

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144477.5 - phase: 1 /pseudo
(206 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
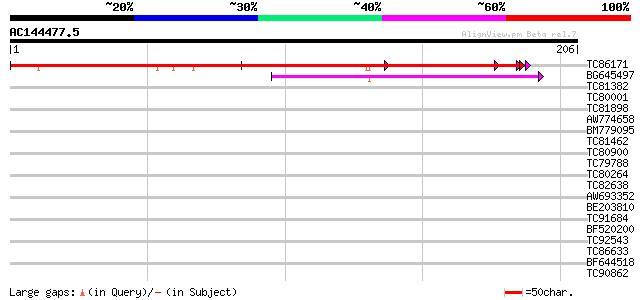
Sequences producing significant alignments: (bits) Value
TC86171 weakly similar to GP|13507547|gb|AAK28636.1 unknown prot... 286 3e-78
BG645497 similar to GP|13507547|gb unknown protein {Arabidopsis ... 52 2e-07
TC81382 weakly similar to GP|14269077|gb|AAK58011.1 verticillium... 33 0.077
TC80001 weakly similar to GP|15451136|gb|AAK96839.1 periaxin-lik... 33 0.100
TC81898 weakly similar to GP|20260576|gb|AAM13186.1 putative rec... 32 0.17
AW774658 similar to GP|2808681|emb| Hcr9-4B {Lycopersicon hirsut... 32 0.22
BM779095 weakly similar to PIR|B84431|B8 probable receptor prote... 32 0.22
TC81462 similar to GP|21536497|gb|AAM60829.1 F-box protein famil... 31 0.38
TC80900 weakly similar to PIR|G84652|G84652 probable receptor-li... 31 0.38
TC79788 weakly similar to GP|21554189|gb|AAM63268.1 putative leu... 30 0.50
TC80264 similar to GP|21689699|gb|AAM67471.1 putative F-box fami... 30 0.50
TC82638 weakly similar to PIR|T00712|T00712 protein kinase homol... 30 0.65
AW693352 weakly similar to GP|20466532|g unknown protein {Arabid... 30 0.85
BE203810 weakly similar to PIR|A96574|A96 protein F12M16.30 [imp... 30 0.85
TC91684 weakly similar to PIR|A96574|A96574 protein F12M16.30 [i... 29 1.4
BF520200 weakly similar to GP|6714444|gb|A putative disease resi... 29 1.4
TC92543 weakly similar to GP|6041804|gb|AAF02124.1| putative pro... 29 1.4
TC86633 weakly similar to SP|Q00874|D100_ARATH DNA-damage-repair... 29 1.4
BF644518 weakly similar to GP|9294230|dbj| disease resistance pr... 29 1.4
TC90862 similar to GP|21740496|emb|CAD40820. OSJNBa0006B20.11 {O... 28 2.5
>TC86171 weakly similar to GP|13507547|gb|AAK28636.1 unknown protein
{Arabidopsis thaliana}, partial (89%)
Length = 2343
Score = 286 bits (733), Expect = 3e-78
Identities = 159/191 (83%), Positives = 166/191 (86%), Gaps = 4/191 (2%)
Frame = +1
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHM----FPFL 56
LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFS L + F
Sbjct: 1057 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSGLTGLKRLSLAFN 1236
Query: 57 QVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISG 116
++ A + LT+L+ +IGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISG
Sbjct: 1237 KITDACLVHLKGLTKLEYLNLDSCQIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISG 1416
Query: 117 LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGA 176
LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGA
Sbjct: 1417 LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGA 1596
Query: 177 RITDSGTTYLR 187
RITDSGTTYLR
Sbjct: 1597 RITDSGTTYLR 1629
Score = 63.9 bits (154), Expect = 4e-11
Identities = 61/197 (30%), Positives = 93/197 (46%), Gaps = 11/197 (5%)
Frame = +1
Query: 1 LQKLSTLNV-EGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQV- 58
L L++L++ + C++T S L L L+L RC GF F L+ + L +
Sbjct: 763 LSNLTSLSIRKSCAVTPDGMRAFSNLVNLEKLDLERCSDIHGGFVHFKGLKKL-ESLNIG 939
Query: 59 ---------LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNS 109
++A G ++L LQ I D G+ L GL L +L + + +
Sbjct: 940 CCKCVTDSDMKAISG-FINLKELQISN---SSITDLGISYLRGLQKLSTLNVEGCSITAA 1107
Query: 110 GIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLI 169
YIS L L LNL+ ++D+G ++ GLT LK L+L +ITDA L +L L+ L
Sbjct: 1108CFEYISALAALACLNLNRCGLSDDGFEKFSGLTGLKRLSLAFNKITDACLVHLKGLTKLE 1287
Query: 170 TLDLFGARITDSGTTYL 186
L+L +I D G L
Sbjct: 1288YLNLDSCQIGDEGLVNL 1338
Score = 58.9 bits (141), Expect = 1e-09
Identities = 52/183 (28%), Positives = 87/183 (47%), Gaps = 5/183 (2%)
Frame = +1
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHM----FPFL 56
L L +L + + + YIS L L LNL+ ++D+G ++ L ++
Sbjct: 1345 LTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDAR 1524
Query: 57 QVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISG 116
Q+ A + L+ L T F +I D G L L+SL + + ++G++ I
Sbjct: 1525 QITDAGLANLTSLSGLITLDLFGARITDSGTTYLRSFKNLQSLEICGGLLTDAGVKNIRE 1704
Query: 117 LNKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFG 175
+ L LNLS +TD L+ + G+T L+SLN+ ++T+ GL L L L TL L
Sbjct: 1705 IVSLTQLNLSQNCKLTDKTLELISGMTALRSLNVSNSRVTNEGLRYLKPLKNLRTLSLES 1884
Query: 176 ARI 178
++
Sbjct: 1885 CKV 1893
Score = 55.1 bits (131), Expect = 2e-08
Identities = 35/106 (33%), Positives = 53/106 (49%), Gaps = 1/106 (0%)
Frame = +1
Query: 85 EGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLLGLTN 143
+G+ + L L+ L L + G + GL KLE LN+ VTD+ +K + G N
Sbjct: 814 DGMRAFSNLVNLEKLDLERCSDIHGGFVHFKGLKKLESLNIGCCKCVTDSDMKAISGFIN 993
Query: 144 LKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSGTTYLRCM 189
LK L + ITD G++ L L L TL++ G IT + Y+ +
Sbjct: 994 LKELQISNSSITDLGISYLRGLQKLSTLNVEGCSITAACFEYISAL 1131
Score = 47.4 bits (111), Expect = 4e-06
Identities = 42/149 (28%), Positives = 71/149 (47%), Gaps = 11/149 (7%)
Frame = +1
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVLQ 60
L L +LN++ IT A +++L+ L L+L ++D G + F LQ L+
Sbjct: 1489 LTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSG----TTYLRSFKNLQSLE 1656
Query: 61 A*RG*-----------VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNS 109
G ++ LT+L K+ D+ L ++G+T L+SL +S++ V N
Sbjct: 1657 ICGGLLTDAGVKNIREIVSLTQLNLSQNC--KLTDKTLELISGMTALRSLNVSNSRVTNE 1830
Query: 110 GIRYISGLNKLEDLNLSFTSVTDNGLKRL 138
G+RY+ L L L+L V +K+L
Sbjct: 1831 GLRYLKPLKNLRTLSLESCKVNAADIKKL 1917
>BG645497 similar to GP|13507547|gb unknown protein {Arabidopsis thaliana},
partial (18%)
Length = 765
Score = 51.6 bits (122), Expect = 2e-07
Identities = 34/100 (34%), Positives = 57/100 (57%), Gaps = 1/100 (1%)
Frame = -2
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLDARQI 154
L+SL + + ++G++ I L+ L LNLS S +TD ++ + GLT L SLNL +I
Sbjct: 425 LRSLEICSGGLTDAGVKNIKELSSLMCLNLSQNSNLTDKTVELIAGLTALVSLNLSNTRI 246
Query: 155 TDAGLANLTSLSGLITLDLFGARITDSGTTYLRCMYIYQL 194
T AGL +L +L L +L L ++T + + +++ L
Sbjct: 245 TSAGLQHLKTLKNLRSLTLESCKVTANDIKKFKLIHLPNL 126
>TC81382 weakly similar to GP|14269077|gb|AAK58011.1 verticillium wilt
disease resistance protein Ve2 {Lycopersicon
esculentum}, partial (16%)
Length = 1974
Score = 33.1 bits (74), Expect = 0.077
Identities = 23/88 (26%), Positives = 44/88 (49%)
Frame = +3
Query: 88 VNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSL 147
+N+ L L LSD + + + ++ LE LNL + ++T + L T+L+ L
Sbjct: 513 INVNRSNGLSGLDLSDNLLDDEIPLVVCNISSLEFLNLGYNNLTGIIPQCLAESTSLQVL 692
Query: 148 NLDARQITDAGLANLTSLSGLITLDLFG 175
NL + +N + S +++L+L+G
Sbjct: 693 NLQMNRFHGTLPSNFSKHSKIVSLNLYG 776
>TC80001 weakly similar to GP|15451136|gb|AAK96839.1 periaxin-like protein
{Arabidopsis thaliana}, partial (38%)
Length = 532
Score = 32.7 bits (73), Expect = 0.100
Identities = 32/106 (30%), Positives = 48/106 (45%), Gaps = 2/106 (1%)
Frame = -1
Query: 83 GDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLT 142
GD G++ +G +L L + G SG ISG L S S+ D+G+ GL
Sbjct: 448 GDSGILGSSGFGNSGTLNLGSSCFGISGTLGISGFGNSGPLGCS--SLGDSGILGSSGLG 275
Query: 143 NLKSLNLDARQITDAGLANLT--SLSGLITLDLFGARITDSGTTYL 186
N +LNL + ++G ++ +SG FG +SGT L
Sbjct: 274 NSGTLNLGSSGFGNSGTLGISIFGISGTFGKSDFG----NSGTLIL 149
Score = 30.0 bits (66), Expect = 0.65
Identities = 28/101 (27%), Positives = 44/101 (42%)
Frame = -1
Query: 83 GDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLT 142
G G + ++G + S L + G+SGI SG LNL + +G + G
Sbjct: 514 GISGTLGISGF--VNSGTLGCSNFGDSGILGSSGFGNSGTLNLGSSCFGISGTLGISGFG 341
Query: 143 NLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSGT 183
N S L + D+G+ + L TL+L + +SGT
Sbjct: 340 N--SGPLGCSSLGDSGILGSSGLGNSGTLNLGSSGFGNSGT 224
>TC81898 weakly similar to GP|20260576|gb|AAM13186.1 putative receptor-like
protein kinase {Arabidopsis thaliana}, partial (29%)
Length = 1103
Score = 32.0 bits (71), Expect = 0.17
Identities = 28/86 (32%), Positives = 41/86 (47%)
Frame = +3
Query: 88 VNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSL 147
+NL+G+ L + DT +G LNKL L+LS +T LT+LKSL
Sbjct: 222 LNLSGIGLTGPI--PDTTIGK--------LNKLHSLDLSNNKITTLP-SDFWSLTSLKSL 368
Query: 148 NLDARQITDAGLANLTSLSGLITLDL 173
NL + I+ + N+ + L DL
Sbjct: 369 NLSSNHISGSLTNNIGNFGLLENFDL 446
>AW774658 similar to GP|2808681|emb| Hcr9-4B {Lycopersicon hirsutum}, partial
(4%)
Length = 665
Score = 31.6 bits (70), Expect = 0.22
Identities = 24/60 (40%), Positives = 31/60 (51%), Gaps = 1/60 (1%)
Frame = +1
Query: 96 LKSLVLSDTEV-GNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQI 154
L+ L LSD ++ G G L KLE L LS ++ + L L GLT L +L LD I
Sbjct: 70 LRLLDLSDNDIQGWIGNEDFPRLTKLETLGLSSNNLNSSILSSLNGLTALTTLYLDFNNI 249
>BM779095 weakly similar to PIR|B84431|B8 probable receptor protein kinase
[imported] - Arabidopsis thaliana, partial (15%)
Length = 721
Score = 31.6 bits (70), Expect = 0.22
Identities = 27/101 (26%), Positives = 46/101 (44%), Gaps = 3/101 (2%)
Frame = +3
Query: 82 IGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVT---DNGLKRL 138
+ D ++L+ T LKSL L+ + + + LNKL+ L+LS +T + L +
Sbjct: 186 LSDSFPISLSNCTSLKSLNLASNFISGGIPKALGQLNKLQSLDLSHNQITGWIPSELSNV 365
Query: 139 LGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARIT 179
G +L L L IT + S + L +D+ +T
Sbjct: 366 CG--SLLELKLSFNNITGSVPFGFNSCTWLQLVDISNNNMT 482
Score = 29.6 bits (65), Expect = 0.85
Identities = 28/87 (32%), Positives = 42/87 (48%), Gaps = 1/87 (1%)
Frame = +3
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQIT 155
L L LS + +S +S L+ LNL+ ++ K L L L+SL+L QIT
Sbjct: 156 LLQLDLSGNNLSDSFPISLSNCTSLKSLNLASNFISGGIPKALGQLNKLQSLDLSHNQIT 335
Query: 156 DAGLANLTSLSG-LITLDLFGARITDS 181
+ L+++ G L+ L L IT S
Sbjct: 336 GWIPSELSNVCGSLLELKLSFNNITGS 416
Score = 28.5 bits (62), Expect = 1.9
Identities = 28/84 (33%), Positives = 41/84 (48%)
Frame = +3
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQIT 155
L+SL LS + S L L+LS +++D+ L T+LKSLNL + I+
Sbjct: 84 LQSLDLSFNNLTGSISDIKIDCKSLLQLDLSGNNLSDSFPISLSNCTSLKSLNLASNFIS 263
Query: 156 DAGLANLTSLSGLITLDLFGARIT 179
L L+ L +LDL +IT
Sbjct: 264 GGIPKALGQLNKLQSLDLSHNQIT 335
>TC81462 similar to GP|21536497|gb|AAM60829.1 F-box protein family AtFBL4
{Arabidopsis thaliana}, partial (67%)
Length = 1452
Score = 30.8 bits (68), Expect = 0.38
Identities = 28/65 (43%), Positives = 32/65 (49%), Gaps = 2/65 (3%)
Frame = +2
Query: 98 SLVLSDTEVGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLLG-LTNLKSLNLDARQIT 155
SL LSD N I G KLE L L + S VT GL L +LKSL+L +
Sbjct: 452 SLCLSD----NGLIALADGFPKLEKLKLIWCSNVTSFGLSSLASKCASLKSLDLQGCYVG 619
Query: 156 DAGLA 160
D GLA
Sbjct: 620 DQGLA 634
>TC80900 weakly similar to PIR|G84652|G84652 probable receptor-like protein
kinase [imported] - Arabidopsis thaliana, partial (24%)
Length = 1054
Score = 30.8 bits (68), Expect = 0.38
Identities = 24/78 (30%), Positives = 38/78 (47%)
Frame = +2
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQIT 155
LK + L + + I L L LNL + ++T + L LTNL+ L L ++T
Sbjct: 20 LKWIYLGYNNLSREIPKNIGNLVSLNHLNLVYNNLTGPIPESLGNLTNLQYLFLYLNKLT 199
Query: 156 DAGLANLTSLSGLITLDL 173
++ +L LI+LDL
Sbjct: 200 GPIPKSIFNLKNLISLDL 253
Score = 28.1 bits (61), Expect = 2.5
Identities = 30/112 (26%), Positives = 48/112 (42%)
Frame = +2
Query: 68 HLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSF 127
+LT LQ ++ K+ ++ L L SL LSD + + L KLE L+L
Sbjct: 152 NLTNLQYLFLYLNKLTGPIPKSIFNLKNLISLDLSDNYLSGEISNLVVNLQKLEILHLFS 331
Query: 128 TSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARIT 179
+ T + L +L+ L L + ++T L + L LDL +T
Sbjct: 332 NNFTGKIPNTITSLPHLQVLQLWSNKLTGEIPQTLGIHNNLTILDLSSNNLT 487
>TC79788 weakly similar to GP|21554189|gb|AAM63268.1 putative leucine-rich
repeat disease resistance protein {Arabidopsis
thaliana}, partial (52%)
Length = 1484
Score = 30.4 bits (67), Expect = 0.50
Identities = 26/96 (27%), Positives = 41/96 (42%)
Frame = +3
Query: 86 GLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLK 145
G T T + + L S IS L +L L+LS + + + L+NLK
Sbjct: 291 GFTCTTDSTHINQITLDPASYSGSLTPLISKLTQLITLDLSDNNFFGSIPSSISSLSNLK 470
Query: 146 SLNLDARQITDAGLANLTSLSGLITLDLFGARITDS 181
+L L + + ++ SL L +LDL +T S
Sbjct: 471 TLTLRSNSFSGPIPPSIISLKSLESLDLSHNSLTGS 578
>TC80264 similar to GP|21689699|gb|AAM67471.1 putative F-box family protein
AtFBL3 {Arabidopsis thaliana}, partial (4%)
Length = 1305
Score = 30.4 bits (67), Expect = 0.50
Identities = 29/84 (34%), Positives = 44/84 (51%), Gaps = 5/84 (5%)
Frame = +3
Query: 104 TEVGNSGIRYIS-GLNKLEDLNL-SFTSVTDNGLKRLL-GLTNLKSLNLD-ARQITDAGL 159
T + + G+ +I+ KL +L+L + D+GL L G L LNL +ITDAGL
Sbjct: 834 TNISDIGLAHIACNCPKLTELDLYRCVRIGDDGLAALTTGCNKLAMLNLAYCNRITDAGL 1013
Query: 160 ANLTSLSGLITLDLFG-ARITDSG 182
+++L L +L G + IT G
Sbjct: 1014KCISNLGELSDFELRGLSNITSIG 1085
>TC82638 weakly similar to PIR|T00712|T00712 protein kinase homolog F22O13.7
- Arabidopsis thaliana, partial (27%)
Length = 1074
Score = 30.0 bits (66), Expect = 0.65
Identities = 23/77 (29%), Positives = 33/77 (41%)
Frame = +1
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQIT 155
++ L LS + S I L L LNL + K + LT+LKSL++ T
Sbjct: 214 VEKLNLSHMNLSGSVSNEIQSLKSLTFLNLCCNGFESSLSKHITNLTSLKSLDVSQNFFT 393
Query: 156 DAGLANLTSLSGLITLD 172
L S L+TL+
Sbjct: 394 GGFPLGLGKASELLTLN 444
>AW693352 weakly similar to GP|20466532|g unknown protein {Arabidopsis
thaliana}, partial (10%)
Length = 499
Score = 29.6 bits (65), Expect = 0.85
Identities = 27/79 (34%), Positives = 37/79 (46%), Gaps = 2/79 (2%)
Frame = +1
Query: 117 LNKLEDLNLSFTSVTD--NGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLF 174
L LE L+LSF + + L L ++K N ++ A LTSLS L LDL
Sbjct: 4 LTNLEYLDLSFNKMKTLPPEISSLKALISMKVANNKLVELPPA----LTSLSRLENLDLS 171
Query: 175 GARITDSGTTYLRCMYIYQ 193
R+T G+ L M+ Q
Sbjct: 172 NNRLTSLGSLELSSMHRLQ 228
>BE203810 weakly similar to PIR|A96574|A96 protein F12M16.30 [imported] -
Arabidopsis thaliana, partial (18%)
Length = 490
Score = 29.6 bits (65), Expect = 0.85
Identities = 20/64 (31%), Positives = 30/64 (46%)
Frame = +1
Query: 81 KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLG 140
K D L GLT +++LVL + Y+ + L+ L+LSF +T L G
Sbjct: 292 KGSDSPFPQLIGLTNIETLVLRSCNLIGEVPDYLGHITTLKSLDLSFNKLTGPIPNTLGG 471
Query: 141 LTNL 144
L N+
Sbjct: 472 LKNI 483
>TC91684 weakly similar to PIR|A96574|A96574 protein F12M16.30 [imported] -
Arabidopsis thaliana, partial (20%)
Length = 1085
Score = 28.9 bits (63), Expect = 1.4
Identities = 17/53 (32%), Positives = 27/53 (50%)
Frame = +3
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMF 53
+ LS L + C+I+ A EY+ L L ++L+ LS F LQ+M+
Sbjct: 537 MSNLSKLVLRNCNISGALPEYLGKLTNLKVIDLSNNKLSGQIPVSFDGLQNMY 695
>BF520200 weakly similar to GP|6714444|gb|A putative disease resistance
protein {Arabidopsis thaliana}, partial (8%)
Length = 621
Score = 28.9 bits (63), Expect = 1.4
Identities = 26/91 (28%), Positives = 43/91 (46%)
Frame = +2
Query: 83 GDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLT 142
G E L LLKS+ LS I+ L +L LNLS ++T + LT
Sbjct: 38 GVEQLFINNRFVLLKSIDLSSNHFSEEIPPEIANLIQLVSLNLSRNNLTGKIPSNIGRLT 217
Query: 143 NLKSLNLDARQITDAGLANLTSLSGLITLDL 173
+L+ L+L ++ + ++L+ + L LD+
Sbjct: 218 SLEFLDLSQNKLFGSIPSSLSQIDRLGGLDV 310
>TC92543 weakly similar to GP|6041804|gb|AAF02124.1| putative protein kinase
{Arabidopsis thaliana}, partial (10%)
Length = 685
Score = 28.9 bits (63), Expect = 1.4
Identities = 23/81 (28%), Positives = 42/81 (51%), Gaps = 3/81 (3%)
Frame = +1
Query: 89 NLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLN 148
+L + +LK L L+ + S + + L+ L+LS S+T K + + NL ++
Sbjct: 61 SLGQMKVLKFLSLAGNNLSGSIPTSLGQMYSLQVLDLSTNSLTGEIPKFIENMRNLTNVL 240
Query: 149 LDARQITD---AGLANLTSLS 166
L+ ++ AGL N+T+LS
Sbjct: 241 LNNNNLSGHIPAGLVNVTTLS 303
>TC86633 weakly similar to SP|Q00874|D100_ARATH DNA-damage-repair/toleration
protein DRT100 precursor. [Mouse-ear cress] {Arabidopsis
thaliana}, partial (93%)
Length = 1407
Score = 28.9 bits (63), Expect = 1.4
Identities = 25/85 (29%), Positives = 45/85 (52%), Gaps = 1/85 (1%)
Frame = +3
Query: 96 LKSLVLSDTEVGNSGI-RYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQI 154
L +LV++D + + I I+ L+ L L+L+ ++ N + L +L LNL I
Sbjct: 339 LTTLVVADWKSISGEIPSCITSLSSLRILDLTGNKISGNIPGNIGKLQHLTVLNLADNAI 518
Query: 155 TDAGLANLTSLSGLITLDLFGARIT 179
+ ++ +SGL+ LDL G +I+
Sbjct: 519 SGEIPMSIVRISGLMHLDLAGNQIS 593
>BF644518 weakly similar to GP|9294230|dbj| disease resistance protein-like
{Arabidopsis thaliana}, partial (12%)
Length = 446
Score = 28.9 bits (63), Expect = 1.4
Identities = 16/42 (38%), Positives = 26/42 (61%)
Frame = -1
Query: 109 SGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLD 150
+ I Y L +L+LSF S+T+N + LT+L++L+LD
Sbjct: 143 NSIHYRKX*QNLCELHLSFNSLTENIPTEMNRLTDLEALHLD 18
>TC90862 similar to GP|21740496|emb|CAD40820. OSJNBa0006B20.11 {Oryza
sativa}, partial (14%)
Length = 1090
Score = 28.1 bits (61), Expect = 2.5
Identities = 17/40 (42%), Positives = 23/40 (57%)
Frame = +3
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSD 40
L+K+S L+ E CS +CF+ +S L L L GLSD
Sbjct: 837 LEKISVLSDEYCSFRKSCFQQLSFLPLLLELVF---GLSD 947
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.335 0.148 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,724,100
Number of Sequences: 36976
Number of extensions: 67975
Number of successful extensions: 641
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 592
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 628
length of query: 206
length of database: 9,014,727
effective HSP length: 92
effective length of query: 114
effective length of database: 5,612,935
effective search space: 639874590
effective search space used: 639874590
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144477.5


