

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
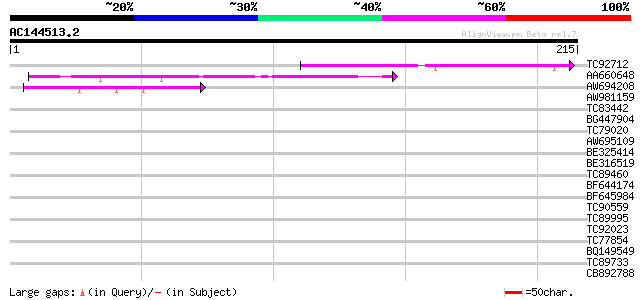
Query= AC144513.2 - phase: 0
(215 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC92712 similar to PIR|T48228|T48228 probable protein kinase - A... 78 3e-15
AA660648 weakly similar to GP|16323147|gb At1g10380/F14N23_32 {A... 58 2e-09
AW694208 weakly similar to GP|5669670|gb|A receptor-like kinase ... 45 3e-05
AW981159 weakly similar to GP|11071985|dbj putative receptor kin... 38 0.003
TC83442 weakly similar to GP|13377498|gb|AAK20737.1 TAK19-1 {Tri... 35 0.017
BG447904 35 0.028
TC79020 34 0.037
AW695109 33 0.063
BE325414 32 0.14
BE316519 similar to GP|15293303|gb| unknown protein {Arabidopsis... 32 0.24
TC89460 similar to GP|11071990|dbj|BAB17335. putative receptor k... 32 0.24
BF644174 weakly similar to PIR|E96721|E967 hypothetical protein ... 32 0.24
BF645984 similar to GP|11071992|dbj putative receptor kinase {Or... 31 0.31
TC90559 similar to PIR|T43291|T43291 laminin alpha chain - Caeno... 30 0.91
TC89995 similar to GP|18425245|gb|AAL69423.1 Putative wall-assoc... 30 0.91
TC92023 similar to GP|20198310|gb|AAM15518.1 Expressed protein {... 29 1.2
TC77854 similar to PIR|T51592|T51592 galactokinase (EC 2.7.1.6) ... 29 1.6
BQ149549 similar to PIR|B86270|B86 hypothetical protein AAF81295... 28 2.7
TC89733 weakly similar to GP|19571157|dbj|BAB86580. hypothetical... 27 5.9
CB892788 27 5.9
>TC92712 similar to PIR|T48228|T48228 probable protein kinase - Arabidopsis
thaliana, partial (2%)
Length = 511
Score = 77.8 bits (190), Expect = 3e-15
Identities = 39/106 (36%), Positives = 61/106 (56%), Gaps = 2/106 (1%)
Frame = +3
Query: 111 DESLPFNISTLNTVMLLNCSDNILQSPLNCSSNSICRQFEEKVEEGNGCM-NTLCCHYLK 169
+ +LPFNI++ NT++ LNC+ ++L+SPLNCS+ S C + C LCC Y
Sbjct: 6 NNTLPFNITSSNTIVYLNCTRDLLRSPLNCSAASACHAYINATVP--TCQTGPLCCTYRT 179
Query: 170 DSVMNSHKIRLRVGSCTAYTCLVDFKPDEPFKKWNY-GIELQWKPP 214
NS++IR+R C+AY+ V+ +W+ G+E+QW P
Sbjct: 180 GGSSNSYQIRVRSSGCSAYSSFVNLDSGLAVNRWSRPGLEIQWMSP 317
>AA660648 weakly similar to GP|16323147|gb At1g10380/F14N23_32 {Arabidopsis
thaliana}, partial (42%)
Length = 637
Score = 58.2 bits (139), Expect = 2e-09
Identities = 47/143 (32%), Positives = 67/143 (45%), Gaps = 3/143 (2%)
Frame = +1
Query: 8 SLKMVILLLFLLITGLGSVINASPCS-NCGDFEVPYPLSTNDDCGDKRYK--IYCNNDSL 64
SL ++ + LFLL S+I + C NCG + YP + CGD R++ I C+ L
Sbjct: 49 SLPLLTIFLFLL----PSLIKSQMCQRNCGKETLKYPFGSGPGCGDPRFQPHITCSQQKL 216
Query: 65 EFLSATGTYYKILKIDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNTV 124
F + TG+ Y I ID + + I P + TC S L G LD + PF
Sbjct: 217 TFTTHTGS-YPITSIDYANQVIYISDPTM--STC-SCTLPSKGFGLDWNAPFTFDDSTIF 384
Query: 125 MLLNCSDNILQSPLNCSSNSICR 147
L++CS N S+SIC+
Sbjct: 385 ALVDCSMN---------SSSICK 426
>AW694208 weakly similar to GP|5669670|gb|A receptor-like kinase {Zea mays},
partial (6%)
Length = 658
Score = 44.7 bits (104), Expect = 3e-05
Identities = 27/76 (35%), Positives = 38/76 (49%), Gaps = 7/76 (9%)
Frame = +2
Query: 6 WNSLKMVILLLFLLITGLGS---VINASPCSNCGDFE-VPYPLSTNDD---CGDKRYKIY 58
W + +V+LL F L+ GS +A P S+CG + YP DD CGD RY++
Sbjct: 74 WVRVILVLLLPFSLLIQQGSSSTTTSACPPSSCGKISNISYPFRLKDDPIHCGDSRYELS 253
Query: 59 CNNDSLEFLSATGTYY 74
C N+ +G YY
Sbjct: 254 CENNVTTLNLYSGKYY 301
>AW981159 weakly similar to GP|11071985|dbj putative receptor kinase {Oryza
sativa (japonica cultivar-group)}, partial (4%)
Length = 739
Score = 37.7 bits (86), Expect = 0.003
Identities = 25/81 (30%), Positives = 36/81 (43%), Gaps = 10/81 (12%)
Frame = -1
Query: 4 CIWNSLKMVILLLFLLITGLGSVINAS------PCSNCGDF-EVPYPLSTNDD---CGDK 53
C +S ++LLF I + I + P S+CG + YP D CGDK
Sbjct: 697 CSSSSYVQTLVLLFASILLISHQIVFAKHHQPWPTSSCGKITNITYPFRLKTDPNHCGDK 518
Query: 54 RYKIYCNNDSLEFLSATGTYY 74
RY++ CN + +G YY
Sbjct: 517 RYELDCNENGPSLTMFSGKYY 455
>TC83442 weakly similar to GP|13377498|gb|AAK20737.1 TAK19-1 {Triticum
aestivum}, partial (6%)
Length = 674
Score = 35.4 bits (80), Expect = 0.017
Identities = 28/105 (26%), Positives = 46/105 (43%), Gaps = 10/105 (9%)
Frame = +3
Query: 6 WNSLKMVILLLFLLITGLG----SVINASPC--SNCGDFE-VPYPLSTNDD---CGDKRY 55
W + +V+LL F + G S C S+CG + YP DD CGD RY
Sbjct: 135 WMRVILVLLLPFSFLFQQGTCSASTEQEHTCLPSSCGKISNISYPFRLKDDPIHCGDSRY 314
Query: 56 KIYCNNDSLEFLSATGTYYKILKIDTSANKLVIKPPNIFKHTCYS 100
++ C N+ +G Y+ + I+ + + + P + + C S
Sbjct: 315 ELSCANNVTTLYLYSGKYH-VQSINYNNFTIRLVDPGVQQSNCSS 446
>BG447904
Length = 211
Score = 34.7 bits (78), Expect = 0.028
Identities = 17/48 (35%), Positives = 24/48 (49%), Gaps = 4/48 (8%)
Frame = +3
Query: 31 PCSNCGDF-EVPYPLSTNDD---CGDKRYKIYCNNDSLEFLSATGTYY 74
P S+CG + +P DD CGD RY++ C N+ +G YY
Sbjct: 57 PPSSCGKITNITHPFRLKDDPTSCGDPRYELSCENNITMLTLLSGKYY 200
>TC79020
Length = 1268
Score = 34.3 bits (77), Expect = 0.037
Identities = 24/76 (31%), Positives = 37/76 (48%), Gaps = 8/76 (10%)
Frame = +2
Query: 7 NSLKMVILLLFLLIT--GLGSVINASPCSN-CGDFEVPYPLSTND---DCGDKRYKIYC- 59
+SL V+LLL L+ + + + CS CG + P D +CGD+RY + C
Sbjct: 152 SSLTFVVLLLILIFSYNKVYGDEHQDSCSRWCGVHNIRSPFRLKDSPKECGDERYNLSCE 331
Query: 60 -NNDSLEFLSATGTYY 74
NN + + + G YY
Sbjct: 332 DNNQLILYHDSFGKYY 379
>AW695109
Length = 603
Score = 33.5 bits (75), Expect = 0.063
Identities = 16/48 (33%), Positives = 24/48 (49%), Gaps = 4/48 (8%)
Frame = +1
Query: 31 PCSNCGDFE-VPYPLSTNDD---CGDKRYKIYCNNDSLEFLSATGTYY 74
P S+CG + +P +D CGD RY++ C N+ +G YY
Sbjct: 148 PSSSCGTIRNIKHPFRLQNDPANCGDPRYELSCENNITTLTLFSGKYY 291
>BE325414
Length = 508
Score = 32.3 bits (72), Expect = 0.14
Identities = 20/60 (33%), Positives = 27/60 (44%), Gaps = 9/60 (15%)
Frame = +1
Query: 24 GSVINASPCSN-CGDFEVPYPLSTNDD---CGDKRYKIYCNNDS-----LEFLSATGTYY 74
G V + CS CG + +P D CGDKRY + C +++ EF G YY
Sbjct: 31 GDVEHQESCSRRCGVHSISHPFRLKDSPEKCGDKRYILSCEDNNQLILYYEFEEYHGKYY 210
>BE316519 similar to GP|15293303|gb| unknown protein {Arabidopsis
thaliana}, partial (26%)
Length = 292
Score = 31.6 bits (70), Expect = 0.24
Identities = 18/64 (28%), Positives = 33/64 (51%), Gaps = 3/64 (4%)
Frame = +1
Query: 13 ILLLFLLITGLGSVINASPCSNCGDFEVPYPLSTNDDCGDKRYK--IYCNN-DSLEFLSA 69
++ LF ++T S + + +CG + YP S +D CG Y+ + C++ LE +
Sbjct: 19 LIYLFSILTKSQSTLCRT---SCGSIPIQYPFSIDDGCGSPYYRFILSCSDTQKLELRTP 189
Query: 70 TGTY 73
+G Y
Sbjct: 190SGRY 201
>TC89460 similar to GP|11071990|dbj|BAB17335. putative receptor kinase
{Oryza sativa (japonica cultivar-group)}, partial (3%)
Length = 720
Score = 31.6 bits (70), Expect = 0.24
Identities = 28/127 (22%), Positives = 52/127 (40%), Gaps = 20/127 (15%)
Frame = +1
Query: 31 PCSNCGDF-EVPYPLSTNDD---CGDKRYKIYCNNDSLEFLSATGTYYKILKIDTSANKL 86
P S+CG + +P +D CGD +Y++ C N+ +G YY + +I+ +
Sbjct: 202 PPSSCGKITNISHPFRLMNDPTSCGDPKYELSCENNITVLTLFSGKYY-VQEINYVNYTI 378
Query: 87 VIKPPNIFKHTCYS--------SDLIGGGLVLDESLPFNISTL--------NTVMLLNCS 130
+ P I + C S S+ + ++ P+ I + ++ LNCS
Sbjct: 379 RLVDPVIEEGNCSSIPPYFLTKSNFTSSFIHINNEDPYQIMDVWYYSDHRYGHIIYLNCS 558
Query: 131 DNILQSP 137
+ P
Sbjct: 559 KQVNDDP 579
>BF644174 weakly similar to PIR|E96721|E967 hypothetical protein T17F3.6
[imported] - Arabidopsis thaliana, partial (13%)
Length = 362
Score = 31.6 bits (70), Expect = 0.24
Identities = 15/62 (24%), Positives = 29/62 (46%), Gaps = 2/62 (3%)
Frame = +1
Query: 33 SNCGDFEV--PYPLSTNDDCGDKRYKIYCNNDSLEFLSATGTYYKILKIDTSANKLVIKP 90
S C F+ P+P ST CG +++ C+ F++ + IL ++ +++ P
Sbjct: 58 SPCPPFQSTPPFPFSTTPGCGHPSFQLTCSTPH-SFITINNLTFSILSYKPNSTSIILSP 234
Query: 91 PN 92
N
Sbjct: 235 HN 240
>BF645984 similar to GP|11071992|dbj putative receptor kinase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 654
Score = 31.2 bits (69), Expect = 0.31
Identities = 20/74 (27%), Positives = 33/74 (44%), Gaps = 4/74 (5%)
Frame = +3
Query: 31 PCSNCGDFE-VPYPLSTNDD---CGDKRYKIYCNNDSLEFLSATGTYYKILKIDTSANKL 86
P S+CG + YP +D CGD RY++ C ++ L Y + I+ + +
Sbjct: 168 PPSSCGKISNIKYPFRLKNDPATCGDPRYELSCEK-NITMLKLLSGKYHVKSINYNNYTI 344
Query: 87 VIKPPNIFKHTCYS 100
+ P I + C S
Sbjct: 345 RLVDPGIKEGDCSS 386
>TC90559 similar to PIR|T43291|T43291 laminin alpha chain - Caenorhabditis
elegans, partial (1%)
Length = 920
Score = 29.6 bits (65), Expect = 0.91
Identities = 16/59 (27%), Positives = 28/59 (47%)
Frame = -1
Query: 71 GTYYKILKIDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNTVMLLNC 129
GT + I+ ++L+I P +TCY + + G + S F+ S + +LNC
Sbjct: 317 GTTFSIMLSSLMNSRLIIPP---LMYTCYLTSFVLGVFFVGFSFSFSFSRVKFSRILNC 150
>TC89995 similar to GP|18425245|gb|AAL69423.1 Putative wall-associated
protein kinase {Oryza sativa}, partial (4%)
Length = 840
Score = 29.6 bits (65), Expect = 0.91
Identities = 46/206 (22%), Positives = 76/206 (36%), Gaps = 21/206 (10%)
Frame = +1
Query: 12 VILLLFLLITGLGSVINASP-----CSN-CGDFEVPYPLSTNDDC--------GDKRYKI 57
++LL ++T + + + P C N CGD ++PYP ++ D Y +
Sbjct: 64 ILLLTIFMVTIVFASNESFPSSLPGCKNTCGDVKIPYPFGISNSLIPNHGPCFLDPFYNL 243
Query: 58 YCNNDSLEFLSATGTYYKILKIDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLDESL--- 114
C ND+ K+L D + + + + SS+ G ++S
Sbjct: 244 TCENDT-----------KLLWSDIQVSNISVIEGQLELLFFVSSNCGNGNGSYNQSFLDI 390
Query: 115 ---PFNISTLNTVMLLNCSDNILQSPLNCSSNSICRQFEEKVEEGNGCMNTLCCHYLKDS 171
F++ST L + C+S R EK GC+ T+C D
Sbjct: 391 FDGRFSLSTRENKFL----------TVGCNSIGFLRSTYEKETFYTGCL-TIC-----DG 522
Query: 172 VMNSHKIRLRVGSCTAY-TCLVDFKP 196
N R+ G+C+ C VD P
Sbjct: 523 TRN----RIENGACSGIGCCQVDIPP 588
>TC92023 similar to GP|20198310|gb|AAM15518.1 Expressed protein {Arabidopsis
thaliana}, partial (66%)
Length = 580
Score = 29.3 bits (64), Expect = 1.2
Identities = 15/32 (46%), Positives = 18/32 (55%)
Frame = +3
Query: 58 YCNNDSLEFLSATGTYYKILKIDTSANKLVIK 89
Y N D L LS YYKIL++D AN I+
Sbjct: 234 YLNFDFLSTLSKPKDYYKILEVDYDANDDTIR 329
>TC77854 similar to PIR|T51592|T51592 galactokinase (EC 2.7.1.6) [validated]
- Arabidopsis thaliana, partial (92%)
Length = 1930
Score = 28.9 bits (63), Expect = 1.6
Identities = 13/31 (41%), Positives = 21/31 (66%)
Frame = +1
Query: 3 PCIWNSLKMVILLLFLLITGLGSVINASPCS 33
P +W SL+ ++LL ++ITGL +V+ CS
Sbjct: 880 PILWQSLRRLLLLPQIIITGLLNVV*LLLCS 972
>BQ149549 similar to PIR|B86270|B86 hypothetical protein AAF81295.1
[imported] - Arabidopsis thaliana, partial (29%)
Length = 635
Score = 28.1 bits (61), Expect = 2.7
Identities = 25/78 (32%), Positives = 38/78 (48%), Gaps = 14/78 (17%)
Frame = +3
Query: 3 PCIWNSLKMVILLLF--LLITGLGSVIN------------ASPCSNCGDFEVPYPLSTND 48
P N K+ LLLF L + +GSV+ A+ CSN G ++ ++++
Sbjct: 54 PSSINIYKLYSLLLFISLFVVNMGSVLFLIMLILPLVYNVAADCSN-GACKLGDECNSDE 230
Query: 49 DCGDKRYKIYCNNDSLEF 66
DCG +YC + SLEF
Sbjct: 231 DCGT---GLYCFDCSLEF 275
>TC89733 weakly similar to GP|19571157|dbj|BAB86580. hypothetical protein
{Oryza sativa (japonica cultivar-group)}, partial (27%)
Length = 768
Score = 26.9 bits (58), Expect = 5.9
Identities = 11/26 (42%), Positives = 14/26 (53%)
Frame = +1
Query: 190 CLVDFKPDEPFKKWNYGIELQWKPPN 215
CL+ +PD F KW +WKP N
Sbjct: 322 CLIHGRPDSDFLKW------RWKPDN 381
>CB892788
Length = 886
Score = 26.9 bits (58), Expect = 5.9
Identities = 15/64 (23%), Positives = 31/64 (48%), Gaps = 12/64 (18%)
Frame = +2
Query: 13 ILLLFLLITGLGSVINASPCSN-----CGDFEVPYPL-----STNDDC--GDKRYKIYCN 60
+ +L +++ +G+ + A +N CG E+PYP +T+ +C G + + CN
Sbjct: 491 VYVLAMVVVLVGTTVAADSATNTCSHTCGGQEIPYPFGIDDNNTSSECFLGRRLIPLTCN 670
Query: 61 NDSL 64
+
Sbjct: 671 ESKI 682
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.139 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,517,952
Number of Sequences: 36976
Number of extensions: 153766
Number of successful extensions: 821
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 816
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 820
length of query: 215
length of database: 9,014,727
effective HSP length: 92
effective length of query: 123
effective length of database: 5,612,935
effective search space: 690391005
effective search space used: 690391005
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144513.2


