

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144503.9 + phase: 0 /pseudo
(346 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
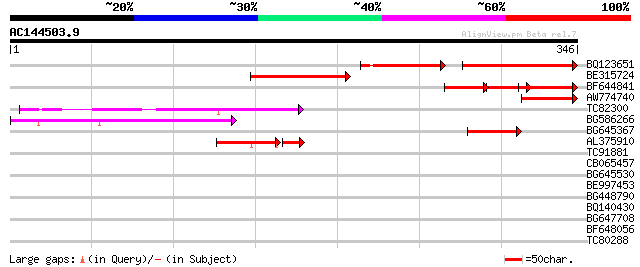
Score E
Sequences producing significant alignments: (bits) Value
BQ123651 86 5e-28
BE315724 118 4e-27
BF644841 similar to GP|18175662|gb| putative receptor protein ki... 47 3e-15
AW774740 65 5e-11
TC82300 56 2e-08
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 54 1e-07
BG645367 51 7e-07
AL375910 43 3e-06
TC91881 38 0.005
CB065457 27 0.051
BG645530 similar to SP|P42347|P3K1_ Phosphatidylinositol 3-kinas... 33 0.21
BE997453 28 0.23
BG448790 29 0.24
BQ140430 similar to GP|21592347|gb| unknown {Arabidopsis thalian... 28 5.2
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 28 6.8
BF648056 similar to GP|14532658|gb unknown protein {Arabidopsis ... 27 8.9
TC80288 weakly similar to PIR|T06029|T06029 hypothetical protein... 27 8.9
>BQ123651
Length = 654
Score = 85.5 bits (210), Expect(2) = 5e-28
Identities = 41/70 (58%), Positives = 53/70 (75%)
Frame = -2
Query: 277 WAVKSQHENQVSVAEMRMLRWMSRKTRHDRIKNDTIRERVGVAPIVEKLVENRLR*FGHV 336
WA HE ++S A+ RMLRWM TR D+I+ND+I RVG+APIVEK++E+RLR FG+V
Sbjct: 248 WAE*CHHERRISEAKKRMLRWMCGNTRRDKIRNDSISVRVGIAPIVEKMIESRLRWFGNV 69
Query: 337 ERRPVDAVVR 346
RP D+VVR
Sbjct: 68 WGRPGDSVVR 39
Score = 56.6 bits (135), Expect(2) = 5e-28
Identities = 30/52 (57%), Positives = 37/52 (70%)
Frame = -3
Query: 215 YLGSFV*NDGEIEADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAV 266
YLGS + + + DV++RIQA +K R GVLCDKKVPLK+KGKFY T V
Sbjct: 421 YLGSVI-QK*QKQRDVNNRIQARLMK*RSPLGVLCDKKVPLKIKGKFYLTTV 269
>BE315724
Length = 392
Score = 118 bits (295), Expect = 4e-27
Identities = 58/61 (95%), Positives = 59/61 (96%)
Frame = -2
Query: 148 DVVLVGESREEVNGRLETWRQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHII 207
+VVLVGESREEVNGRLETWRQALEAY FRLSRS TEYMECNFSGRRSRSTLEVKVGDHII
Sbjct: 184 NVVLVGESREEVNGRLETWRQALEAYEFRLSRSKTEYMECNFSGRRSRSTLEVKVGDHII 5
Query: 208 P 208
P
Sbjct: 4 P 2
>BF644841 similar to GP|18175662|gb| putative receptor protein kinase
{Arabidopsis thaliana}, partial (1%)
Length = 676
Score = 47.0 bits (110), Expect(3) = 3e-15
Identities = 19/27 (70%), Positives = 23/27 (84%)
Frame = +2
Query: 266 VRPALLYGTECWAVKSQHENQVSVAEM 292
+RPA+LY ECWAVK+QHEN+VS EM
Sbjct: 158 LRPAMLYAMECWAVKNQHENKVSYVEM 238
Score = 42.7 bits (99), Expect(3) = 3e-15
Identities = 21/36 (58%), Positives = 27/36 (74%)
Frame = +1
Query: 311 TIRERVGVAPIVEKLVENRLR*FGHVERRPVDAVVR 346
T ERV V+ IVE +V+N+LR FG+V RRP+ VVR
Sbjct: 298 TTLERVWVSSIVENMVDNKLRWFGYVXRRPIHFVVR 405
Score = 28.9 bits (63), Expect(3) = 3e-15
Identities = 12/27 (44%), Positives = 18/27 (66%)
Frame = +3
Query: 292 MRMLRWMSRKTRHDRIKNDTIRERVGV 318
M+ML W+ KTR +N+ IRE +G+
Sbjct: 243 MKMLCWICGKTRRYPFQNNNIRESLGI 323
>AW774740
Length = 739
Score = 64.7 bits (156), Expect = 5e-11
Identities = 32/34 (94%), Positives = 32/34 (94%)
Frame = +2
Query: 313 RERVGVAPIVEKLVENRLR*FGHVERRPVDAVVR 346
RERVGVAPIVEKLVENR R FGHVERRPVDAVVR
Sbjct: 2 RERVGVAPIVEKLVENRFRWFGHVERRPVDAVVR 103
>TC82300
Length = 589
Score = 56.2 bits (134), Expect = 2e-08
Identities = 56/176 (31%), Positives = 75/176 (41%), Gaps = 3/176 (1%)
Frame = +2
Query: 7 IERRLRKEARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDLEKTYDRVP 66
IE +LR N F F+PGR TM+ L++ + + D +
Sbjct: 131 IENKLRGHQHHI-NLFSFIPGRSTMK------------------LYICYEN*W*DVDGIN 253
Query: 67 REILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTLSPYLFT 126
W L KG I MYE + SV T G EDFPIT GV+ GS LSPY
Sbjct: 254 *TYTWLLLM*KGNNI--------MYERGTASVTTLGGNVEDFPITKGVN*GSVLSPYFSI 409
Query: 127 ---LVLDVLTEHIQELAPRCMLFADVVLVGESREEVNGRLETWRQALEAYGFRLSR 179
V + + ++ ++VLV ES E +N L+ Q LE Y RL+R
Sbjct: 410 GSGFVY*T*AREVVTGSQGMLVGNEIVLVEES*EAINKILKIR*QVLETYDNRLTR 577
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 53.5 bits (127), Expect = 1e-07
Identities = 37/143 (25%), Positives = 66/143 (45%), Gaps = 5/143 (3%)
Frame = -3
Query: 1 KLWERVIERRLRKEAR--VTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHL---VF 55
K+ +++ +R++ R ++ +Q F+PGR + + + +++ R H+ V
Sbjct: 736 KIIAKILSKRMQPLLRSIISPSQSAFVPGRAISDNVLITHKILHYLRQSGAKKHVSMAVK 557
Query: 56 IDLEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVH 115
D+ K YDR+ L + L + G +I I + S S G + G+
Sbjct: 556 TDMTKAYDRIAWNFLREVLTRLGFHGIWISWIMECVSTVSYSFLINGGPQGRVLPSRGLR 377
Query: 116 QGSTLSPYLFTLVLDVLTEHIQE 138
QG LSPYLF L +VL+ Q+
Sbjct: 376 QGDPLSPYLFILCTEVLSGLCQQ 308
>BG645367
Length = 739
Score = 50.8 bits (120), Expect = 7e-07
Identities = 20/33 (60%), Positives = 28/33 (84%)
Frame = +1
Query: 280 KSQHENQVSVAEMRMLRWMSRKTRHDRIKNDTI 312
K+QH+N++S +EMRML WM KTRH RI+N+T+
Sbjct: 640 KNQHKNKISTSEMRMLHWMCGKTRHGRIRNETL 738
>AL375910
Length = 244
Score = 43.1 bits (100), Expect(2) = 3e-06
Identities = 28/48 (58%), Positives = 31/48 (64%), Gaps = 9/48 (18%)
Frame = -1
Query: 127 LVLDVLTEHIQELAPRCMLF-------ADVVLVGESREEVNG--RLET 165
+ LDVLTE+IQELA RCM F D VL GE REE+ G RLET
Sbjct: 181 IFLDVLTENIQELALRCMFFFFLFFFTDDAVLHGELREELMGG*RLET 38
Score = 25.0 bits (53), Expect(2) = 3e-06
Identities = 12/14 (85%), Positives = 12/14 (85%)
Frame = -2
Query: 167 RQALEAYGFRLSRS 180
RQALEAYGF LS S
Sbjct: 42 RQALEAYGFCLSGS 1
>TC91881
Length = 860
Score = 38.1 bits (87), Expect = 0.005
Identities = 19/49 (38%), Positives = 32/49 (64%)
Frame = +3
Query: 261 FYRTAVRPALLYGTECWAVKSQHENQVSVAEMRMLRWMSRKTRHDRIKN 309
+Y T+++ +++ +CWAVKSQ E S+A+MRML + ++IKN
Sbjct: 315 YYCTSLK-LVIFRIDCWAVKSQ*ERMTSIAKMRMLW*ICGHIGRNKIKN 458
Score = 29.3 bits (64), Expect = 2.3
Identities = 15/35 (42%), Positives = 20/35 (56%)
Frame = +1
Query: 168 QALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKV 202
Q L F L+RS EY+ C FS + +L+VKV
Sbjct: 16 QTLNFNDFCLTRSKLEYIGCRFSRGHTNVSLDVKV 120
>CB065457
Length = 831
Score = 27.3 bits (59), Expect(2) = 0.051
Identities = 11/16 (68%), Positives = 13/16 (80%)
Frame = +3
Query: 230 VSHRIQAEWLKWRRAS 245
V+H IQ WLKW+RAS
Sbjct: 48 VNH*IQDGWLKWQRAS 95
Score = 26.2 bits (56), Expect(2) = 0.051
Identities = 12/14 (85%), Positives = 12/14 (85%)
Frame = +1
Query: 213 FKYLGSFV*NDGEI 226
FKYLGS V NDGEI
Sbjct: 1 FKYLGSKVHNDGEI 42
>BG645530 similar to SP|P42347|P3K1_ Phosphatidylinositol 3-kinase root
isoform (EC 2.7.1.137) (PI3-kinase) (PtdIns-3-kinase)
(PI3K), partial (7%)
Length = 821
Score = 32.7 bits (73), Expect = 0.21
Identities = 11/19 (57%), Positives = 14/19 (72%)
Frame = +1
Query: 262 YRTAVRPALLYGTECWAVK 280
YRT +RP + Y T+CW VK
Sbjct: 757 YRTVIRPMIFYETKCWGVK 813
>BE997453
Length = 795
Score = 28.1 bits (61), Expect(2) = 0.23
Identities = 13/24 (54%), Positives = 16/24 (66%)
Frame = +2
Query: 321 IVEKLVENRLR*FGHVERRPVDAV 344
I KLV L FGHV RRP++A+
Sbjct: 83 IANKLVRPCLGWFGHVRRRPIEAL 154
Score = 23.1 bits (48), Expect(2) = 0.23
Identities = 12/25 (48%), Positives = 14/25 (56%)
Frame = +1
Query: 293 RMLRWMSRKTRHDRIKNDTIRERVG 317
RML MS TR DRI+N +G
Sbjct: 1 RML*RMSGHTR*DRIRNKVFERNLG 75
>BG448790
Length = 652
Score = 28.9 bits (63), Expect(2) = 0.24
Identities = 17/46 (36%), Positives = 25/46 (53%), Gaps = 1/46 (2%)
Frame = +2
Query: 145 LFADVVLVG-ESREEVNGRLETWRQALEAYGFRLSRSTTEYMECNF 189
+FA +V VG +S N +L+ W +E + F S S E +EC F
Sbjct: 374 VFASLVKVG*KS*GYRNEKLKVWSHIVETHSFG*SXSKIENIECKF 511
Score = 22.3 bits (46), Expect(2) = 0.24
Identities = 12/40 (30%), Positives = 23/40 (57%)
Frame = +3
Query: 198 LEVKVGDHIIPQVTRFKYLGSFV*NDGEIEADVSHRIQAE 237
++ K+G+ I+PQV+ K + +D +IE +H + E
Sbjct: 534 VKAKIGEQIMPQVSFSKKV-----SDVDIE*GFNHEFKRE 638
>BQ140430 similar to GP|21592347|gb| unknown {Arabidopsis thaliana}, partial
(31%)
Length = 409
Score = 28.1 bits (61), Expect = 5.2
Identities = 12/40 (30%), Positives = 22/40 (55%)
Frame = +3
Query: 152 VGESREEVNGRLETWRQALEAYGFRLSRSTTEYMECNFSG 191
+ ESR E+ GR+++ +Q L+ + +L Y + F G
Sbjct: 168 LNESRSELLGRIQSLKQDLQGWRSKLDTQVKVYRDTKFHG 287
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 27.7 bits (60), Expect = 6.8
Identities = 12/21 (57%), Positives = 15/21 (71%)
Frame = +1
Query: 113 GVHQGSTLSPYLFTLVLDVLT 133
G+ QG LSPYLF L +VL+
Sbjct: 13 GLRQGDPLSPYLFILCANVLS 75
>BF648056 similar to GP|14532658|gb unknown protein {Arabidopsis thaliana},
partial (76%)
Length = 652
Score = 27.3 bits (59), Expect = 8.9
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = +3
Query: 167 RQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHII 207
R L AYG ++ S++EY++ N S S +V DH +
Sbjct: 141 RGGLGAYGGKILGSSSEYVQSNISRYFSDPQYYFQVNDHYV 263
>TC80288 weakly similar to PIR|T06029|T06029 hypothetical protein T28I19.100
- Arabidopsis thaliana, partial (14%)
Length = 1460
Score = 27.3 bits (59), Expect = 8.9
Identities = 19/68 (27%), Positives = 31/68 (44%)
Frame = +1
Query: 240 KWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECWAVKSQHENQVSVAEMRMLRWMS 299
K R + ++K K+K K R ++ +L +K NQ + +RM WM
Sbjct: 541 KNRMKKKISMEQKCRRKMKAKVKRWKMKEEMLR*-----MKMIMRNQRQIMIVRMRLWMK 705
Query: 300 RKTRHDRI 307
RKT+ R+
Sbjct: 706 RKTKKKRV 729
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,716,407
Number of Sequences: 36976
Number of extensions: 122147
Number of successful extensions: 704
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 700
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 704
length of query: 346
length of database: 9,014,727
effective HSP length: 97
effective length of query: 249
effective length of database: 5,428,055
effective search space: 1351585695
effective search space used: 1351585695
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144503.9


