

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144513.4 - phase: 0 /pseudo
(1015 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
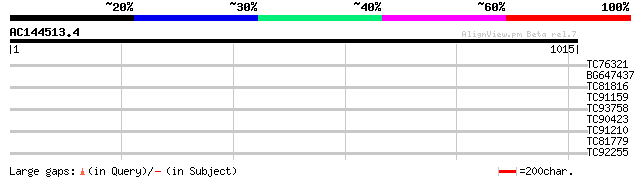
TC76321 homologue to GP|16033631|gb|AAL13304.1 leucine zipper-co... 41 0.002
BG647437 similar to PIR|G84555|G84 hypothetical protein At2g1773... 31 2.7
TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana taba... 30 3.6
TC91159 weakly similar to GP|20197705|gb|AAD17402.2 putative RIN... 30 3.6
TC93758 weakly similar to GP|22002164|gb|AAM88648.1 putative rec... 30 4.7
TC90423 similar to GP|20466586|gb|AAM20610.1 unknown protein {Ar... 30 4.7
TC91210 similar to GP|2291080|gb|AAC96017.1| defective gibberell... 30 6.1
TC81779 30 6.1
TC92255 weakly similar to GP|4049888|gb|AAC97848.1| ORF MSV023 A... 29 8.0
>TC76321 homologue to GP|16033631|gb|AAL13304.1 leucine zipper-containing
protein {Euphorbia esula}, partial (92%)
Length = 1468
Score = 41.2 bits (95), Expect = 0.002
Identities = 25/79 (31%), Positives = 45/79 (56%)
Frame = +3
Query: 236 KTIIKWLKLVGVKTMVTALRSLK*LLVNWIAEVNKNMGTFLRM*KILKRPSLILKIVFLL 295
K + L+L+ +K ++ RSL+ LL+N +A +++++ +FLR +L +VF
Sbjct: 321 KLCFRNLRLITIKLILLGTRSLRKLLINLMALLDRSLLSFLRGLVLLN------SLVFFS 482
Query: 296 KKSLDGSRRKKIFWMVCWS 314
KSL+G RK W + +S
Sbjct: 483 TKSLEGDLRKPTLWWLRFS 539
>BG647437 similar to PIR|G84555|G84 hypothetical protein At2g17730 [imported]
- Arabidopsis thaliana, partial (48%)
Length = 693
Score = 30.8 bits (68), Expect = 2.7
Identities = 31/101 (30%), Positives = 48/101 (46%), Gaps = 1/101 (0%)
Frame = +3
Query: 647 FMVSKLLLMPPALLTFFMQMAIFSFVRLTLMRPLLYPTFFLNTK-LFLARKLILISQIWS 705
F++ LLL P +L F TL L+ F++ + +FL L W
Sbjct: 3 FIII*LLLKPKSLYHQFQ-------TERTLPILLVLVVLFVSIRAIFLCENL----DKWI 149
Query: 706 LAQIPVLVLKIYSELISLSTSRIIFLSIWGFLLSLEDLKLK 746
L QI VL++ + S L+ +S +++ L FLLSLE L+
Sbjct: 150 LYQIHVLLVLLCSVLLGISLTKLNNLVPSQFLLSLETFSLR 272
>TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana tabacum},
partial (13%)
Length = 663
Score = 30.4 bits (67), Expect = 3.6
Identities = 36/115 (31%), Positives = 54/115 (46%), Gaps = 6/115 (5%)
Frame = -2
Query: 645 VTFMVSKLLLMPPALLTFFMQMAIFSFVR--LTLMRPLLYPTFF----LNTKLFLARKLI 698
+ F++S+ LL+ L FF + +F+F R + P Y T F L LFL L+
Sbjct: 644 LVFILSRFLLL--LFLVFFFLLLLFTFFRGFFYINIPYRYFTLFFLILLLLFLFLILILL 471
Query: 699 LISQIWSLAQIPVLVLKIYSELISLSTSRIIFLSIWGFLLSLEDLKLKILSLLWI 753
++ I L I V +L ++ L+ L L F L L L L +L LL+I
Sbjct: 470 FLTSI--LTFIIVFILVLFLLLVFLRFLLFTILVFVFFSLLLLFLFLLLLILLFI 312
>TC91159 weakly similar to GP|20197705|gb|AAD17402.2 putative RING-H2 zinc
finger protein {Arabidopsis thaliana}, partial (21%)
Length = 695
Score = 30.4 bits (67), Expect = 3.6
Identities = 14/32 (43%), Positives = 23/32 (71%)
Frame = +2
Query: 18 IKNLSHVFSMVLSLILLLLIVLLVIAIDQVVL 49
I + HVFS+ L IL++LIV L++++D + L
Sbjct: 464 ISIIQHVFSLGLKHILIVLIVELLLSLDDIYL 559
>TC93758 weakly similar to GP|22002164|gb|AAM88648.1 putative receptor-like
protein kinase {Oryza sativa (japonica cultivar-group)},
partial (3%)
Length = 814
Score = 30.0 bits (66), Expect = 4.7
Identities = 26/89 (29%), Positives = 42/89 (46%), Gaps = 7/89 (7%)
Frame = -2
Query: 674 LTLMRP--LLYPTFFLNTKLFLARKL-----ILISQIWSLAQIPVLVLKIYSELISLSTS 726
+T M+P ++Y + NT ++AR + LI+ +WS + LI + T
Sbjct: 324 VTEMKPK*MIYVLLYRNTPTYVARTVNTLFASLINPVWS------------NILI*IWTI 181
Query: 727 RIIFLSIWGFLLSLEDLKLKILSLLWIEF 755
R I S W ++K ILS+LWI +
Sbjct: 180 RCIICSAWNM-----NVKGAILSVLWISY 109
>TC90423 similar to GP|20466586|gb|AAM20610.1 unknown protein {Arabidopsis
thaliana}, partial (14%)
Length = 1114
Score = 30.0 bits (66), Expect = 4.7
Identities = 16/41 (39%), Positives = 22/41 (53%)
Frame = +2
Query: 643 IRVTFMVSKLLLMPPALLTFFMQMAIFSFVRLTLMRPLLYP 683
+R ++ LL+ +L FM+M IFSF R MR L P
Sbjct: 809 VREGLLIFNTLLLQIFMLLHFMEMVIFSFTREN*MRNLCEP 931
>TC91210 similar to GP|2291080|gb|AAC96017.1| defective gibberellin
3B-hydroxylase {Pisum sativum}, partial (44%)
Length = 659
Score = 29.6 bits (65), Expect = 6.1
Identities = 17/40 (42%), Positives = 24/40 (59%)
Frame = +2
Query: 353 TL*LTHMKKFGEITITFMRSLLPILRRCLLVLDLLFPLML 392
TL H K F +T+ + P+LRR LLV +LFPL++
Sbjct: 170 TLISIHYKNFLNLTLGTTLMITPLLRRVLLV--VLFPLLI 283
>TC81779
Length = 478
Score = 29.6 bits (65), Expect = 6.1
Identities = 18/39 (46%), Positives = 24/39 (61%)
Frame = +2
Query: 101 ILLICYIILENMIVGLFLGILT*YYIVLKS*EEILLFLY 139
I LIC ++LEN V G++ *YYIVL * I +F +
Sbjct: 362 ISLICLLVLENGDV--VFGVVK*YYIVLAI*RYISIFQF 472
>TC92255 weakly similar to GP|4049888|gb|AAC97848.1| ORF MSV023 ALI motif
gene family protein {Melanoplus sanguinipes
entomopoxvirus}, partial (8%)
Length = 886
Score = 29.3 bits (64), Expect = 8.0
Identities = 16/40 (40%), Positives = 23/40 (57%)
Frame = -3
Query: 701 SQIWSLAQIPVLVLKIYSELISLSTSRIIFLSIWGFLLSL 740
S +S+ +L+LK S + LSTS + SIW F+ SL
Sbjct: 179 SSSFSVFNASILLLKWASSFLFLSTSCLWLASIWNFINSL 60
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.368 0.167 0.619
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 36,509,345
Number of Sequences: 36976
Number of extensions: 608841
Number of successful extensions: 10600
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 2673
Number of HSP's successfully gapped in prelim test: 596
Number of HSP's that attempted gapping in prelim test: 7524
Number of HSP's gapped (non-prelim): 3945
length of query: 1015
length of database: 9,014,727
effective HSP length: 106
effective length of query: 909
effective length of database: 5,095,271
effective search space: 4631601339
effective search space used: 4631601339
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC144513.4


