

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144503.4 - phase: 0
(366 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
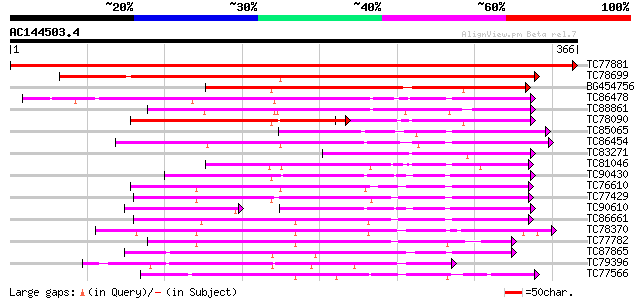
Score E
Sequences producing significant alignments: (bits) Value
TC77881 homologue to GP|15528439|emb|CAC69137. MEK map kinase ki... 752 0.0
TC78699 weakly similar to PIR|G96761|G96761 probable MAP kinase ... 268 2e-72
BG454756 homologue to GP|12718822|db NQK1 MAPKK {Nicotiana tabac... 162 2e-40
TC86478 similar to GP|3688193|emb|CAA08995.1 MAP3K alpha 1 prote... 132 2e-31
TC88861 weakly similar to GP|17027283|gb|AAL34137.1 putative pro... 125 3e-29
TC78090 GP|15528441|emb|CAC69138. MAP kinase kinase {Medicago sa... 114 6e-26
TC85065 similar to GP|1255448|dbj|BAA09057.1 mitogen-activated p... 114 8e-26
TC86454 homologue to GP|4567091|gb|AAD23582.1| SNF-1-like serine... 112 2e-25
TC83271 similar to GP|15528441|emb|CAC69138. MAP kinase kinase {... 109 1e-24
TC81046 similar to GP|17529280|gb|AAL38867.1 putative protein ki... 108 3e-24
TC90430 similar to GP|18766642|gb|AAL79042.1 NIMA-related protei... 108 4e-24
TC76610 similar to PIR|T50802|T50802 serine/threonine protein ki... 107 1e-23
TC77429 similar to GP|15215664|gb|AAK91377.1 AT5g25110/T11H3_120... 105 2e-23
TC90610 similar to GP|18766642|gb|AAL79042.1 NIMA-related protei... 89 6e-23
TC86661 similar to GP|9798599|dbj|BAB11737.1 serine/threonine pr... 104 6e-23
TC78370 similar to GP|5001828|gb|AAD37165.1| 3-phosphoinositide-... 103 8e-23
TC77782 similar to GP|13448037|gb|AAK26845.1 SOS2-like protein k... 102 2e-22
TC87865 homologue to GP|13936371|gb|AAK40361.1 CTR1-like protein... 102 3e-22
TC79396 similar to GP|13605881|gb|AAK32926.1 At2g11520/F14P14.15... 101 5e-22
TC77566 similar to GP|310580|gb|AAA34017.1|| protein kinase 2 {G... 99 3e-21
>TC77881 homologue to GP|15528439|emb|CAC69137. MEK map kinase kinsae
{Medicago sativa subsp. x varia}, complete
Length = 1546
Score = 752 bits (1941), Expect = 0.0
Identities = 366/366 (100%), Positives = 366/366 (100%)
Frame = +2
Query: 1 MRPIQLPPPTSTGSATASGSPNNNNSRPQRRRRDHLTLPLPQRDTNLAVPLPLPPSGGSG 60
MRPIQLPPPTSTGSATASGSPNNNNSRPQRRRRDHLTLPLPQRDTNLAVPLPLPPSGGSG
Sbjct: 218 MRPIQLPPPTSTGSATASGSPNNNNSRPQRRRRDHLTLPLPQRDTNLAVPLPLPPSGGSG 397
Query: 61 GGGNGSGSGSGGASQQLVIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHE 120
GGGNGSGSGSGGASQQLVIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHE
Sbjct: 398 GGGNGSGSGSGGASQQLVIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHE 577
Query: 121 ESVRRQIHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLEGKHIPQENQLAD 180
ESVRRQIHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLEGKHIPQENQLAD
Sbjct: 578 ESVRRQIHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLEGKHIPQENQLAD 757
Query: 181 VARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTI 240
VARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTI
Sbjct: 758 VARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTI 937
Query: 241 AYMSPERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQ 300
AYMSPERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQ
Sbjct: 938 AYMSPERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQ 1117
Query: 301 PPEAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVRNGSNHNQSPPNMHQLLPPP 360
PPEAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVRNGSNHNQSPPNMHQLLPPP
Sbjct: 1118PPEAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVRNGSNHNQSPPNMHQLLPPP 1297
Query: 361 PRSQSS 366
PRSQSS
Sbjct: 1298PRSQSS 1315
Score = 28.1 bits (61), Expect = 5.6
Identities = 15/27 (55%), Positives = 15/27 (55%)
Frame = -2
Query: 36 LTLPLPQRDTNLAVPLPLPPSGGSGGG 62
L LPLP P PLPP GGSG G
Sbjct: 426 LPLPLP------FPPPPLPPDGGSGSG 364
>TC78699 weakly similar to PIR|G96761|G96761 probable MAP kinase T9L24.32
[imported] - Arabidopsis thaliana, partial (71%)
Length = 1119
Score = 268 bits (686), Expect = 2e-72
Identities = 142/318 (44%), Positives = 203/318 (63%), Gaps = 8/318 (2%)
Frame = +3
Query: 33 RDHLTLPLPQ-RDTNLAVPLPLPPSGGSGGGGNGSGSGSGGASQQLVIPFSELERLNRIG 91
R+ L LP+ D P+PLP + + + + G I + E+L+ +G
Sbjct: 54 RNSPKLRLPEISDHRPRFPVPLPANIYKQPSTSATTASVAGGDN---ISAGDFEKLSVLG 224
Query: 92 SGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDD-VNVVKCHEMYD 150
G+GGTVYKV H++ YALK+ + + + RR+ E+ ILR D NVVK H ++
Sbjct: 225 HGNGGTVYKVRHKLTSIIYALKINHYDSDPTTRRRALTEVNILRRATDCTNVVKYHGSFE 404
Query: 151 H-NAEIQVLLEYMDGGSLEGKHIP----QENQLADVARQILRGLAYLHRRHIVHRDIKPS 205
++ +L+EYMD GSLE E++L+ VAR IL GL YLH R+I HRDIKPS
Sbjct: 405 KPTGDVCILMEYMDSGSLETALKTTGTFSESKLSTVARDILNGLTYLHARNIAHRDIKPS 584
Query: 206 NLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQYDAYAGDIW 265
N+L+N + +VKIADFGV + + +T++ CNS VGT AYMSPER + ++ G Y+ ++ DIW
Sbjct: 585 NILVNIKNEVKIADFGVSKFMGRTLEACNSYVGTCAYMSPERFDPEVYGGNYNGFSADIW 764
Query: 266 SLGVSILEFYMGRFPF-AVGRQGDWASLMCAICMSQPPEAPTTASPEFRDFVSRCLQRDP 324
SLG+++ E Y+G FPF G++ DWASLMCAIC S PP P TAS EFR+FV CL+++
Sbjct: 765 SLGLTLFELYVGYFPFLQSGQRPDWASLMCAICFSDPPSLPETASSEFRNFVECCLKKES 944
Query: 325 SRRWTASRLLSHPFLVRN 342
RW+A++LL+HPFL ++
Sbjct: 945 GERWSAAQLLTHPFLCKD 998
>BG454756 homologue to GP|12718822|db NQK1 MAPKK {Nicotiana tabacum}, partial
(60%)
Length = 648
Score = 162 bits (410), Expect = 2e-40
Identities = 100/220 (45%), Positives = 133/220 (60%), Gaps = 10/220 (4%)
Frame = +2
Query: 127 IHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSL----EGKHIPQENQLADVA 182
I +E++I + +VV C+ + +N I ++LEYMD GSL + E LA V
Sbjct: 2 IVQELKINQASQCPHVVVCYHSFYNNGVISLVLEYMDRGSLVDVIRQVNTILEPYLAVVC 181
Query: 183 RQILRGLAYLHR-RHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIA 241
+Q+L+GL YLH RH++HRDIKPSNLL+N + +VKI DFGV +L TM ++ VGT
Sbjct: 182 KQVLQGLVYLHNERHVIHRDIKPSNLLVNHKGEVKITDFGVSAMLASTMGQRDTFVGTYN 361
Query: 242 YMSPERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGR-QGDWAS---LMCAIC 297
YMSPERI+ D Y+ DIWSLG+ +LE +GRFP+ Q W S L+ AI
Sbjct: 362 YMSPERISGSTYD-----YSCDIWSLGMVVLECAIGRFPYIQSEDQQAWPSFYELLQAIV 526
Query: 298 MSQPPEAPTTA-SPEFRDFVSRCLQRDPSRRWTASRLLSH 336
S PP AP SPEF FVS C+++DP R T+ LL H
Sbjct: 527 ESPPPSAPPDQFSPEFCSFVSSCIKKDPRERSTSLELLDH 646
>TC86478 similar to GP|3688193|emb|CAA08995.1 MAP3K alpha 1 protein kinase
{Brassica napus}, partial (52%)
Length = 2266
Score = 132 bits (332), Expect = 2e-31
Identities = 108/344 (31%), Positives = 175/344 (50%), Gaps = 13/344 (3%)
Frame = +2
Query: 9 PTSTGSATASGSPNNNNSRPQRRRRDHLTLPLP---QRD-TNLAVPLPLPPSGGSGGGGN 64
PTS A+ P + S P R L L P Q D + PLPLPP GS +
Sbjct: 743 PTSPLHPNATRGPTSPTS-PLHPRLQGLNLDSPTGKQDDGRSQCHPLPLPP--GSPTSPS 913
Query: 65 GSGSGSGGASQQLVIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIY----GHHE 120
+ S + + S+ ++ +G G+ G VY + NG+ A+K + +
Sbjct: 914 SALSNTRSPFENSSPNLSKWKKGKLLGRGTFGHVYLGFNSENGQMCAIKEVRVGCDDQNS 1093
Query: 121 ESVRRQIHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLEG---KHIP-QEN 176
+ +Q+H+EI +L + N+V+ + V LEY+ GGS+ ++ P +E
Sbjct: 1094 KECLKQLHQEIDLLNQLSHPNIVQYLGSELGEESLSVYLEYVSGGSIHKLLQEYGPFKEP 1273
Query: 177 QLADVARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSS 236
+ + RQI+ GLAYLH R+ VHRDIK +N+L++ ++K+ADFG+ + + S
Sbjct: 1274 VIQNYTRQIVSGLAYLHGRNTVHRDIKGANILVDPNGEIKLADFGMAKHITSAASML-SF 1450
Query: 237 VGTIAYMSPERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAI 296
G+ +M+PE + +N Y + DIWSLG +++E + P++ Q + + + I
Sbjct: 1451 KGSPYWMAPEVV---MNTNGY-SLPVDIWSLGCTLIEMAASKPPWS---QYEGVAAIFKI 1609
Query: 297 CMSQP-PEAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFL 339
S+ P P S + ++F+ CLQRDPS R TA +LL HPF+
Sbjct: 1610 GNSKDMPIIPEHLSNDAKNFIMLCLQRDPSARPTAQKLLEHPFI 1741
>TC88861 weakly similar to GP|17027283|gb|AAL34137.1 putative protein kinase
{Oryza sativa (japonica cultivar-group)}, partial (34%)
Length = 1847
Score = 125 bits (313), Expect = 3e-29
Identities = 90/263 (34%), Positives = 144/263 (54%), Gaps = 13/263 (4%)
Frame = +1
Query: 90 IGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVR----RQIHREIQILRDVDDVNVVKC 145
IG GS G+VY + G + ALK + ++ +Q+ +EI+IL + N+V+
Sbjct: 1042 IGRGSFGSVYHATNLETGASCALKEVDLVPDDPKSTDCIKQLDQEIRILGQLHHPNIVEY 1221
Query: 146 HEMYDHNAEIQVLLEYMDGGSLEG---KH--IPQENQLADVARQILRGLAYLHRRHIVHR 200
+ + + +EY+ GSL+ H + E+ + + R IL GLAYLH +HR
Sbjct: 1222 YGSEVVGDRLCIYMEYVHPGSLQKFMQDHCGVMTESVVRNFTRHILSGLAYLHSTKTIHR 1401
Query: 201 DIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDIND--GQYD 258
DIK +NLL+++ VK+ADFGV +IL + S G+ +M+PE + + +
Sbjct: 1402 DIKGANLLVDASGIVKLADFGVSKILTEKSYEL-SLKGSPYWMAPELMMAAMKNETNPTV 1578
Query: 259 AYAGDIWSLGVSILEFYMGRFPFA--VGRQGDWASLMCAICMSQPPEAPTTASPEFRDFV 316
A A DIWSLG +I+E G+ P++ G Q + L + P+ P T SPE +DF+
Sbjct: 1579 AMAVDIWSLGCTIIEMLTGKPPWSEFPGHQAMFKVL------HRSPDIPKTLSPEGQDFL 1740
Query: 317 SRCLQRDPSRRWTASRLLSHPFL 339
+C QR+P+ R +A+ LL+HPF+
Sbjct: 1741 EQCFQRNPADRPSAAVLLTHPFV 1809
>TC78090 GP|15528441|emb|CAC69138. MAP kinase kinase {Medicago sativa subsp.
x varia}, complete
Length = 1677
Score = 114 bits (285), Expect = 6e-26
Identities = 63/148 (42%), Positives = 96/148 (64%), Gaps = 6/148 (4%)
Frame = +2
Query: 79 IPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVD 138
+ ++++ + +G G+GG V V H+ + +ALK+I + EESVR+QI +E++I +
Sbjct: 413 LSLADIDIVKVVGKGNGGVVQLVQHKWTNQFFALKIIQMNIEESVRKQIAKELKINQAAQ 592
Query: 139 DVNVVKCHEMYDHNAEIQVLLEYMDGGSL-----EGKHIPQENQLADVARQILRGLAYL- 192
VV C++ + N I ++LEYMDGGS+ + K IP E L+ + +Q+L+GL YL
Sbjct: 593 CPYVVVCYQSFYDNGVISIILEYMDGGSMADLLKKVKTIP-EPYLSAICKQVLKGLIYLH 769
Query: 193 HRRHIVHRDIKPSNLLINSRKQVKIADF 220
H RHI+HRD+KPSNLLIN +VKI F
Sbjct: 770 HERHIIHRDLKPSNLLINHTGEVKIT*F 853
Score = 94.4 bits (233), Expect = 6e-20
Identities = 56/134 (41%), Positives = 78/134 (57%), Gaps = 5/134 (3%)
Frame = +3
Query: 211 SRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQYDAYAGDIWSLGVS 270
++ ++++ DFGV I+ T N+ +GT YMSPERIN + Y+ Y DIWSLG+
Sbjct: 825 TQARLRLPDFGVSAIMESTSGQANTFIGTYNYMSPERING--SQRGYN-YKSDIWSLGLI 995
Query: 271 ILEFYMGRFPFAVGRQGD-WAS---LMCAICMSQPPEAPTTA-SPEFRDFVSRCLQRDPS 325
+LE MGRFP+ Q + W S L+ I PP AP+ S EF F+S CLQ+DP
Sbjct: 996 LLECAMGRFPYTPPDQSERWESIFELIETIVDKPPPSAPSEQFSSEFCSFISACLQKDPG 1175
Query: 326 RRWTASRLLSHPFL 339
R +A L+ PF+
Sbjct: 1176SRLSAQELMELPFI 1217
>TC85065 similar to GP|1255448|dbj|BAA09057.1 mitogen-activated protein
kinase {Arabidopsis thaliana}, partial (26%)
Length = 789
Score = 114 bits (284), Expect = 8e-26
Identities = 68/178 (38%), Positives = 104/178 (58%), Gaps = 2/178 (1%)
Frame = +2
Query: 174 QENQLADVARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPC 233
+++Q++ RQIL GL YLH R+IVHRDIK +N+L+++ VK+ADFG+ + ++
Sbjct: 14 RDSQVSAYTRQILHGLKYLHDRNIVHRDIKCANILVDANGSVKVADFGLS*AIK--LNDV 187
Query: 234 NSSVGTIAYMSPERINTDINDGQYDAYA--GDIWSLGVSILEFYMGRFPFAVGRQGDWAS 291
S GT +M+PE + G+ Y DIWSLG ++LE G+ P+A + S
Sbjct: 188 KSCQGTAFWMAPEVVR-----GKVKGYGLPADIWSLGCTVLEMLTGQVPYA---PMECIS 343
Query: 292 LMCAICMSQPPEAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVRNGSNHNQS 349
M I + P P T S + RDF+ +CL+ +P R TA++LL H F+ R+ S + S
Sbjct: 344 AMFRIGKGELPPVPDTLSRDARDFILQCLKVNPDDRPTAAQLLDHKFVQRSFSQSSGS 517
>TC86454 homologue to GP|4567091|gb|AAD23582.1| SNF-1-like serine/threonine
protein kinase {Glycine max}, partial (90%)
Length = 2194
Score = 112 bits (280), Expect = 2e-25
Identities = 85/293 (29%), Positives = 139/293 (47%), Gaps = 10/293 (3%)
Frame = +3
Query: 69 GSGGASQQLVIPFSELERLNR-IGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQ- 126
GS G V F ++ + +G GS G V H + G A+K++ +++ +
Sbjct: 66 GSAGPGGGNVNAFLRNYKMGKTLGIGSFGKVKIAEHVLTGHKVAIKILNRRKIKNMEMEE 245
Query: 127 -IHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLEGKHIP----QENQLADV 181
+ REI+ILR ++++ +E+ + +I V++EY+ G L + QE++
Sbjct: 246 KVRREIKILRLFMHHHIIRLYEVVETPTDIYVVMEYVKSGELFDYIVEKGRLQEDEARSF 425
Query: 182 ARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIA 241
+QI+ G+ Y HR +VHRD+KP NLL++S+ VKIADFG+ I+ +S G+
Sbjct: 426 FQQIISGVEYCHRNMVVHRDLKPENLLLDSKWSVKIADFGLSNIMRDG-HFLKTSCGSPN 602
Query: 242 YMSPERINTDINDGQYDAYAG---DIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICM 298
Y +PE I+ + YAG D+WS GV + G PF + +
Sbjct: 603 YAAPEVISGKL-------YAGPEVDVWSCGVILYALLCGTLPF----DDENIPNLFKKIK 749
Query: 299 SQPPEAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVRNGSNHNQSPP 351
P+ SP RD + R L DP +R T + HP+ + + PP
Sbjct: 750 GGIYTLPSHLSPGARDLIPRLLVVDPMKRMTIPEIRQHPWFQLHLPRYLAVPP 908
>TC83271 similar to GP|15528441|emb|CAC69138. MAP kinase kinase {Medicago
sativa subsp. x varia}, partial (42%)
Length = 683
Score = 109 bits (273), Expect = 1e-24
Identities = 62/142 (43%), Positives = 82/142 (57%), Gaps = 5/142 (3%)
Frame = +1
Query: 203 KPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQYDAYAG 262
KPSN+LIN R +VKI DFGV I+ T N+ +GT YMSPERI ++ Y+ Y
Sbjct: 10 KPSNILINHRGEVKITDFGVSAIMESTSGQANTFIGTYNYMSPERIKASDSEQGYN-YKS 186
Query: 263 DIWSLGVSILEFYMGRFPFAVGRQGD-WASLMC---AICMSQPPEA-PTTASPEFRDFVS 317
DIWSLG+ +L+ G+FP+ + W +L AI P A P SPEF F+S
Sbjct: 187 DIWSLGLMLLKCATGKFPYTPPDNSEGWENLFLLIEAIVEKPSPSAPPDECSPEFCSFIS 366
Query: 318 RCLQRDPSRRWTASRLLSHPFL 339
CLQ++P R + LL HPF+
Sbjct: 367 ACLQKNPRDRPSTRNLLRHPFV 432
>TC81046 similar to GP|17529280|gb|AAL38867.1 putative protein kinase
{Arabidopsis thaliana}, partial (44%)
Length = 921
Score = 108 bits (270), Expect = 3e-24
Identities = 71/225 (31%), Positives = 118/225 (51%), Gaps = 13/225 (5%)
Frame = +2
Query: 127 IHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGS---LEGKHIPQ---ENQLAD 180
I RE+Q + +D N+++ H + + V++ +M GGS + P+ E +A
Sbjct: 23 IRREVQTMSLIDHPNLLRAHCSFTAGHSLWVVMPFMSGGSCLHIMKSSFPEGFDEPVIAT 202
Query: 181 VARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMD---PCNSSV 237
V R++L+ L YLH +HRD+K N+L+++ VK+ADFGV + T D N+ V
Sbjct: 203 VLREVLKALVYLHAHGHIHRDVKAGNILLDANGSVKMADFGVSACMFDTGDRQRSRNTFV 382
Query: 238 GTIAYMSPERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAIC 297
GT +M+PE + ++ YD + DIWS G++ LE G PF+ + ++
Sbjct: 383 GTPCWMAPE-VMQQLHG--YD-FKADIWSFGITALELAHGHAPFS---KYPPMKVLLMTL 541
Query: 298 MSQPP----EAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPF 338
+ PP E S F++ V+ CL +DP +R ++ +LL H F
Sbjct: 542 QNAPPGLDYERDKRFSKSFKELVATCLVKDPKKRPSSEKLLKHHF 676
>TC90430 similar to GP|18766642|gb|AAL79042.1 NIMA-related protein kinase
{Populus x canescens}, partial (45%)
Length = 976
Score = 108 bits (269), Expect = 4e-24
Identities = 78/248 (31%), Positives = 124/248 (49%), Gaps = 9/248 (3%)
Frame = +1
Query: 101 VVHRINGRAYALKVI-YGHHEESVRRQIHREIQILRDVDDVNVVKCHEMY-DHNAEIQVL 158
V H+ + Y LK I + RR H+E++++ V + +V+ + + + + ++
Sbjct: 136 VRHKHEKKKYVLKKIRLARQTDRTRRSAHQEMELISKVRNPFIVEYKDSWVEKGCFVCII 315
Query: 159 LEYMDGGSL-------EGKHIPQENQLADVARQILRGLAYLHRRHIVHRDIKPSNLLINS 211
+ Y +GG + G + P+E +L Q+L L YLH HI+HRD+K SN+ +
Sbjct: 316 IGYCEGGDMAETVKKANGVNFPEE-KLCKWLVQLLMALDYLHVNHILHRDVKCSNIFLTK 492
Query: 212 RKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQYDAYAGDIWSLGVSI 271
+ +++ DFG+ ++L D +S VGT +YM PE + DI G DIWSLG +
Sbjct: 493 NQDIRLGDFGLAKLLTSD-DLASSIVGTPSYMCPELL-ADIPYGS----KSDIWSLGCCV 654
Query: 272 LEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPEFRDFVSRCLQRDPSRRWTAS 331
E R F + D +L+ I S PT S FR V L+++P R TA
Sbjct: 655 YEMAAHRPAF---KAFDIQALIHKINKSIVSPLPTMYSSAFRGLVKSMLRKNPELRPTAG 825
Query: 332 RLLSHPFL 339
LL+HP L
Sbjct: 826 ELLNHPHL 849
>TC76610 similar to PIR|T50802|T50802 serine/threonine protein kinase-like
protein - Arabidopsis thaliana, partial (73%)
Length = 1799
Score = 107 bits (266), Expect = 1e-23
Identities = 75/271 (27%), Positives = 131/271 (47%), Gaps = 11/271 (4%)
Frame = +2
Query: 79 IPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHH--EESVRRQIHREIQILRD 136
I F++ E +G G+ VY + A+KVI +E + +QI RE+ ++R
Sbjct: 311 ILFNKYEIGKTLGQGNFAKVYHGRNIATNENVAIKVIKKEKLKKERLMKQIKREVSVMRL 490
Query: 137 VDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLEGKHIP---QENQLADVARQILRGLAYLH 193
V ++V+ E+ + A++ +++EY+ GG L K +EN +Q++ + + H
Sbjct: 491 VRHPHIVELKEVMANKAKVFMVVEYVKGGELFAKVAKGKMEENIARKYFQQLISAVDFCH 670
Query: 194 RRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQ------TMDPCNSSVGTIAYMSPER 247
R + HRD+KP NLL++ + +K++DFG+ + +Q + PC GT AY++PE
Sbjct: 671 SRGVTHRDLKPENLLLDENEDLKVSDFGLSALPDQRRSDGMLLTPC----GTPAYVAPEV 838
Query: 248 INTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTT 307
+ YD DIWS GV + G PF QG+ + + P
Sbjct: 839 LKKI----GYDGSKADIWSCGVILFALLCGYLPF----QGENVMRIYSKSFKADYVLPEW 994
Query: 308 ASPEFRDFVSRCLQRDPSRRWTASRLLSHPF 338
SP + +S L DP +R++ ++ P+
Sbjct: 995 ISPGAKKLISNLLVVDPEKRFSIPDIMKDPW 1087
>TC77429 similar to GP|15215664|gb|AAK91377.1 AT5g25110/T11H3_120
{Arabidopsis thaliana}, partial (79%)
Length = 1869
Score = 105 bits (263), Expect = 2e-23
Identities = 76/267 (28%), Positives = 123/267 (45%), Gaps = 9/267 (3%)
Frame = +3
Query: 81 FSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHH--EESVRRQIHREIQILRDVD 138
F + E +G G+ VY G A+KV+ +E + QI REI ++R V
Sbjct: 261 FGKYEMGRVLGKGTFAKVYYAKEISTGEGVAIKVVDKDKVKKEGMMEQIKREISVMRLVK 440
Query: 139 DVNVVKCHEMYDHNAEIQVLLEYMDGGSL-----EGKHIPQENQLADVARQILRGLAYLH 193
N+V E+ +I ++EY GG L +GK +E +Q++ + Y H
Sbjct: 441 HPNIVNLKEVMATKTKILFVMEYARGGELFEKVAKGKF--KEELARKYFQQLISAVDYCH 614
Query: 194 RRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDP--CNSSVGTIAYMSPERINTD 251
R + HRD+KP NLL++ + +K++DFG+ + ++ GT AY++PE
Sbjct: 615 SRGVSHRDLKPENLLLDENENLKVSDFGLSALPEHLRQDGLLHTQCGTPAYVAPE----V 782
Query: 252 INDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPE 311
+ Y + D WS GV + G PF Q + M + + P SPE
Sbjct: 783 VRKRGYSGFKADTWSCGVILYALLAGFLPF----QHENLISMYNKVFKEEYQFPPWFSPE 950
Query: 312 FRDFVSRCLQRDPSRRWTASRLLSHPF 338
+ +S+ L DP RR T S +++ P+
Sbjct: 951 SKRLISKILVADPERRITISSIMNVPW 1031
>TC90610 similar to GP|18766642|gb|AAL79042.1 NIMA-related protein kinase
{Populus x canescens}, partial (46%)
Length = 1507
Score = 89.0 bits (219), Expect(2) = 6e-23
Identities = 59/165 (35%), Positives = 86/165 (51%)
Frame = +2
Query: 175 ENQLADVARQILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCN 234
E +L Q+L L YLH HI+HRD+K SN+ + + +++ DFG+ ++L D +
Sbjct: 566 EERLCKWLVQLLMALDYLHANHILHRDVKCSNIFLTKDQDIRLGDFGLAKMLTSD-DLAS 742
Query: 235 SSVGTIAYMSPERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMC 294
S VGT +YM PE + DI G DIWSLG + E + F + D +L+
Sbjct: 743 SIVGTPSYMCPELL-ADIPYGS----KSDIWSLGCCVYEMAAHKPAF---KALDMQALIN 898
Query: 295 AICMSQPPEAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFL 339
I S PT S FR V L+++P R +A+ LL+HP L
Sbjct: 899 KINKSLVAPLPTMYSGTFRSMVKSMLRKNPELRPSAADLLNHPHL 1033
Score = 36.2 bits (82), Expect(2) = 6e-23
Identities = 22/87 (25%), Positives = 40/87 (45%), Gaps = 10/87 (11%)
Frame = +3
Query: 75 QQLVIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVI-YGHHEESVRRQIHREIQI 133
+ L + + E L +IG GS + V H+ + Y LK I + +RR H+E+++
Sbjct: 240 RNLELDMEQYEVLEQIGKGSFASALLVRHKHENKRYVLKKIRLARQSDRIRRSAHQEMEL 419
Query: 134 LRDVDDVNVVK---------CHEMYDH 151
+ V + +V+ C +Y H
Sbjct: 420 ISKVRNPFIVEYKDSWVEKGCFVLYSH 500
>TC86661 similar to GP|9798599|dbj|BAB11737.1 serine/threonine protein
kinase {Arabidopsis thaliana}, partial (58%)
Length = 1840
Score = 104 bits (259), Expect = 6e-23
Identities = 73/266 (27%), Positives = 124/266 (46%), Gaps = 8/266 (3%)
Frame = +2
Query: 81 FSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEES--VRRQIHREIQILRDVD 138
F + E +G G+ VY + NG++ A+KVI + + + REI I+ +
Sbjct: 296 FGKYELGKLLGCGAFAKVYHARNTENGQSVAVKVINKKKISATGLAGHVKREISIMSKLR 475
Query: 139 DVNVVKCHEMYDHNAEIQVLLEYMDGGSLEGKHIPQENQLADVAR----QILRGLAYLHR 194
N+V+ HE+ +I ++E+ GG L K + D++R Q++ + Y H
Sbjct: 476 HPNIVRLHEVLATKTKIYFVMEFAKGGELFAKIANKGRFSEDLSRRLFQQLISAVGYCHS 655
Query: 195 RHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTM--DPCNSSVGTIAYMSPERINTDI 252
+ HRD+KP NLL++ + +K++DFG+ + Q ++ GT AY++PE +
Sbjct: 656 HGVFHRDLKPENLLLDDKGNLKVSDFGLSAVKEQVRIDGMLHTLCGTPAYVAPE----IL 823
Query: 253 NDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPEF 312
YD DIWS G+ + G PF +M S + P SP+
Sbjct: 824 AKRGYDGAKVDIWSCGIILFVLVAGYLPF----NDPNLMVMYRKIYSGEFKCPRWFSPDL 991
Query: 313 RDFVSRCLQRDPSRRWTASRLLSHPF 338
+ F+SR + +P R T +L P+
Sbjct: 992 KRFLSRLMDTNPETRITVDEILRDPW 1069
>TC78370 similar to GP|5001828|gb|AAD37165.1| 3-phosphoinositide-dependent
protein kinase-1 {Arabidopsis thaliana}, partial (91%)
Length = 2137
Score = 103 bits (258), Expect = 8e-23
Identities = 88/326 (26%), Positives = 143/326 (42%), Gaps = 28/326 (8%)
Frame = +1
Query: 56 SGGSGGGGNGSGSGSGGASQQLVIP-----FSELERLNRIGSGSGGTVYKVVHRINGRAY 110
S S GGG G+G+ S P + E G GS V + + G Y
Sbjct: 376 SSSSNGGGGGAGNVQRSKSFAFRAPHENYTIQDFELGKIYGVGSYSKVVRAKKKDTGVVY 555
Query: 111 ALKVIYGHH--EESVRRQIHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLE 168
A+K++ +E+ + E +L +D +V+ + + + + LE +GG L
Sbjct: 556 AMKIMDKKFITKENKTAYVKLERIVLDQLDHPGIVRLYFTFQDTFSLYMALESCEGGELF 735
Query: 169 GKHIPQENQLADVAR----QILRGLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGR 224
+ + D A+ +++ L Y+H ++HRDIKP NLL+ + +KIADFG +
Sbjct: 736 DQITRKSRLTQDEAQFYAAEVVDALEYIHSLGVIHRDIKPENLLLTTEGHIKIADFGSVK 915
Query: 225 ILNQTM----------DPCNSSVGTIAYMSPERINTDINDGQYDAYAGDIWSLGVSILEF 274
+ + D + VGT AY+ PE +N+ + D+W+LG ++ +
Sbjct: 916 PMQDSQITVLPNAASDDKACTFVGTAAYVPPEVLNS-----SPATFGNDLWALGCTLYQM 1080
Query: 275 YMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPEFRDFVSRCLQRDPSRRWTA---- 330
G PF +W L+ +++ P S E RD + R L DPSRR A
Sbjct: 1081LSGTSPFK--DASEW--LIFQRIIAREIRFPDYFSDEARDLIDRLLDLDPSRRPGAGPDG 1248
Query: 331 -SRLLSHPFL--VRNGSNHNQSPPNM 353
+ L SHPF V + Q+PP +
Sbjct: 1249YAILKSHPFFKGVDWNNLRAQTPPKL 1326
>TC77782 similar to GP|13448037|gb|AAK26845.1 SOS2-like protein kinase PKS6
{Arabidopsis thaliana}, partial (77%)
Length = 1236
Score = 102 bits (255), Expect = 2e-22
Identities = 73/250 (29%), Positives = 120/250 (47%), Gaps = 12/250 (4%)
Frame = +2
Query: 90 IGSGSGGTVYKVVHRINGRAYALKVIYGHH--EESVRRQIHREIQILRDVDDVNVVKCHE 147
IG GS V + NG A+K++ +H + ++ Q+ REI ++ ++ NV+K E
Sbjct: 209 IGEGSFAKVKLAKNVENGDYVAIKILDRNHVLKHNMMDQLKREISAMKIINHPNVIKIFE 388
Query: 148 MYDHNAEIQVLLEYMDGGSLEGKHIPQENQLADVAR----QILRGLAYLHRRHIVHRDIK 203
+ +I ++LE ++GG L K D AR Q++ + Y H R + HRD+K
Sbjct: 389 VMASKTKIYIVLELVNGGELFDKIATHGRLKEDEARSYFQQLINAVDYCHSRGVYHRDLK 568
Query: 204 PSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQYDAYAGD 263
P NLL+++ +K++DFG+ Q + ++ GT Y++PE +ND Y + D
Sbjct: 569 PENLLLDTNGVLKVSDFGLSTYSQQEDELLRTACGTPNYVAPE----VLNDRGYVGSSSD 736
Query: 264 IWSLGVSILEFYMGRFPF------AVGRQGDWASLMCAICMSQPPEAPTTASPEFRDFVS 317
IWS GV + G PF A+ R+ A C P+ SPE + ++
Sbjct: 737 IWSCGVILFVLMAGYLPFDEPNLIALYRKIGRADFKC----------PSWFSPEAKKLLT 886
Query: 318 RCLQRDPSRR 327
L +P R
Sbjct: 887 SILNPNPLTR 916
>TC87865 homologue to GP|13936371|gb|AAK40361.1 CTR1-like protein kinase
{Rosa hybrid cultivar}, partial (31%)
Length = 1190
Score = 102 bits (253), Expect = 3e-22
Identities = 73/264 (27%), Positives = 130/264 (48%), Gaps = 11/264 (4%)
Frame = +3
Query: 75 QQLVIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVR-RQIHREIQI 133
+ L IP+S+L +IGSGS GTV++ NG A+K++ + R ++ RE+ I
Sbjct: 30 EDLAIPWSDLVLKEKIGSGSFGTVHRA--EWNGSDVAVKILMEQDFHAERFKEFMREVAI 203
Query: 134 LRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLE-------GKHIPQENQLADVARQIL 186
++ + N+V + ++ EY+ GSL K + E + +A +
Sbjct: 204 MKHLRHPNIVLLMGAVTQPPNLSIVTEYLSRGSLYRLLHRPGAKEVLDERRRLSMAYDVA 383
Query: 187 RGLAYLHRRH--IVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMS 244
+G+ YLH+R+ IVHRD+K NLL++ + VK+ DFG+ R+ T S+ GT +M+
Sbjct: 384 KGMNYLHKRNPPIVHRDLKSPNLLVDKKYTVKVCDFGLSRLKANTFLSSKSAAGTPEWMA 563
Query: 245 PERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPP-E 303
PE + + ++ + DI+S GV + E + P+ + A ++ A+ E
Sbjct: 564 PEVLRDEPSNEK-----SDIYSFGVIMWEIATLQQPWG---NLNPAQVVAAVGFKHKRLE 719
Query: 304 APTTASPEFRDFVSRCLQRDPSRR 327
P +P+ + C +P +R
Sbjct: 720 IPRNLNPQIAAIIEACWANEPWKR 791
>TC79396 similar to GP|13605881|gb|AAK32926.1 At2g11520/F14P14.15
{Arabidopsis thaliana}, partial (53%)
Length = 1658
Score = 101 bits (251), Expect = 5e-22
Identities = 79/261 (30%), Positives = 121/261 (46%), Gaps = 20/261 (7%)
Frame = +1
Query: 48 AVPLPLPPSGGSGGGGNGSGSGSGGASQQLVIPFSELERLNR-------IGSGSGGTVYK 100
A PL +PPS S S Q L + S++ + R IG G GTVYK
Sbjct: 217 ASPLRVPPSPS-----RFSMSPKLSRLQSLHLNLSQVSKATRNFSETLQIGEGGFGTVYK 381
Query: 101 VVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLE 160
H +G A+K H ES+R + E+++L +D N+VK D E ++ E
Sbjct: 382 A-HLDDGLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGYIDKGNERILITE 558
Query: 161 YMDGGSLE-------GKHIPQENQLADVARQILRGLAYLH---RRHIVHRDIKPSNLLIN 210
++ G+L GK I NQ ++A + GL YLH + I+HRD+K SN+L+
Sbjct: 559 FVANGTLREHLDGLRGK-ILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLT 735
Query: 211 SRKQVKIADFG---VGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQYDAYAGDIWSL 267
+ K+ADFG +G + N GT+ Y+ PE + T + D++S
Sbjct: 736 ESMRAKVADFGFAKLGPVNNDHTHISTKVKGTVGYLDPEYMKT-----YHLTPKSDVYSF 900
Query: 268 GVSILEFYMGRFPFAVGRQGD 288
G+ +LE GR P + + +
Sbjct: 901 GILLLEILTGRRPVELKKSAE 963
>TC77566 similar to GP|310580|gb|AAA34017.1|| protein kinase 2 {Glycine
max}, partial (88%)
Length = 1502
Score = 98.6 bits (244), Expect = 3e-21
Identities = 77/275 (28%), Positives = 126/275 (45%), Gaps = 17/275 (6%)
Frame = +1
Query: 85 ERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNVVK 144
E L +GSG+ G + G ALK I + + + REI R + N+++
Sbjct: 79 EPLKELGSGNFGVARLAKDKNTGELVALKYI--ERGKKIDENVQREIINHRSLRHPNIIR 252
Query: 145 CHEMYDHNAEIQVLLEYMDGGSLEGKHIPQENQLADVAR----QILRGLAYLHRRHIVHR 200
E+ + + ++LEY GG L + D AR Q++ G++Y H I HR
Sbjct: 253 FKEVLLTPSHLAIVLEYASGGELFERICSAGRFSEDEARYFFQQLISGVSYCHSMEICHR 432
Query: 201 DIKPSNLLI--NSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQYD 258
D+K N L+ N ++KI DFG + P S+VGT AY++PE ++ +YD
Sbjct: 433 DLKLENTLLDGNPSPRLKICDFGYSKSAILHSQP-KSTVGTPAYIAPEVLSRK----EYD 597
Query: 259 AYAGDIWSLGVSILEFYMGRFPF-----------AVGRQGDWASLMCAICMSQPPEAPTT 307
D+WS GV++ +G +PF +GR + + S P +
Sbjct: 598 GKIADVWSCGVTLYVMLVGAYPFEEPEDPRNFRKTIGR-------IIGVQYSIPDYVRIS 756
Query: 308 ASPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVRN 342
A E R+ +SR DP++R T + +P+ ++N
Sbjct: 757 A--ECRNLLSRIFVADPAKRVTIPEIKQNPWFLKN 855
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.136 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,605,346
Number of Sequences: 36976
Number of extensions: 211008
Number of successful extensions: 3667
Number of sequences better than 10.0: 623
Number of HSP's better than 10.0 without gapping: 2749
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3240
length of query: 366
length of database: 9,014,727
effective HSP length: 97
effective length of query: 269
effective length of database: 5,428,055
effective search space: 1460146795
effective search space used: 1460146795
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144503.4


