

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
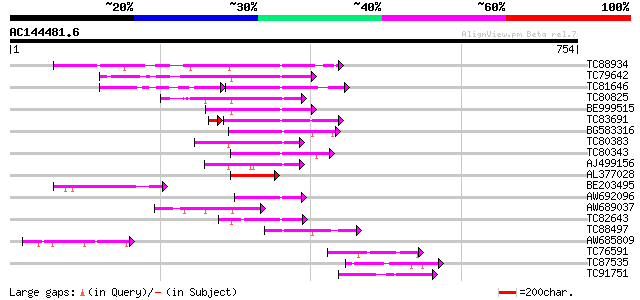
Query= AC144481.6 - phase: 0
(754 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC88934 similar to PIR|T00434|T00434 probable kinesin heavy chai... 121 9e-28
TC79642 similar to PIR|T07397|T07397 kinesin heavy chain-like pr... 110 2e-24
TC81646 similar to GP|13536981|dbj|BAB40709. BY-2 kinesin-like p... 69 7e-21
TC80825 similar to PIR|F84614|F84614 probable kinesin heavy chai... 86 4e-17
BE999515 similar to PIR|T07397|T0 kinesin heavy chain-like prote... 86 5e-17
TC83691 similar to PIR|T00434|T00434 probable kinesin heavy chai... 78 7e-17
BG583316 homologue to GP|11994617|db contains similarity to kine... 79 6e-15
TC80383 similar to PIR|C85065|C85065 kinesin-like protein [impor... 76 5e-14
TC80343 similar to PIR|C96661|C96661 kinesin-like protein 73641... 69 9e-12
AJ499156 similar to GP|10176794|db kinesin-like protein {Arabido... 63 4e-10
AL377028 SP|O23826|K12 125 kDa kinesin-related protein. [Common ... 59 5e-09
BE203495 weakly similar to PIR|T48258|T48 kinesin-like protein -... 55 1e-07
AW692096 homologue to GP|10140692|gb kinesin-like protein {Oryza... 53 4e-07
AW689037 similar to GP|10177316|d kinesin-like protein {Arabidop... 53 5e-07
TC82643 similar to GP|10998143|dbj|BAB03114. kinesin (centromere... 52 8e-07
TC88497 similar to GP|11994617|dbj|BAB02754. contains similarity... 51 1e-06
AW685809 similar to PIR|E84792|E8 probable kinesin heavy chain [... 50 3e-06
TC76591 similar to PIR|B85063|B85063 hypothetical protein AT4g05... 45 8e-05
TC87535 similar to GP|19310542|gb|AAL85004.1 unknown protein {Ar... 43 4e-04
TC91751 similar to GP|2739382|gb|AAC14505.1| unknown protein {Ar... 43 5e-04
>TC88934 similar to PIR|T00434|T00434 probable kinesin heavy chain
[imported] - Arabidopsis thaliana, partial (49%)
Length = 2193
Score = 121 bits (304), Expect = 9e-28
Identities = 116/406 (28%), Positives = 181/406 (44%), Gaps = 20/406 (4%)
Frame = +3
Query: 59 IEVIARIRDY-PDRKDKPLSVLQASSNSRSIRV--KADFGYRDFTLDGVSVSEEEELDLF 115
I V R+R + P + + +V +I V K G R F + V + ++F
Sbjct: 87 IRVYCRVRPFLPGQPNHSSTVENIEDGVITINVPSKNGKGRRSFNFNKVFGPSAAQGEVF 266
Query: 116 YKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFG----CSKQAGIVYRALRDILGDGD 171
++ + V G I YG TGSGK+ TM G K G+ YRAL D+ +
Sbjct: 267 AD--MQPLVRSVLDGFNVCIFAYGQTGSGKTFTMTGPKEITEKSQGVNYRALSDLFYTAN 440
Query: 172 TDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNAS 231
+ D V V ++EIYNE++ DLL T+G +
Sbjct: 441 QRKDTFRYD------------VSVQMIEIYNEQVRDLLVTDG-----------------T 533
Query: 232 KVKLEVM-----GKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVI 286
+LE+ G +A+ + + + + + K R V +T NDRSSRSH +
Sbjct: 534 NKRLEIRSNSQRGLSVPDASLVQVSSTNDVIELMNLGHKNRAVGATALNDRSSRSHSCLT 713
Query: 287 LDVP--------TVGGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESI 338
+ V + G + LVD+AGSE ++++ TG K + IN+ AL V+ S+
Sbjct: 714 VHVQGRDLTSGAVLRGCMHLVDLAGSERVDKSEATGDRLK-EAQHINKSLSALGDVIASL 890
Query: 339 ANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVR 398
A + HVP+R+SKLT LLQDS ++K LM + SP+ + +TISTL++ + +
Sbjct: 891 AQKNQHVPYRNSKLTQLLQDSL-GGQAKTLMFVHISPEANAVGETISTLKFAERVATVEL 1067
Query: 399 GPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAH 444
G K+ G+ + + E I L+ L KE N H
Sbjct: 1068GAARVNKD---------GADVKELKEQIASLKA--ALARKEGNLEH 1172
>TC79642 similar to PIR|T07397|T07397 kinesin heavy chain-like protein (clone
PKCBP) - potato, partial (42%)
Length = 2321
Score = 110 bits (276), Expect = 2e-24
Identities = 83/298 (27%), Positives = 146/298 (48%), Gaps = 9/298 (3%)
Frame = +3
Query: 120 VESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQAGIVYRALRDILGDGDTDSEGSDG 179
V+S ++G + I YG TGSGK+ T++G G+ RA+ ++ DS
Sbjct: 1158 VQSAVDGYNV----CIFAYGQTGSGKTFTIYGSEDNPGLTPRAIAELFRILRRDS----- 1310
Query: 180 DSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMG 239
+ F L + ++E+Y + + DLL +K + +K + G
Sbjct: 1311 -NKYSFSL------KAYMVELYQDTLIDLLLPKN------------AKHSRLDIKKDSTG 1433
Query: 240 KKA-KNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVPTVG----- 293
+N T +S + +++ IQK +RR + T N+ SSRSH ++ + V +
Sbjct: 1434 MVVVENVTVMSISTIEELNYIIQKGSERRHISGTQMNEESSRSHLILSIVVESTNLQSQS 1613
Query: 294 ---GRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDS 350
G+L VD+AGSE ++++G G + K + IN+ AL V+ ++++G H P+R+
Sbjct: 1614 VARGKLSFVDLAGSERVKKSGSMGSQLK-EAQSINKSLSALGDVISALSSGGQHTPYRNH 1790
Query: 351 KLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKEED 408
KLTML+ DS +K LM + SP + +T ++L Y ++ + IV P V ++
Sbjct: 1791 KLTMLMSDSL-GGNAKTLMFVNVSPIESSLDETHNSLMYASRVRSIVNDPSKNVSSKE 1961
>TC81646 similar to GP|13536981|dbj|BAB40709. BY-2 kinesin-like protein 5
{Nicotiana tabacum}, partial (50%)
Length = 1456
Score = 68.9 bits (167), Expect(2) = 7e-21
Identities = 49/164 (29%), Positives = 81/164 (48%)
Frame = +3
Query: 288 DVPTVGGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPF 347
+ T +L +VD+ GSE + + G G A IN AL V+ ++ SHVP+
Sbjct: 495 EAKTETSKLWMVDLGGSERLLKTGARGLTLDEGRA-INLSLSALADVLAALKRKRSHVPY 671
Query: 348 RDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKEE 407
R+SKLT LL+DS D SK+LM++ SP +++ +TI +L + +A+ I P++ +
Sbjct: 672 RNSKLTQLLRDSL-GDGSKVLMLVHISPSEEDVCETICSLNFAKRARAIESIKEVPMESK 848
Query: 408 DSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAHKKLMKKE 451
+ IM+L+ K EK+R ++ E
Sbjct: 849 KQK------------EMKIMELEENIKEAEKQRQSLGDQIQNVE 944
Score = 50.4 bits (119), Expect(2) = 7e-21
Identities = 44/168 (26%), Positives = 70/168 (41%)
Frame = +1
Query: 120 VESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQAGIVYRALRDILGDGDTDSEGSDG 179
VE + G + YG TG+GK+ TM G ++Q GI+ RAL ++ DG
Sbjct: 37 VEPILRSAMDGHNVCVFAYGQTGTGKTFTMDGTNEQPGIIPRALEELFRQA-----SMDG 201
Query: 180 DSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMG 239
SS F +++LE+Y + DLL+ + + K +E+ G
Sbjct: 202 SSSFTF--------SMSMLEVYMGNVRDLLANS-----------NLNIQTDPKGLVEIEG 324
Query: 240 KKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVIL 287
+ + ++ K K + R T N+ SSRSHC + L
Sbjct: 325 -----LSEVQISDCAKAKWWYNKGRRCRSTSWTNVNEASSRSHCFIYL 453
>TC80825 similar to PIR|F84614|F84614 probable kinesin heavy chain
[imported] - Arabidopsis thaliana, partial (16%)
Length = 1771
Score = 86.3 bits (212), Expect = 4e-17
Identities = 68/208 (32%), Positives = 103/208 (48%), Gaps = 14/208 (6%)
Frame = +3
Query: 201 YNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISK-- 258
YNE+I DLL+T ASK +LE+ + + ++ G K+
Sbjct: 60 YNEQIRDLLATGP----------------ASK-RLEIK-QNYEGHHHVPGVVEAKVDNIS 185
Query: 259 ----EIQKVEKRRIVKSTLCNDRSSRSHCMVILDVPTVG--------GRLMLVDMAGSEN 306
+Q R V S N+ SSRSHCM+ + V T +L LVD+AGSE
Sbjct: 186 DVWTVLQAGSNARAVGSNNVNEHSSRSHCMLCIMVKTKNLMNGECTKSKLWLVDLAGSER 365
Query: 307 IEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSK 366
+ + G K + IN+ AL V+ ++A SH+P+R+SKLT LLQDS D SK
Sbjct: 366 LAKTDVQGERLK-EAQNINRSLSALGDVISALAAKSSHIPYRNSKLTHLLQDSLGGD-SK 539
Query: 367 ILMILCASPDPKEIHKTISTLEYGAKAK 394
LM + SP +++ +T+S+L + + +
Sbjct: 540 TLMFVQISPSDQDVGETLSSLNFATRVR 623
Score = 35.0 bits (79), Expect = 0.11
Identities = 33/126 (26%), Positives = 62/126 (49%), Gaps = 8/126 (6%)
Frame = +2
Query: 436 REKERNEAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQR 495
+ K ++ HK L +E+I L ++E + + + +V++ L ++L+ K E C
Sbjct: 773 KAKGKDNIHKNL---QEKIKELEGQIELKTSMQNQSEKQVSQ----LCEKLKGKEETCCT 931
Query: 496 MTNEFVELERK---RMEERILQQQEEVEILRKRLEEIELQLCSSSKQERKD-----ENES 547
+ ++ ELERK +++ Q++ L K+L++ +LQ S KD E +
Sbjct: 932 LQHKVKELERKIKEQLQTETANFQQKAWDLEKKLKD-QLQGSESESSFLKDKIKELERKL 1108
Query: 548 KEMEPN 553
KE E N
Sbjct: 1109KEQEQN 1126
>BE999515 similar to PIR|T07397|T0 kinesin heavy chain-like protein (clone
PKCBP) - potato, partial (13%)
Length = 526
Score = 85.9 bits (211), Expect = 5e-17
Identities = 50/156 (32%), Positives = 87/156 (55%), Gaps = 8/156 (5%)
Frame = +2
Query: 261 QKVEKRRIVKSTLCNDRSSRSHCMVILDVPTVG--------GRLMLVDMAGSENIEQAGQ 312
Q+ +RR T N+ SSRSH ++ + + +V G+L VD+AGSE I+++G
Sbjct: 2 QRGSERRHTAGTQMNEESSRSHRILSIVIESVNLQSQSTARGKLSFVDLAGSERIKKSGS 181
Query: 313 TGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILC 372
G + K + IN+ AL V+ ++++G H+P+R+ KLTML+ DS +K LM +
Sbjct: 182 EGSQLK-EAQSINKSLSALGDVISALSSGGQHIPYRNHKLTMLMSDSL-GGNAKTLMFVN 355
Query: 373 ASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKEED 408
SP + +T ++L Y ++ + IV P + ++
Sbjct: 356 VSPVESSLDETHNSLMYASRVRSIVNDPSKNISSKE 463
>TC83691 similar to PIR|T00434|T00434 probable kinesin heavy chain
[imported] - Arabidopsis thaliana, partial (22%)
Length = 1056
Score = 78.2 bits (191), Expect(2) = 7e-17
Identities = 51/160 (31%), Positives = 88/160 (54%)
Frame = +1
Query: 285 VILDVPTVGGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSH 344
V+L + G + LVD+AGSE ++ TG K + IN+ AL V+ S+A+ ++H
Sbjct: 154 VVLTGSIIRGCMHLVDLAGSERADKTEATGDRLK-EAQHINKSLSALGDVIASLAHKNAH 330
Query: 345 VPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPV 404
VP+R+SKLT LLQD+ ++K LM + SP+P + +T+STL++ + + G T
Sbjct: 331 VPYRNSKLTQLLQDAL-GGQAKTLMFVHISPEPDALGETLSTLKFAERVSTVELG--TAR 501
Query: 405 KEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAH 444
+D++ L +IA + + + E + ++ N H
Sbjct: 502 VNKDNTEVKELKEQIAMLKAALARKDGEAEHIQQPSNSGH 621
Score = 27.7 bits (60), Expect(2) = 7e-17
Identities = 13/18 (72%), Positives = 13/18 (72%)
Frame = +3
Query: 265 KRRIVKSTLCNDRSSRSH 282
K R V ST NDRSSRSH
Sbjct: 111 KNRAVGSTAMNDRSSRSH 164
>BG583316 homologue to GP|11994617|db contains similarity to kinesin
protein~gene_id:MGL6.9 {Arabidopsis thaliana}, partial
(15%)
Length = 725
Score = 79.0 bits (193), Expect = 6e-15
Identities = 51/162 (31%), Positives = 82/162 (50%), Gaps = 13/162 (8%)
Frame = +1
Query: 292 VGGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSK 351
V G++ +D+AGSE + +++ A+IN+ +ALK + ++ N H+PFR SK
Sbjct: 28 VVGKISFIDLAGSERGADTTDNDRQTRIEGAEINKSLLALKECIRALDNDQIHIPFRGSK 207
Query: 352 LTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPH---------T 402
LT +L+DSF + SK +MI C SP T++TL Y + K + + +
Sbjct: 208 LTEVLRDSFVGN-SKTVMISCISPGAGSCEHTLNTLRYADRVKSLSKSGNPRKDAIPNCV 384
Query: 403 PVKEEDSSSTVILGSRIAAMDEFIMK----LQMENKLREKER 440
P +D SST L + + A D + + M K EKE+
Sbjct: 385 PQTNKDVSSTSTLPASVGAEDFIDQRQEKPMDMGRKPLEKEK 510
>TC80383 similar to PIR|C85065|C85065 kinesin-like protein [imported] -
Arabidopsis thaliana, partial (26%)
Length = 897
Score = 75.9 bits (185), Expect = 5e-14
Identities = 49/155 (31%), Positives = 82/155 (52%), Gaps = 8/155 (5%)
Frame = +1
Query: 246 TYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDV--------PTVGGRLM 297
T + +++ + + R V T N++SSRSH + L + V G L
Sbjct: 97 TVVDVESVKEVAFLLNQAANSRSVGKTQMNEQSSRSHFVFTLRIYGVNESTDQQVQGVLN 276
Query: 298 LVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQ 357
L+D+AGSE + ++G TG K +T IN+ +L V+ ++A + H+PFR+SKLT LLQ
Sbjct: 277 LIDLAGSERLSRSGSTGDRLK-ETQAINKSLSSLSDVIFALAKKEDHIPFRNSKLTYLLQ 453
Query: 358 DSFEDDKSKILMILCASPDPKEIHKTISTLEYGAK 392
D SK LM + +PD +++ +L + ++
Sbjct: 454 PCLGGD-SKTLMFVNIAPDQASSGESLCSLRFASR 555
>TC80343 similar to PIR|C96661|C96661 kinesin-like protein 73641-79546
[imported] - Arabidopsis thaliana, partial (17%)
Length = 1406
Score = 68.6 bits (166), Expect = 9e-12
Identities = 50/143 (34%), Positives = 74/143 (50%), Gaps = 5/143 (3%)
Frame = +3
Query: 294 GRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLT 353
G L LVD+AGSE ++++ TG K + N+ AL V+ ++A HVP+R+SKLT
Sbjct: 24 GCLHLVDLAGSERVDRSEATGDRLK-EAHXXNKSLSALGDVIFALAQKSPHVPYRNSKLT 200
Query: 354 MLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKE-----ED 408
LLQ S ++K LM + +PD +TISTL++ + + G KE E
Sbjct: 201 QLLQSSL-GGQAKTLMFVQLNPDVASYSETISTLKFAERVSGVELGAARSNKEGRDVREL 377
Query: 409 SSSTVILGSRIAAMDEFIMKLQM 431
L +A DE I +LQ+
Sbjct: 378 MEQMSFLKDAMARKDEEIERLQL 446
>AJ499156 similar to GP|10176794|db kinesin-like protein {Arabidopsis
thaliana}, partial (22%)
Length = 678
Score = 63.2 bits (152), Expect = 4e-10
Identities = 49/169 (28%), Positives = 79/169 (45%), Gaps = 36/169 (21%)
Frame = +1
Query: 260 IQKVEKRRIVKSTLCNDRSSRSHCMVILDV--------PTVGGRLMLVDMAGSENIEQAG 311
+ + R V T N++SSRSH + L + V G L L+D+AGSE + ++G
Sbjct: 16 LNQAANSRSVGKTQMNEQSSRSHFVFTLRIYGVNESTDQQVQGVLNLIDLAGSERLSRSG 195
Query: 312 QTGFEAK-----------------------MQTA-----KINQGNIALKRVVESIANGDS 343
TG K +QT+ IN+ +L V+ ++A +
Sbjct: 196 STGDRLKETQVCKTIRCFLFYTVLLL*FIIIQTSFTCN*AINKSLSSLSDVIFALAKKED 375
Query: 344 HVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAK 392
H+PFR+SKLT LLQ D SK LM + +PD +++ +L + ++
Sbjct: 376 HIPFRNSKLTYLLQPCLGGD-SKTLMFVNIAPDQASSGESLCSLRFASR 519
>AL377028 SP|O23826|K12 125 kDa kinesin-related protein. [Common tobacco]
{Nicotiana tabacum}, partial (7%)
Length = 223
Score = 59.3 bits (142), Expect = 5e-09
Identities = 30/66 (45%), Positives = 45/66 (67%)
Frame = +2
Query: 294 GRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLT 353
G+L LVD+AGSENI ++G A+ + +IN+ + L RV+ ++ H+P+RDSKLT
Sbjct: 20 GKLNLVDLAGSENISRSGAREGRAR-EAGEINKSLLTLGRVINALVEHLGHIPYRDSKLT 196
Query: 354 MLLQDS 359
LL+DS
Sbjct: 197 RLLRDS 214
>BE203495 weakly similar to PIR|T48258|T48 kinesin-like protein - Arabidopsis
thaliana, partial (21%)
Length = 502
Score = 54.7 bits (130), Expect = 1e-07
Identities = 42/164 (25%), Positives = 70/164 (42%), Gaps = 13/164 (7%)
Frame = +2
Query: 59 IEVIARIRDYPDRK-----DKPLSVLQA------SSNSRSIRVKADFGYRD--FTLDGVS 105
+ VI R+R + + D P+S + S N ++ +K F R + LD
Sbjct: 41 VRVIVRVRPFLPHETSRNCDDPVSCISLLDQDFQSQNDVAVYLKDPFTSRKECYKLDSFF 220
Query: 106 VSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQAGIVYRALRD 165
E+ + +++ V I + G T+ YG TGSGK++TM G +Q G++ A+
Sbjct: 221 DQEDNNVGQIFEREVSPMIPAIFGGCNATVFAYGATGSGKTYTMQGTEEQPGLMPLAMST 400
Query: 166 ILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLL 209
IL C + Q++ E+Y + YDLL
Sbjct: 401 ILS----------------ICQNTSSTAQISYYEVYMDRCYDLL 484
>AW692096 homologue to GP|10140692|gb kinesin-like protein {Oryza sativa
(japonica cultivar-group)}, partial (14%)
Length = 387
Score = 53.1 bits (126), Expect = 4e-07
Identities = 34/97 (35%), Positives = 56/97 (57%), Gaps = 1/97 (1%)
Frame = +1
Query: 299 VDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGD-SHVPFRDSKLTMLLQ 357
+D+AGSE+ + TG K + + IN+ + L V+ ++ G SHVP+RDSKLT LLQ
Sbjct: 1 IDLAGSES-SKTETTGLRRK-EGSYINKSLLTLGTVIGKLSEGKASHVPYRDSKLTRLLQ 174
Query: 358 DSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAK 394
S + +I +P + +T +TL++ ++AK
Sbjct: 175 SSL-SGHGHVSLICTITPASSNMEETHNTLKFASRAK 282
>AW689037 similar to GP|10177316|d kinesin-like protein {Arabidopsis
thaliana}, partial (13%)
Length = 662
Score = 52.8 bits (125), Expect = 5e-07
Identities = 44/164 (26%), Positives = 75/164 (44%), Gaps = 16/164 (9%)
Frame = +3
Query: 193 VQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNA-----SKVKLEVMGKKAKNATY 247
++V+ +EI+ EE++DLL N G S NA ++V +++ + T
Sbjct: 51 IRVSFIEIFKEEVFDLLDPNASKGE--------SVCNAKFAAPARVPIQIRETLSGGITL 206
Query: 248 ISGNEAGKISKE-----IQKVEKRRIVKSTLCNDRSSRSHCMVILDVPTVGG------RL 296
E +KE + + R ST N +SSRSH + + + G +L
Sbjct: 207 AGVTEPEVKTKEEMSSYLSRGSMSRATGSTNMNSQSSRSHAIFTITMEQKNGDDVLCAKL 386
Query: 297 MLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIAN 340
LVD+AGSE ++ G G K + IN+G + L V+ ++ +
Sbjct: 387 HLVDLAGSERAKRTGADGMRLK-EGIHINKGLLTLGNVISALGD 515
>TC82643 similar to GP|10998143|dbj|BAB03114. kinesin (centromere protein)
like heavy chain-like protein {Arabidopsis thaliana},
partial (25%)
Length = 880
Score = 52.0 bits (123), Expect = 8e-07
Identities = 39/132 (29%), Positives = 66/132 (49%), Gaps = 13/132 (9%)
Frame = +2
Query: 278 SSRSHCMVILDVPTVG------------GRLMLVDMAGSENIEQAGQTGFEAKMQTAKIN 325
SSRSH + L V + +L L+D+AGSE+ + +T + + + IN
Sbjct: 11 SSRSHTIFTLTVESSPCGEYIEGEAVTLSQLNLIDLAGSESSK--AETIGMRRREGSYIN 184
Query: 326 QGNIALKRVVESIANGD-SHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTI 384
+ + L V+ + SH+P+RDSKLT +LQ S ++ +I +P +T
Sbjct: 185 KSLLTLGTVISKLTEAKASHIPYRDSKLTRVLQSSL-SGHGRVSLICTVTPSSSSSEETH 361
Query: 385 STLEYGAKAKCI 396
+TL++ +AK I
Sbjct: 362 NTLKFAHRAKHI 397
>TC88497 similar to GP|11994617|dbj|BAB02754. contains similarity to kinesin
protein~gene_id:MGL6.9 {Arabidopsis thaliana}, partial
(26%)
Length = 1310
Score = 51.2 bits (121), Expect = 1e-06
Identities = 38/139 (27%), Positives = 65/139 (46%), Gaps = 11/139 (7%)
Frame = +2
Query: 340 NGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRG 399
N H+PFR SKLT +L+DSF + SK +MI C SP+ T++TL Y + K + +
Sbjct: 113 NDQIHIPFRGSKLTEVLRDSFVGN-SKTVMISCISPNAGSCEHTLNTLRYADRVKSLSKS 289
Query: 400 PH---------TPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAHKKLMKK 450
+ P ++ ST L A D + + +E + + +K+++
Sbjct: 290 GNPRKDQAPNPVPQSNKEVLSTSSLPDSACAEDVYYQR-------QEVKTGDMGRKVIEN 448
Query: 451 EEEI--AALRTKVETAPAS 467
E + +A V+ P+S
Sbjct: 449 ENSLYSSAAAADVDKQPSS 505
>AW685809 similar to PIR|E84792|E8 probable kinesin heavy chain [imported] -
Arabidopsis thaliana, partial (14%)
Length = 651
Score = 50.1 bits (118), Expect = 3e-06
Identities = 44/174 (25%), Positives = 73/174 (41%), Gaps = 25/174 (14%)
Frame = +2
Query: 18 ITPRSKHRLNFNGVKPAPTP---PHPHPSPHPNFNNKDSPP--------EHPIEVIARIR 66
+TP ++ P+P P P P P+ D P E ++V+ R R
Sbjct: 101 MTPDHSKKVGLGMTIPSPAPFLTPRPERR-RPDSRGSDRNPNRQDNKDKETNVQVLLRCR 277
Query: 67 DYPDRKDKPL--SVLQASSNSRSIRVKADFGYRD----FTLDGVSVSEEEELDLFYKKFV 120
D + + V+ + N R + V + F D V + ++ + Y + +
Sbjct: 278 PLSDEEQRSNVPKVVSCNENKREVTVMHTIANKQVEKVFNFDKVFGPKSQQRSI-YDQAI 454
Query: 121 ESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSK--------QAGIVYRALRDI 166
+N V G CT+ YG TG+GK++TM G K +AG++ RA+R I
Sbjct: 455 APIVNEVLDGFNCTVFAYGQTGTGKTYTMEGGMKNKGGDLTAEAGVIPRAVRQI 616
>TC76591 similar to PIR|B85063|B85063 hypothetical protein AT4g05020
[imported] - Arabidopsis thaliana, partial (87%)
Length = 2283
Score = 45.4 bits (106), Expect = 8e-05
Identities = 33/137 (24%), Positives = 63/137 (45%), Gaps = 9/137 (6%)
Frame = -3
Query: 423 DEFIMKLQMENKLREKERNEAHKKLMKKEEEIAALRTKVE------TAPASEEEINLKVN 476
D F ++ + + K++ + ++ + VE + P S + +L +
Sbjct: 472 DWFHNAASLDGATGNTRQQRSESKIIARRNDVNLISGIVEIPQETCSCPTSSQHQHLFL- 296
Query: 477 ERTRHLRQELEKKLEECQRMTNEFVELERKRMEERILQQQEEVEILRKRLEEIELQLCSS 536
RH R L CQ+ T + E ++E R++ EI RK LEE+E+ CS
Sbjct: 295 --LRHRRFRCSNSLTVCQKTTTTYGGAEENKLELRVII----TEIFRKALEELEIPHCSK 134
Query: 537 SKQE---RKDENESKEM 550
SK E ++++NE++++
Sbjct: 133 SKVEGGKKREKNEARQL 83
>TC87535 similar to GP|19310542|gb|AAL85004.1 unknown protein {Arabidopsis
thaliana}, partial (86%)
Length = 893
Score = 43.1 bits (100), Expect = 4e-04
Identities = 38/140 (27%), Positives = 69/140 (49%), Gaps = 9/140 (6%)
Frame = +1
Query: 447 LMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNEFVELERK 506
L K++EE++ R+ + T A EEEI K E ++ +L + EE +R+ R+
Sbjct: 238 LSKEDEEMS--RSALSTFRAKEEEIEKKKMEVRDKVQFQLGRVEEETKRLAT-----IRE 396
Query: 507 RMEERILQQQEEVEILRKRLEEIELQL------CSSSKQERKDENES---KEMEPNGFMR 557
+E ++EV ++RKR++ + +L C ++E KD E+ K E +
Sbjct: 397 ELEALADPMRKEVSLVRKRIDSVNKELKPLGHTCQKKEKEYKDALEAFNEKNREKVQLIT 576
Query: 558 KLLSVYKSTDDLGMVKSMDL 577
KL+ + ++ L M K +L
Sbjct: 577 KLMELVGESERLRMKKLEEL 636
>TC91751 similar to GP|2739382|gb|AAC14505.1| unknown protein {Arabidopsis
thaliana}, partial (38%)
Length = 1200
Score = 42.7 bits (99), Expect = 5e-04
Identities = 35/132 (26%), Positives = 64/132 (47%)
Frame = +2
Query: 438 KERNEAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMT 497
+ RN ++L K +EEI R + ETA ++ ++ LK + T+ L +EL+
Sbjct: 266 ERRNLVEQELDKADEEIPEYRKQAETAEQTKNQV-LKELDSTKRLIEELK---------- 412
Query: 498 NEFVELERKRMEERILQQQEEVEILRKRLEEIELQLCSSSKQERKDENESKEMEPNGFMR 557
+ LER + EE+ Q +++ E+ + R+EE+E + S K + E + +
Sbjct: 413 ---LNLERAQTEEQ--QARQDSELAKLRVEEMEQGIADESSVAAKAQLEVAKARYTAAIT 577
Query: 558 KLLSVYKSTDDL 569
L +V + D L
Sbjct: 578 DLAAVKEELDAL 613
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.312 0.131 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,171,517
Number of Sequences: 36976
Number of extensions: 315416
Number of successful extensions: 7035
Number of sequences better than 10.0: 211
Number of HSP's better than 10.0 without gapping: 3300
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5782
length of query: 754
length of database: 9,014,727
effective HSP length: 103
effective length of query: 651
effective length of database: 5,206,199
effective search space: 3389235549
effective search space used: 3389235549
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC144481.6


