

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144481.4 + phase: 0
(152 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
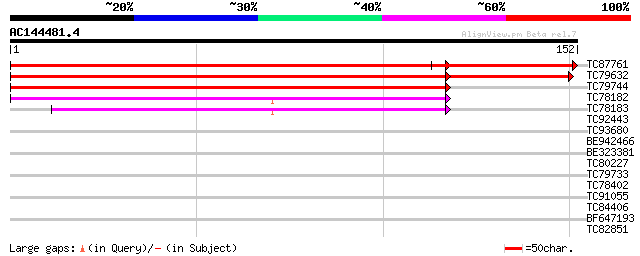
Score E
Sequences producing significant alignments: (bits) Value
TC87761 similar to GP|22296432|dbj|BAC10199. hypothetical protei... 223 4e-74
TC79632 similar to GP|22296432|dbj|BAC10199. hypothetical protei... 199 8e-59
TC79744 weakly similar to GP|9280228|dbj|BAB01718.1 gene_id:MXL8... 93 5e-20
TC78182 weakly similar to GP|21740504|emb|CAD40828. OSJNBa0006B2... 87 3e-18
TC78183 weakly similar to GP|9280228|dbj|BAB01718.1 gene_id:MXL8... 76 6e-15
TC92443 homologue to PIR|A85060|A85060 probable ABC transporter ... 31 0.17
TC93680 similar to GP|15293299|gb|AAK93760.1 unknown protein {Ar... 27 2.4
BE942466 similar to GP|8778726|gb| T25N20.14 {Arabidopsis thalia... 27 3.2
BE323381 similar to GP|11993471|g ubiquitin-specific protease 12... 26 5.4
TC80227 similar to GP|13537198|dbj|BAB40775. histidine kinase {A... 26 5.4
TC79733 similar to PIR|T01797|T01797 hypothetical protein A_TM02... 26 7.0
TC78402 weakly similar to PIR|B84853|B84853 hypothetical protein... 26 7.0
TC91055 weakly similar to GP|10440622|gb|AAG16860.1 putative NBS... 26 7.0
TC84406 similar to GP|12039327|gb|AAG46115.1 putative sugar tran... 25 9.2
BF647193 similar to PIR|T01227|T012 glutathione-regulated potass... 25 9.2
TC82851 similar to GP|22654962|gb|AAM98074.1 AT5g60420/muf9_70 {... 25 9.2
>TC87761 similar to GP|22296432|dbj|BAC10199. hypothetical protein~similar
to Arabidopsis thaliana chromosome 2 At2g33470,
complete
Length = 1050
Score = 223 bits (567), Expect(2) = 4e-74
Identities = 107/118 (90%), Positives = 112/118 (94%)
Frame = +1
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
M +VKSDIGGNISRLESKYLSNP KFNCLYSLVQ+EVETKTAKASSSCT GLLWLTRAMD
Sbjct: 307 MTLVKSDIGGNISRLESKYLSNPTKFNCLYSLVQIEVETKTAKASSSCTNGLLWLTRAMD 486
Query: 61 FLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
FLVA+FRNLIEHADWSMSQACTDSY KTLKKWHGWLASS+ TVAM LAPDRKKFMEV+
Sbjct: 487 FLVALFRNLIEHADWSMSQACTDSYNKTLKKWHGWLASSSFTVAMKLAPDRKKFMEVI 660
Score = 71.6 bits (174), Expect(2) = 4e-74
Identities = 36/39 (92%), Positives = 37/39 (94%)
Frame = +2
Query: 114 FMEVVISMLTLSNFVLAFLLSLKRITSFWLDLDWMN*RL 152
+ EVVISMLTLSNFVLAFLLSLKRITSFWLDLDWM *RL
Sbjct: 659 YREVVISMLTLSNFVLAFLLSLKRITSFWLDLDWMI*RL 775
>TC79632 similar to GP|22296432|dbj|BAC10199. hypothetical protein~similar
to Arabidopsis thaliana chromosome 2 At2g33470,
complete
Length = 1106
Score = 199 bits (507), Expect(2) = 8e-59
Identities = 94/118 (79%), Positives = 107/118 (90%)
Frame = +3
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
M +VKSDIGGNI+RLE+ Y SNP++FN LYSLVQVE+E+KTAK+SSSCT GLLWLTRAMD
Sbjct: 282 MTLVKSDIGGNITRLETLYSSNPSRFNILYSLVQVEIESKTAKSSSSCTNGLLWLTRAMD 461
Query: 61 FLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
FLVA+F+NLIEH DW MSQACTD+Y KTLKKWHGWLASS+ TV M LAPDRKKFMEV+
Sbjct: 462 FLVALFQNLIEHDDWPMSQACTDAYNKTLKKWHGWLASSSFTVVMKLAPDRKKFMEVI 635
Score = 43.5 bits (101), Expect(2) = 8e-59
Identities = 23/34 (67%), Positives = 26/34 (75%)
Frame = +1
Query: 118 VISMLTLSNFVLAFLLSLKRITSFWLDLDWMN*R 151
VISMLT NFV + LLSL RIT+FW L+WM *R
Sbjct: 646 VISMLT*RNFVQSSLLSLNRITNFWRVLNWMI*R 747
>TC79744 weakly similar to GP|9280228|dbj|BAB01718.1 gene_id:MXL8.12~unknown
protein {Arabidopsis thaliana}, partial (66%)
Length = 677
Score = 92.8 bits (229), Expect = 5e-20
Identities = 43/118 (36%), Positives = 72/118 (60%)
Frame = +3
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
MA+++ D+ NI RLE + SNP L ++++E AK SSC+ +WLTR +D
Sbjct: 195 MAVLRQDVYQNIKRLELMHESNPTTNLNLVEILKLEATEGNAKKGSSCSKAFVWLTRTLD 374
Query: 61 FLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
F ++ + L++ + M + +SY TLK WHGW++S+ V VA+ L P+ K F++++
Sbjct: 375 FTSSLLQILLKDPEKKMEKIVEESYEVTLKPWHGWISSTAVRVALRLVPESKTFIDLL 548
>TC78182 weakly similar to GP|21740504|emb|CAD40828. OSJNBa0006B20.19 {Oryza
sativa}, partial (29%)
Length = 810
Score = 87.0 bits (214), Expect = 3e-18
Identities = 46/119 (38%), Positives = 70/119 (58%), Gaps = 1/119 (0%)
Frame = +2
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
MA+++ DI NI RLE+ + SNP + L + + E K S + +WLTR++D
Sbjct: 305 MAVLRQDIHQNIKRLEAIHESNPLTNSNLVEIFKSETSKGNGKKRVSGSKSFVWLTRSLD 484
Query: 61 FLVAVFRNL-IEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
F A+ + L ++ +M QA +SY TLK WHGW+AS+ VA+ L PD K FM+++
Sbjct: 485 FTSALLQALLVKDPKKNMEQAVQESYDATLKPWHGWIASAAYRVAIKLVPDTKTFMDLL 661
>TC78183 weakly similar to GP|9280228|dbj|BAB01718.1 gene_id:MXL8.12~unknown
protein {Arabidopsis thaliana}, partial (45%)
Length = 1125
Score = 75.9 bits (185), Expect = 6e-15
Identities = 40/108 (37%), Positives = 62/108 (57%), Gaps = 1/108 (0%)
Frame = +2
Query: 12 ISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDFLVAVFRNL-I 70
+ RLE+ + SNP + L + + E K S + +WLTR++DF A+ + L +
Sbjct: 500 LQRLEAIHESNPLTNSNLVEIFKSETSKGNGKKRVSGSKSFVWLTRSLDFTSALLQALLV 679
Query: 71 EHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
+ +M QA +SY TLK WHGW+AS+ VA+ L PD K FM+++
Sbjct: 680 KDPKKNMEQAVQESYDATLKPWHGWIASAAYRVAIKLVPDTKTFMDLL 823
>TC92443 homologue to PIR|A85060|A85060 probable ABC transporter [imported]
- Arabidopsis thaliana, partial (29%)
Length = 803
Score = 31.2 bits (69), Expect = 0.17
Identities = 17/41 (41%), Positives = 24/41 (58%)
Frame = +3
Query: 13 SRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLL 53
SR+ SK +S NC LVQV+ + + AK SS C + L+
Sbjct: 120 SRIISKGISAGHSRNCYRGLVQVQSKAENAKNSSQCDSMLI 242
>TC93680 similar to GP|15293299|gb|AAK93760.1 unknown protein {Arabidopsis
thaliana}, partial (14%)
Length = 476
Score = 27.3 bits (59), Expect = 2.4
Identities = 14/38 (36%), Positives = 22/38 (57%)
Frame = -3
Query: 14 RLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTG 51
R S L+ P+K+ S VQ ++ TA+A+ +CT G
Sbjct: 234 RSTSNLLTIPSKYGVKISGVQFSLKLSTAEAA*ACTRG 121
>BE942466 similar to GP|8778726|gb| T25N20.14 {Arabidopsis thaliana}, partial
(4%)
Length = 647
Score = 26.9 bits (58), Expect = 3.2
Identities = 18/53 (33%), Positives = 28/53 (51%), Gaps = 1/53 (1%)
Frame = -2
Query: 66 FRNLIEHADWSMSQACTDSYYKTLKK-WHGWLASSTVTVAMMLAPDRKKFMEV 117
F+NL + D S +CT K + +H +L+ S T ++ L KKFME+
Sbjct: 340 FQNL--YRDIYHSVSCTKKVCKKINTFYHIYLSQSEQTYSIGLESQNKKFMEI 188
>BE323381 similar to GP|11993471|g ubiquitin-specific protease 12
{Arabidopsis thaliana}, partial (9%)
Length = 356
Score = 26.2 bits (56), Expect = 5.4
Identities = 17/46 (36%), Positives = 23/46 (49%), Gaps = 1/46 (2%)
Frame = -1
Query: 9 GGNISRLESKYLSNPAKFNC-LYSLVQVEVETKTAKASSSCTTGLL 53
GG I R + FN L++L++ + TA ASS TGLL
Sbjct: 344 GGGIGRKSCPISISKNNFNSPLFALLETSLNCPTACASS*AVTGLL 207
>TC80227 similar to GP|13537198|dbj|BAB40775. histidine kinase {Arabidopsis
thaliana}, partial (22%)
Length = 1693
Score = 26.2 bits (56), Expect = 5.4
Identities = 20/69 (28%), Positives = 29/69 (41%), Gaps = 1/69 (1%)
Frame = +2
Query: 73 ADWSMSQACTDSYYKTLKK-WHGWLASSTVTVAMMLAPDRKKFMEVVISMLTLSNFVLAF 131
AD+S C Y + K W+GW+ T+ L+ F + L S VL F
Sbjct: 1013 ADFSDDSGCNSGYT*EMPKVWNGWICFKTI*SRTTLSGSFSVFPVI----LKRSRKVLVF 1180
Query: 132 LLSLKRITS 140
L + I+S
Sbjct: 1181 LQRMLSISS 1207
>TC79733 similar to PIR|T01797|T01797 hypothetical protein A_TM021B04.9 -
Arabidopsis thaliana, partial (2%)
Length = 1253
Score = 25.8 bits (55), Expect = 7.0
Identities = 15/50 (30%), Positives = 24/50 (48%)
Frame = -3
Query: 84 SYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVVISMLTLSNFVLAFLL 133
+YY L+ W G A S+ + L P ++ I LSN VL+ ++
Sbjct: 1056 TYYSLLR*WCGTAAGSSSQKPLKLLPCSSVSLKAGILWENLSNSVLSIVM 907
>TC78402 weakly similar to PIR|B84853|B84853 hypothetical protein At2g42370
[imported] - Arabidopsis thaliana, partial (14%)
Length = 1982
Score = 25.8 bits (55), Expect = 7.0
Identities = 16/45 (35%), Positives = 23/45 (50%)
Frame = +2
Query: 99 STVTVAMMLAPDRKKFMEVVISMLTLSNFVLAFLLSLKRITSFWL 143
STV ++L R++ +V+ L L N V LLSL S W+
Sbjct: 71 STVLS*VVLLSFRRRIRLLVVLSLMLGNLVQRILLSLMSWQSNWM 205
>TC91055 weakly similar to GP|10440622|gb|AAG16860.1 putative NBS-LRR type
resistance protein 3' partial {Oryza sativa}, partial
(2%)
Length = 978
Score = 25.8 bits (55), Expect = 7.0
Identities = 11/31 (35%), Positives = 20/31 (64%)
Frame = -2
Query: 42 AKASSSCTTGLLWLTRAMDFLVAVFRNLIEH 72
+KA +S TTGL+ + + ++ + RNL+ H
Sbjct: 860 SKAKTSTTTGLIVTSSTPNDVIFIIRNLLIH 768
>TC84406 similar to GP|12039327|gb|AAG46115.1 putative sugar transporter
{Oryza sativa}, partial (13%)
Length = 753
Score = 25.4 bits (54), Expect = 9.2
Identities = 11/35 (31%), Positives = 19/35 (53%), Gaps = 4/35 (11%)
Frame = +2
Query: 67 RNLIEHADWSMSQACTDSYYKTLKK----WHGWLA 97
R+ +E + + SQAC D + +T WH W++
Sbjct: 419 RSKMESSSRTWSQACIDCWNRTSNSSTGCWHKWIS 523
>BF647193 similar to PIR|T01227|T012 glutathione-regulated potassium-efflux
system protein kefB homolog F6N23.15 - Arabidopsis
thaliana, partial (11%)
Length = 590
Score = 25.4 bits (54), Expect = 9.2
Identities = 13/34 (38%), Positives = 21/34 (61%)
Frame = +3
Query: 99 STVTVAMMLAPDRKKFMEVVISMLTLSNFVLAFL 132
S ++ +LAP KFM +IS+L L ++AF+
Sbjct: 444 SFLSCRKILAPKFVKFMSRIISLLLLLYMLMAFV 545
>TC82851 similar to GP|22654962|gb|AAM98074.1 AT5g60420/muf9_70 {Arabidopsis
thaliana}, partial (14%)
Length = 874
Score = 25.4 bits (54), Expect = 9.2
Identities = 9/31 (29%), Positives = 16/31 (51%)
Frame = +1
Query: 65 VFRNLIEHADWSMSQACTDSYYKTLKKWHGW 95
+F + I+ W +C + + L+KW GW
Sbjct: 376 LFWSYIKRGFWKS*TSCYSNEQQWLRKWSGW 468
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,196,636
Number of Sequences: 36976
Number of extensions: 66323
Number of successful extensions: 524
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 520
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 522
length of query: 152
length of database: 9,014,727
effective HSP length: 88
effective length of query: 64
effective length of database: 5,760,839
effective search space: 368693696
effective search space used: 368693696
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC144481.4


