Expect = 1e-13
Identities = 48/123 (39%), Positives = 61/123 (49%), Gaps = 9/123 (7%)
Frame = +2
Query: 62 VNY--PTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPV 119
VNY PT+PP + +P SPV AP PL S P +APSP + + SP P+
Sbjct: 122 VNYVNPTAPP-SPIVTPPSPVKAPPTPPLVKSPPIVKAPSPPLVKTPPYQSPPVKPTPPI 298
Query: 120 TSSPSPSPSAINSPPSPPPHASPAAAPAI---TPSSISTPPT----KAPSSISTPPTKAP 172
SP PSP + SPP P A +P + TP + +PP+ K P S P KAP
Sbjct: 299 VKSP-PSPPLVKSPPYQSPPIVKAPSPPLVKPTPPIVKSPPSPPLVKTPPYQSPPIVKAP 475
Query: 173 STP 175
TP
Sbjct: 476 PTP 484
Score = 70.1 bits (170), Expect = 6e-13
Identities = 53/153 (34%), Positives = 71/153 (45%), Gaps = 3/153 (1%)
Frame = +2
Query: 27 SPVATP-SPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAV 85
SP+ TP SP +P P V SP P++ APSP P P PP P P+
Sbjct: 152 SPIVTPPSPVKAP-PTPPLVKSP-PIVKAPSP-PLVKTPPYQSPPVK---PTPPIVKSPP 313
Query: 86 TPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPP--SPPPHASPA 143
+P + +PP Q+P APS P+ P+ SP PSP + +PP SPP +P
Sbjct: 314 SPPLVKSPPYQSPPIVKAPSP----PLVKPTPPIVKSP-PSPPLVKTPPYQSPPIVKAPP 478
Query: 144 AAPAITPSSISTPPTKAPSSISTPPTKAPSTPT 176
P I + TPP ++P + PP PT
Sbjct: 479 TPPPI----VKTPPYQSPPIVK-PPVAPSPPPT 562
Score = 64.7 bits (156), Expect = 2e-11
Identities = 50/147 (34%), Positives = 67/147 (45%), Gaps = 19/147 (12%)
Frame = +2
Query: 48 PVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQA----PSPAIA 103
P P++ PSP V P +PP SP A +P + TPP Q+ P+P I
Sbjct: 149 PSPIVTPPSP----VKAPPTPPLVK-----SPPIVKAPSPPLVKTPPYQSPPVKPTPPIV 301
Query: 104 PSTSANSPVASSPV---PVTSSPS-----PSPSAINSPPSP-----PPHASP--AAAPAI 148
S + V S P P+ +PS P+P + SPPSP PP+ SP AP
Sbjct: 302 KSPPSPPLVKSPPYQSPPIVKAPSPPLVKPTPPIVKSPPSPPLVKTPPYQSPPIVKAPPT 481
Query: 149 TPSSISTPPTKAPSSISTPPTKAPSTP 175
P + TPP ++P + P APS P
Sbjct: 482 PPPIVKTPPYQSPPIVK--PPVAPSPP 556
Score = 50.1 bits (118), Expect = 6e-07
Identities = 40/111 (36%), Positives = 47/111 (42%)
Frame = +2
Query: 27 SPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVT 86
+P SP PL SP SP P++ APSP P P SP SP
Sbjct: 287 TPPIVKSPPSPPLVKSPPYQSP-PIVKAPSPPL------VKPTPPIVKSPPSP------- 424
Query: 87 PLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPP 137
PL + TPP Q+P AP T P+ +P P S P P SPP P
Sbjct: 425 PL-VKTPPYQSPPIVKAPPTP--PPIVKTP-PYQSPPIVKPPVAPSPPPTP 565
>TC79359 weakly similar to PIR|S49915|S49915 extensin-like protein - maize,
partial (3%)
Length = 1170
Score = 71.6 bits (174), Expect = 2e-13
Identities = 70/195 (35%), Positives = 91/195 (45%), Gaps = 13/195 (6%)
Frame = +3
Query: 3 SYMVVLMLVASLLVSSTFARSLSSSPVATP------SPAGSPLAISPAVS---SPVPLMN 53
S V+ ML L + A S SS A P SP SP SPA S SP
Sbjct: 177 SLFVLSMLFTVLFAQNAPASSPKSSVTAKPPSSVSVSPTNSPA--SPAKSPTLSPPSQTP 350
Query: 54 APSPSPFAVNYP--TSPPPASFGSPASPVAAPAVTPLS--ISTPPSQAPSPAIAPSTSAN 109
A SPS A P TSPP S +PA++P + STPP SPA +P T+A
Sbjct: 351 AVSPSGSASTPPPATSPPAKSPAVQPPSSVSPAISPSNNVSSTPP--VSSPASSPPTAAV 524
Query: 110 SPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPT 169
SPV+S PV + SP +S P A+PA APA T S +P T +P+S S +
Sbjct: 525 SPVSS---PVEAPSVSSPPEASSAGIPSSSATPADAPAATLPSSKSPGT-SPASSSPETS 692
Query: 170 KAPSTPTNGATMNGF 184
+ P+ + + + F
Sbjct: 693 QGPAAADDSGSRSSF 737
Score = 71.6 bits (174), Expect = 2e-13
Identities = 54/136 (39%), Positives = 76/136 (55%), Gaps = 8/136 (5%)
Frame = +3
Query: 53 NAPSPSPFAVNYPTSPPPASFG-SPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSAN-- 109
NAP+ SP + T+ PP+S SP + A+PA +P +PPSQ +PA++PS SA+
Sbjct: 222 NAPASSP--KSSVTAKPPSSVSVSPTNSPASPAKSPTL--SPPSQ--TPAVSPSGSASTP 383
Query: 110 ----SPVASSP-VPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSI 164
SP A SP V SS SP+ S N+ S PP +SPA++P S + P +APS
Sbjct: 384 PPATSPPAKSPAVQPPSSVSPAISPSNNVSSTPPVSSPASSPPTAAVSPVSSPVEAPSVS 563
Query: 165 STPPTKAPSTPTNGAT 180
S P + P++ AT
Sbjct: 564 SPPEASSAGIPSSSAT 611
Score = 63.5 bits (153), Expect = 5e-11
Identities = 54/148 (36%), Positives = 71/148 (47%), Gaps = 1/148 (0%)
Frame = +3
Query: 26 SSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAV 85
S +TP PA SP A SPAV P + A SPS N +S PP S SPAS AV
Sbjct: 363 SGSASTPPPATSPPAKSPAVQPPSSVSPAISPS----NNVSSTPPVS--SPASSPPTAAV 524
Query: 86 TPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTS-SPSPSPSAINSPPSPPPHASPAA 144
+P+S SP APS S+ +S+ +P +S +P+ +P+A + P SP
Sbjct: 525 SPVS---------SPVEAPSVSSPPEASSAGIPSSSATPADAPAA-----TLPSSKSPGT 662
Query: 145 APAITPSSISTPPTKAPSSISTPPTKAP 172
+PA + S P A S S AP
Sbjct: 663 SPASSSPETSQGPAAADDSGSRSSFGAP 746
>TC77143 similar to GP|559557|gb|AAA51649.1|| arabinogalactan-protein {Pyrus
communis}, partial (30%)
Length = 956
Score = 71.6 bits (174), Expect = 2e-13
Identities = 54/170 (31%), Positives = 77/170 (44%), Gaps = 3/170 (1%)
Frame = +3
Query: 1 MASYMVV-LMLVASLLVSSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSP 59
MASY V + L+ LL +S FA++ ++P PS +P P+P+P
Sbjct: 108 MASYAAVQVFLILGLLATSCFAQAPGAAPTQPPSATPTP----------------PTPAP 239
Query: 60 FAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPV 119
A PT+ PP + TPP+ P PA AP+ + +P S P P
Sbjct: 240 VATP-PTATPPTA-------------------TPPTATPPPAAAPTPATPAPATSPPAPT 359
Query: 120 TSSPSPSP-SAINSPPSPPPHASPAAAPAITPSSISTPPT-KAPSSISTP 167
+S +P+P S +SPP+P P PA P S PP+ A SI+ P
Sbjct: 360 PTSDAPTPDSTSSSPPAPGP-----GGPAPGPGSTDAPPSPSAAFSINKP 494
Score = 53.1 bits (126), Expect = 7e-08
Identities = 37/125 (29%), Positives = 60/125 (47%), Gaps = 5/125 (4%)
Frame = +3
Query: 78 SPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSP- 136
+P AAP P + TPP+ AP +A +A P A+ P T++P P+ + + P+P
Sbjct: 177 APGAAPTQPPSATPTPPTPAP---VATPPTATPPTATPP---TATPPPAAAPTPATPAPA 338
Query: 137 --PPHASPAAAPAITPSSISTPPTKAPSSISTPP--TKAPSTPTNGATMNGFNVAGYAAA 192
PP +P + S+ S+PP P + P T AP +P+ ++N +A A +
Sbjct: 339 TSPPAPTPTSDAPTPDSTSSSPPAPGPGGPAPGPGSTDAPPSPSAAFSINKPIMAATALS 518
Query: 193 VVFIA 197
A
Sbjct: 519 AAIFA 533
Score = 51.2 bits (121), Expect = 3e-07
Identities = 46/149 (30%), Positives = 67/149 (44%), Gaps = 16/149 (10%)
Frame = +3
Query: 46 SSPVPLMNAPSPSPFAVNYP-TSPPPASFGSPASPVAAPAVTPLSI----------STPP 94
S+ + + + SP+ + +P +P P S AS A L + P
Sbjct: 15 STTLSISHLFSPTKYKHLFPFQTPKPLSITKMASYAAVQVFLILGLLATSCFAQAPGAAP 194
Query: 95 SQAPSPAIAPSTSANSPVASSP--VPVTSSP---SPSPSAINSPPSPPPHASPAAAPAIT 149
+Q PS P T A PVA+ P P T++P +P P+A +P +P P SP PA T
Sbjct: 195 TQPPSATPTPPTPA--PVATPPTATPPTATPPTATPPPAAAPTPATPAPATSP---PAPT 359
Query: 150 PSSISTPPTKAPSSISTPPTKAPSTPTNG 178
P+S + P S+ S+PP P P G
Sbjct: 360 PTSDAPTP---DSTSSSPPAPGPGGPAPG 437
>TC77148 ENOD20
Length = 1108
Score = 70.9 bits (172), Expect = 3e-13
Identities = 51/140 (36%), Positives = 74/140 (52%), Gaps = 5/140 (3%)
Frame = +3
Query: 68 PPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVP-----VTSS 122
PPP S +P S P S+ +PPS +PSP+ +PS S + S+P+P +S
Sbjct: 429 PPPPSPPTPRSSTPIPHPPRRSLPSPPSPSPSPSPSPSPSPSP--RSTPIPHPRKRSPAS 602
Query: 123 PSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMN 182
PSPSPS SP P SP+ AP+ + S S P+ +PS S P+ APS ++G+
Sbjct: 603 PSPSPSL---SKSPSPSESPSLAPSPSDSVASLAPSSSPSDES--PSPAPSPSSSGSKGG 767
Query: 183 GFNVAGYAAAVVFIAALILM 202
G AG+ V IA ++ +
Sbjct: 768 G---AGHGFLEVSIAMMMFL 818
Score = 58.2 bits (139), Expect = 2e-09
Identities = 36/107 (33%), Positives = 56/107 (51%), Gaps = 4/107 (3%)
Frame = +3
Query: 43 PAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPA-SPVAAPAVTPLSIS---TPPSQAP 98
P S P P + P P P + P+ P P+ SP+ SP +P TP+ +P S +P
Sbjct: 432 PPPSPPTPRSSTPIPHPPRRSLPSPPSPSPSPSPSPSPSPSPRSTPIPHPRKRSPASPSP 611
Query: 99 SPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAA 145
SP+++ S S + + +P P S S +PS+ S SP P SP+++
Sbjct: 612 SPSLSKSPSPSESPSLAPSPSDSVASLAPSSSPSDESPSPAPSPSSS 752
Score = 58.2 bits (139), Expect = 2e-09
Identities = 44/126 (34%), Positives = 60/126 (46%), Gaps = 3/126 (2%)
Frame = +3
Query: 41 ISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSP 100
++P +SSP P + P+P + P PP S SP SP +P+ +P PS +P
Sbjct: 405 VAPVLSSPPPPPSPPTPRS-STPIP-HPPRRSLPSPPSPSPSPSPSP-----SPSPSPRS 563
Query: 101 AIAPSTSANSPVASSPVPVTS---SPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPP 157
P SP + SP P S SPS SPS SP + + AP+ +PS S P
Sbjct: 564 TPIPHPRKRSPASPSPSPSLSKSPSPSESPSLAPSPSD----SVASLAPSSSPSDESPSP 731
Query: 158 TKAPSS 163
+PSS
Sbjct: 732 APSPSS 749
Score = 53.5 bits (127), Expect = 5e-08
Identities = 43/114 (37%), Positives = 58/114 (50%), Gaps = 3/114 (2%)
Frame = +3
Query: 22 RSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVA 81
RSL S P +PSP+ SP SP +PSP + +P SPASP
Sbjct: 489 RSLPSPPSPSPSPSPSP--------SP-----SPSPRSTPIPHPRK------RSPASPSP 611
Query: 82 APAVTPLSISTPPSQAPSPAIAPSTS---ANSPVASSPVPVTSSPSPSPSAINS 132
+P S+S PS + SP++APS S A+ +SSP + SP+PSPS+ S
Sbjct: 612 SP-----SLSKSPSPSESPSLAPSPSDSVASLAPSSSPSDESPSPAPSPSSSGS 758
Score = 50.4 bits (119), Expect = 5e-07
Identities = 30/84 (35%), Positives = 46/84 (54%), Gaps = 12/84 (14%)
Frame = +3
Query: 102 IAPSTSANSPVASSPVPVTSSPSPSPS--AINSPPSPPPHASPAAAPAITPSSISTP--- 156
+AP S+ P S P P +S+P P P ++ SPPSP P SP+ +P+ +P S P
Sbjct: 405 VAPVLSSPPPPPSPPTPRSSTPIPHPPRRSLPSPPSPSPSPSPSPSPSPSPRSTPIPHPR 584
Query: 157 ------PTKAPSSISTP-PTKAPS 173
P+ +PS +P P+++PS
Sbjct: 585 KRSPASPSPSPSLSKSPSPSESPS 656
>TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like protein
{Arabidopsis thaliana}, partial (16%)
Length = 814
Score = 68.9 bits (167), Expect = 1e-12
Identities = 45/116 (38%), Positives = 60/116 (50%), Gaps = 5/116 (4%)
Frame = +3
Query: 56 SPSPFAVNYPTSPPPASFGSP--ASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVA 113
SP P P+SPPP++ +P +P A P TP + S+PP A PST +NSP
Sbjct: 6 SPPPAT---PSSPPPSTPANPPPVTPSAPPPSTPATPSSPPPSTTPSAPPPSTPSNSP-- 170
Query: 114 SSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAP---SSIST 166
P P T+ PS + PSPP P++ + +P S TPP +P SSIST
Sbjct: 171 --PSPPTTPAISPPSGGGTTPSPPSRTPPSSDDSPSPPSSKTPPPPSPPSSSSIST 332
Score = 67.0 bits (162), Expect = 5e-12
Identities = 55/141 (39%), Positives = 71/141 (50%), Gaps = 10/141 (7%)
Frame = +3
Query: 66 TSPPPASFGSP--ASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSP 123
+SPPPA+ SP ++P P VTP S PP PST A SSP P T+
Sbjct: 3 SSPPPATPSSPPPSTPANPPPVTP---SAPP---------PSTPATP---SSPPPSTTPS 137
Query: 124 SPSPSA-INSPPSPP--PHASPAAAPAITPSSIS-TPPTK----APSSISTPPTKAPSTP 175
+P PS NSPPSPP P SP + TPS S TPP+ +P S TPP P +P
Sbjct: 138 APPPSTPSNSPPSPPTTPAISPPSGGGTTPSPPSRTPPSSDDSPSPPSSKTPP---PPSP 308
Query: 176 TNGATMNGFNVAGYAAAVVFI 196
+ ++++ V G A V +
Sbjct: 309 PSSSSISTGTVIGIAVGAVVV 371
Score = 66.2 bits (160), Expect = 8e-12
Identities = 54/132 (40%), Positives = 69/132 (51%), Gaps = 1/132 (0%)
Frame = +3
Query: 25 SSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPA 84
SS P ATPS SP +PA PV +AP PS A P+SPPP++ +P A P
Sbjct: 3 SSPPPATPS---SPPPSTPANPPPVT-PSAPPPSTPAT--PSSPPPST-----TPSAPPP 149
Query: 85 VTPLSISTPPSQAPSPAIAP-STSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPA 143
TP + +PPS +PAI+P S +P S P +S SPSP + +PP P P +S
Sbjct: 150 STPSN--SPPSPPTTPAISPPSGGGTTPSPPSRTPPSSDDSPSPPSSKTPPPPSPPSS-- 317
Query: 144 AAPAITPSSIST 155
SSIST
Sbjct: 318 -------SSIST 332
Score = 38.9 bits (89), Expect = 0.001
Identities = 43/170 (25%), Positives = 61/170 (35%), Gaps = 19/170 (11%)
Frame = +3
Query: 25 SSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPA 84
S+ P +TPS + +PA+S P PSP P+ PP+S SP+ P
Sbjct: 135 SAPPPSTPSNSPPSPPTTPAISPPSGGGTTPSP-------PSRTPPSSDDSPSPP----- 278
Query: 85 VTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPS-------------------P 125
S TPP +P + + ST +A V V S
Sbjct: 279 ----SSKTPPPPSPPSSSSISTGTVIGIAVGAVVVLVFFSICCICFRKKKRRRRDEEYYG 446
Query: 126 SPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTP 175
+ PP+ P P P + + PP + +S PP AP P
Sbjct: 447 QQNYQQPPPAQRPKVEPYGGPPQQWQNNAHPP--SDHVVSKPPPPAPIPP 590
Score = 35.8 bits (81), Expect = 0.012
Identities = 44/176 (25%), Positives = 59/176 (33%), Gaps = 41/176 (23%)
Frame = +3
Query: 27 SPVATPS---PAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPV--A 81
SP TP+ P+G SP +P ++PSP P SPP +S S + + A
Sbjct: 174 SPPTTPAISPPSGGGTTPSPPSRTPPSSDDSPSPPSSKTPPPPSPPSSSSISTGTVIGIA 353
Query: 82 APAVTPLSIST---------------------------PPSQAPS------PAIAPSTSA 108
AV L + PP+Q P P +A
Sbjct: 354 VGAVVVLVFFSICCICFRKKKRRRRDEEYYGQQNYQQPPPAQRPKVEPYGGPPQQWQNNA 533
Query: 109 NSP---VASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAP 161
+ P V S P P P PS + PP PP S + S P +P
Sbjct: 534 HPPSDHVVSKPPPPAPIPPRPPSHVAPPPPPPAFISSSGGSGSNYSGGELLPPPSP 701
>BQ751419 weakly similar to GP|21629340|gb L509.2 {Leishmania major}, partial
(1%)
Length = 766
Score = 68.2 bits (165), Expect = 2e-12
Identities = 64/191 (33%), Positives = 91/191 (47%), Gaps = 29/191 (15%)
Frame = +1
Query: 18 STFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPA 77
+T+ SSSP A PSP + + +P +P P +P+P+ A +P PAS + A
Sbjct: 34 TTWQPPASSSPSARPSPPPTTASSTPP-GAPTP---SPTPTRRATLNGNTPAPASPTAQA 201
Query: 78 SPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSP-------SPSPSAI 130
+P P+ S ST + P+ + STSA+SP A+SP + SP +P+PSA
Sbjct: 202 TPPTGPSTAAPSPSTCTTTGPTSS---STSASSPAATSPRTTSPSPHSSGTPPAPAPSAS 372
Query: 131 NSPPSPPPHA---------SPAAAPAI------TPSSISTPP------TKAPSSISTPP- 168
S S PP A SP A PA T +S TPP K +S TP
Sbjct: 373 RSSTSRPPRARARRAASRSSPWATPARACTTAPTSASRQTPPPSPRTAAKPKASRCTPSR 552
Query: 169 TKAPSTPTNGA 179
++AP+ P+ A
Sbjct: 553 SRAPTEPSKMA 585
Score = 58.9 bits (141), Expect = 1e-09
Identities = 47/134 (35%), Positives = 62/134 (46%), Gaps = 18/134 (13%)
Frame = +1
Query: 57 PSPFAVNYPTS-PPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSA----NSP 111
P P A N T+ PPAS P A P+ P + S+ P AP+P+ P+ A N+P
Sbjct: 7 PQPAASNLHTTWQPPASSS----PSARPSPPPTTASSTPPGAPTPSPTPTRRATLNGNTP 174
Query: 112 VASSPV--------PVTSSPSPSPSAINSPPSPPPHASPAAA--PAIT---PSSISTPPT 158
+SP P T++PSPS P S AS AA P T P S TPP
Sbjct: 175 APASPTAQATPPTGPSTAAPSPSTCTTTGPTSSSTSASSPAATSPRTTSPSPHSSGTPPA 354
Query: 159 KAPSSISTPPTKAP 172
APS+ + ++ P
Sbjct: 355 PAPSASRSSTSRPP 396
Score = 46.6 bits (109), Expect = 7e-06
Identities = 51/159 (32%), Positives = 72/159 (45%), Gaps = 10/159 (6%)
Frame = +1
Query: 18 STFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSP-PPASFGSP 76
ST A S S+ P+ + S A SPA +SP SPSP + P +P P AS S
Sbjct: 220 STAAPSPSTCTTTGPT-SSSTSASSPAATSP----RTTSPSPHSSGTPPAPAPSASRSST 384
Query: 77 ASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVP---VTSSPSPS---PSAI 130
+ P A A S S+P + +PA A +T+ S +P P + P S PS
Sbjct: 385 SRPPRARARRAASRSSPWA---TPARACTTAPTSASRQTPPPSPRTAAKPKASRCTPSRS 555
Query: 131 NSPPSPPPHASPAAAP---AITPSSISTPPTKAPSSIST 166
+P P A+ AAA A P + + P +A +S+
Sbjct: 556 RAPTEPSKMAAAAAAAGTRATRPRATAAAPHRAALRLSS 672
Score = 45.8 bits (107), Expect = 1e-05
Identities = 36/112 (32%), Positives = 52/112 (46%), Gaps = 16/112 (14%)
Frame = +1
Query: 83 PAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSP---SAIN----SPPS 135
PA + L + P + SP+ PS + ASS P +PSP+P + +N +P S
Sbjct: 13 PAASNLHTTWQPPASSSPSARPSPPPTT--ASSTPPGAPTPSPTPTRRATLNGNTPAPAS 186
Query: 136 PPPHASPAAAPAITPSSISTPPTKAPSSIST---------PPTKAPSTPTNG 178
P A+P P+ S ST T P+S ST P T +PS ++G
Sbjct: 187 PTAQATPPTGPSTAAPSPSTCTTTGPTSSSTSASSPAATSPRTTSPSPHSSG 342
Score = 42.7 bits (99), Expect = 9e-05
Identities = 25/61 (40%), Positives = 34/61 (54%)
Frame = +1
Query: 123 PSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMN 182
P P+ S +++ PP +SP+A P S PPT A S+ PT +P TPT AT+N
Sbjct: 7 PQPAASNLHTTWQPPASSSPSARP-------SPPPTTASSTPPGAPTPSP-TPTRRATLN 162
Query: 183 G 183
G
Sbjct: 163 G 165
Score = 42.0 bits (97), Expect = 2e-04
Identities = 27/92 (29%), Positives = 44/92 (47%), Gaps = 9/92 (9%)
Frame = +1
Query: 98 PSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASP---------AAAPAI 148
P PA + + P ASS SP P+ ++ P +P P +P APA
Sbjct: 7 PQPAASNLHTTWQPPASSSPSARPSPPPTTASSTPPGAPTPSPTPTRRATLNGNTPAPA- 183
Query: 149 TPSSISTPPTKAPSSISTPPTKAPSTPTNGAT 180
+P++ +TPPT ++ +P T + PT+ +T
Sbjct: 184 SPTAQATPPTGPSTAAPSPSTCTTTGPTSSST 279
>TC76716 similar to PIR|T10863|T10863 extensin precursor - kidney bean,
partial (48%)
Length = 1275
Score = 67.4 bits (163), Expect = 4e-12
Identities = 53/162 (32%), Positives = 64/162 (38%), Gaps = 14/162 (8%)
Frame = +3
Query: 28 PVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPA-SFGSPASPV---AAP 83
PV +P P + P V SP P SP P YP+ PPP + SP PV +P
Sbjct: 219 PVYSPPPVYKYKSPPPPVYSPPPPYKYKSPPPPPYKYPSPPPPVYKYKSPPPPVYKYKSP 398
Query: 84 AVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPS-------P 136
P PS P P + S PV S P PV SP P + PP P
Sbjct: 399 PPPPKKPYKYPS--PPPPVYKYKSPPPPVYSPPPPVYKYKSPPPPVHSPPPPYKYKSPPP 572
Query: 137 PPHASPAAAPAITPSSISTPPT---KAPSSISTPPTKAPSTP 175
PP+ P+ P + PP K+P P K PS P
Sbjct: 573 PPYKYPSPPPPVYKYKSPPPPVYKYKSPPPPPKKPYKYPSPP 698
Score = 58.5 bits (140), Expect = 2e-09
Identities = 50/153 (32%), Positives = 61/153 (39%), Gaps = 6/153 (3%)
Frame = +3
Query: 29 VATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPA-SFGSPASPVAAPAVTP 87
+ TPSP P + S P P+ SP P Y + PPP + SP P P P
Sbjct: 6 INTPSP---PPPVYKYKSPPPPVYKYKSPPPPVYKYKSPPPPVYKYKSPPPPPKKPYKYP 176
Query: 88 LSISTPPSQAPSPAIAPSTSANSPVAS--SPVPVTSSPSPSPSAINSPPSPPPHASPAAA 145
S PP + P + PV SP P SP P P SPP PPP+ P+
Sbjct: 177 ---SPPPPVYKYKSPPPPVYSPPPVYKYKSPPPPVYSPPP-PYKYKSPP-PPPYKYPSPP 341
Query: 146 PAITPSSISTPPT---KAPSSISTPPTKAPSTP 175
P + PP K+P P K PS P
Sbjct: 342 PPVYKYKSPPPPVYKYKSPPPPPKKPYKYPSPP 440
Score = 50.1 bits (118), Expect = 6e-07
Identities = 45/138 (32%), Positives = 54/138 (38%), Gaps = 1/138 (0%)
Frame = +3
Query: 32 PSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPA-SFGSPASPVAAPAVTPLSI 90
P P + P V SP P SP P YP+ PPP + SP PV P
Sbjct: 489 PPPVYKYKSPPPPVHSPPPPYKYKSPPPPPYKYPSPPPPVYKYKSPPPPVYKYKSPP--- 659
Query: 91 STPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITP 150
PP + P +P S P PV SP P ++SP PPPH ++ P P
Sbjct: 660 --PPPKKPYKYPSPPPPVYK-YKSPPPPVYKYKSPPPP-VHSP--PPPHYVYSSPP---P 812
Query: 151 SSISTPPTKAPSSISTPP 168
S PP S PP
Sbjct: 813 PVYSPPPPHYIYSSPPPP 866
Score = 45.8 bits (107), Expect = 1e-05
Identities = 47/156 (30%), Positives = 56/156 (35%), Gaps = 8/156 (5%)
Frame = +3
Query: 28 PVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTP 87
P PSP P + S P P+ + P P Y + PPP SP P + P
Sbjct: 417 PYKYPSP---PPPVYKYKSPPPPVYSPPPP---VYKYKSPPPPVH--SPPPPYKYKSPPP 572
Query: 88 LSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAP- 146
PP + PSP P SP PV SP P P PSPPP +P
Sbjct: 573 -----PPYKYPSPP-PPVYKYKSP--PPPVYKYKSPPPPPKKPYKYPSPPPPVYKYKSPP 728
Query: 147 -------AITPSSISTPPTKAPSSISTPPTKAPSTP 175
+ P S PP S PP +P P
Sbjct: 729 PPVYKYKSPPPPVHSPPPPHYVYSSPPPPVYSPPPP 836
>BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like protein
(clone pMG14) - common tobacco (fragment), partial (11%)
Length = 632
Score = 67.0 bits (162), Expect = 5e-12
Identities = 59/161 (36%), Positives = 81/161 (49%), Gaps = 5/161 (3%)
Frame = +1
Query: 7 VLMLVASLLVSSTFARSLSSSPVATPSPAGSPLAISPAVSS-PVPLMNAPSPSPFAVNYP 65
+L L +S +S +L SP A P+ SP A SP+ + P L + P PSP
Sbjct: 175 LLSLPSSPPSTSPSPHALLRSPTAPPAAPLSPPASSPSSPTRPSSLFSPPPPSPSR---- 342
Query: 66 TSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPV---PVTSS 122
+ PPP S SP P+ L ++T PS P P+ P A+SP S PV P ++S
Sbjct: 343 SRPPPPSTPSPLLPLPL-----LPLATEPSPFP-PSSPPPRPASSPSLSPPVLLFPPSAS 504
Query: 123 PSP-SPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPS 162
P P SPS S +PP S A+P +P S ST P++ P+
Sbjct: 505 PPPWSPSRPPSALAPPSRPSLPASPPSSPRS-STRPSRFPA 624
Score = 65.9 bits (159), Expect = 1e-11
Identities = 58/194 (29%), Positives = 92/194 (46%), Gaps = 27/194 (13%)
Frame = +1
Query: 18 STFARSL-SSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSP 76
++F+R+L ++ + PS +PL P + SPVP ++ P+SPP S SP
Sbjct: 52 ASFSRTLVQNNRPSQPSK*STPL---PFLLSPVPSWPTVVTRRRLLSLPSSPPSTS-PSP 219
Query: 77 ASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVAS-----SPVPVTSSP-------- 123
+ + +P P + +PP+ +PS PS+ + P S P P T SP
Sbjct: 220 HALLRSPTAPPAAPLSPPASSPSSPTRPSSLFSPPPPSPSRSRPPPPSTPSPLLPLPLLP 399
Query: 124 ---SPSPSAINSPPSPPPHASPAAAPAIT--PSSISTP---PTKAPSSISTP-----PTK 170
PSP +SPP P P +SP+ +P + P S S P P++ PS+++ P P
Sbjct: 400 LATEPSPFPPSSPP-PRPASSPSLSPPVLLFPPSASPPPWSPSRPPSALAPPSRPSLPAS 576
Query: 171 APSTPTNGATMNGF 184
PS+P + + F
Sbjct: 577 PPSSPRSSTRPSRF 618
>BQ751748 similar to PIR|T47216|T47 probable V-ATPase 20K chain [imported] -
Neurospora crassa, partial (82%)
Length = 642
Score = 66.6 bits (161), Expect = 6e-12
Identities = 52/151 (34%), Positives = 73/151 (47%), Gaps = 3/151 (1%)
Frame = -1
Query: 27 SPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFG---SPASPVAAP 83
+P +PSPA + + + S VP ++ SP PPPAS S +SP ++
Sbjct: 633 APTLSPSPATTSASATS*CSRAVPARSSRSPP--------RPPPASTAISVSTSSPSSST 478
Query: 84 AVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPA 143
P S + PP+ + P APS+S S V+S+ VT SP P P + S + P SP
Sbjct: 477 RSPPSSPTLPPASSSRPCSAPSSS--SIVSSTSPMVTLSP*PRP--VTSSRACPSLTSPT 310
Query: 144 AAPAITPSSISTPPTKAPSSISTPPTKAPST 174
+ PA T S A SS +TP +PST
Sbjct: 309 SGPASTAPSTPAVAASASSS*TTPVASSPST 217
Score = 42.7 bits (99), Expect = 9e-05
Identities = 44/135 (32%), Positives = 60/135 (43%), Gaps = 22/135 (16%)
Frame = -1
Query: 68 PPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPV-TSSPSPS 126
PPPA SP+ PA T S ++ S+A PA + + P AS+ + V TSSPS S
Sbjct: 642 PPPAPTLSPS-----PATTSASATS*CSRAV-PARSSRSPPRPPPASTAISVSTSSPSSS 481
Query: 127 PSAINSPPSPPPHASPAAAPAITPSSISTPPTKAP---------------------SSIS 165
+ S P+ PP +S + P PSS S + +P S S
Sbjct: 480 TRSPPSSPTLPPASS--SRPCSAPSSSSIVSSTSPMVTLSP*PRPVTSSRACPSLTSPTS 307
Query: 166 TPPTKAPSTPTNGAT 180
P + APSTP A+
Sbjct: 306 GPASTAPSTPAVAAS 262
Score = 42.7 bits (99), Expect = 9e-05
Identities = 43/118 (36%), Positives = 60/118 (50%), Gaps = 8/118 (6%)
Frame = -1
Query: 17 SSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSP 76
S+ + S SS +T SP SP + PA SS P +P + + +S P SP
Sbjct: 522 STAISVSTSSPSSSTRSPPSSP-TLPPASSS------RPCSAPSSSSIVSSTSPMVTLSP 364
Query: 77 -----ASPVAAPAVTPLSISTPPSQAPS-PAIAPSTSAN--SPVASSPVPVTSSPSPS 126
S A P++T + S P S APS PA+A S S++ +PVASSP SS +P+
Sbjct: 363 *PRPVTSSRACPSLTSPT-SGPASTAPSTPAVAASASSS*TTPVASSPSTSRSSTAPA 193
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.306 0.120 0.341
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,862,584
Number of Sequences: 36976
Number of extensions: 195021
Number of successful extensions: 16763
Number of sequences better than 10.0: 1707
Number of HSP's better than 10.0 without gapping: 4492
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 8783
length of query: 202
length of database: 9,014,727
effective HSP length: 91
effective length of query: 111
effective length of database: 5,649,911
effective search space: 627140121
effective search space used: 627140121
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144517.1


