

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144475.2 - phase: 0 /pseudo
(656 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
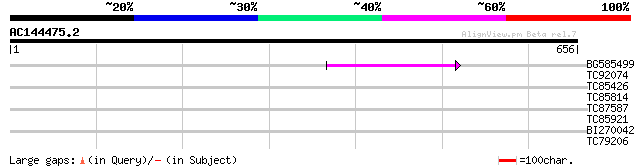
BG585499 98 1e-20
TC92074 32 0.59
TC85426 similar to SP|O23758|NLTP_CICAR Nonspecific lipid-transf... 31 1.3
TC85814 similar to SP|P48631|FD62_SOYBN Omega-6 fatty acid desat... 31 1.3
TC87587 similar to GP|20259233|gb|AAM14332.1 putative ankyrin pr... 29 6.6
TC85921 weakly similar to GP|11908074|gb|AAG41466.1 unknown prot... 29 6.6
BI270042 similar to GP|17979097|gb putative N-acetylornithine de... 28 8.6
TC79206 similar to GP|20339364|gb|AAM19355.1 UOS1 {Pisum sativum... 28 8.6
>BG585499
Length = 792
Score = 97.8 bits (242), Expect = 1e-20
Identities = 69/156 (44%), Positives = 82/156 (52%), Gaps = 1/156 (0%)
Frame = +1
Query: 367 SQFKVLILCNIKIARFWKVTGKVFGSGMAHIASKPLFGWLRTSAFLQISKGVNGVVEYLL 426
S+ +VL + + + W V GK FG G I K L G L FLQ V GV + L
Sbjct: 166 SKLRVLTIYLFRTNQLWIVIGKCFGDGGDRIVLKLLCG*LHMVVFLQTIVVVGGVRGFWL 345
Query: 427 IARIVEMKMKLFFMCFAIASTLPRYGFTSFHMIL*LISF-LSIAGIGSSITSSRKTLGRT 485
+VEM KLFFMCF IA RYG +I *LISF L IAGIG S T ++ +
Sbjct: 346 HVPVVEMLTKLFFMCFVIAGQRLRYGLDLCLLIG*LISFPLMIAGIGFSKTLAKDRMEFL 525
Query: 486 L*VGRQLS*QHVGTCGSGGIKLFLKLILLGRTTQFL 521
G QLS* HVG CG G KLFLK + TQ +
Sbjct: 526 SLSGSQLS*LHVGICGLGETKLFLKKDFKDQITQLM 633
>TC92074
Length = 423
Score = 32.3 bits (72), Expect = 0.59
Identities = 27/99 (27%), Positives = 52/99 (52%), Gaps = 4/99 (4%)
Frame = +3
Query: 31 IYFLLMIYFFLQRPQLNRLIVFCIALIFFVKPLVKR*IE---IKLVYISPRMLIPNFARI 87
IYF+L++ FF+ ++I+F + V+ + ++ + LV+++ R +
Sbjct: 15 IYFMLLLSFFVYFLSQPKVILFHAQHMLNVREICVNFLKLCGVLLVFVNVRKSNK*VEK* 194
Query: 88 FYITQGLIRLIAWACILGLILLLEGV*EASL-IILLTKF 125
FYIT I ++ I+ LILL +*+ + II+L K+
Sbjct: 195 FYITCNCIIILEGTLIIFLILLFVFL*QVQICIIILQKY 311
>TC85426 similar to SP|O23758|NLTP_CICAR Nonspecific lipid-transfer protein
precursor (LTP). [Chickpea Garbanzo] {Cicer arietinum},
complete
Length = 696
Score = 31.2 bits (69), Expect = 1.3
Identities = 20/46 (43%), Positives = 26/46 (56%)
Frame = +2
Query: 571 WGLLLDVAAFFVIQTVDGSKATLRRLELVMPYMLRCGDYIWVWTWL 616
W L+L VA F+ QTV S L L ++ Y L G+ WV+TWL
Sbjct: 512 WRLILYVALSFLYQTVFDS---LLPLHVLFLYEL-LGEIFWVFTWL 637
>TC85814 similar to SP|P48631|FD62_SOYBN Omega-6 fatty acid desaturase
endoplasmic reticulum isozyme 2 (EC 1.14.99.-).
[Soybean], complete
Length = 1897
Score = 31.2 bits (69), Expect = 1.3
Identities = 29/114 (25%), Positives = 47/114 (40%)
Frame = +3
Query: 336 LTLLIKFLRFQSLMMMMVQTLLGGVEPIPTISQFKVLILCNIKIARFWKVTGKVFGSGMA 395
LT+L L S M+ M +P I+ L L ++ +G++
Sbjct: 447 LTVLSALLSIHSPMLFMTSP*PSSSITLPPITSSTFLAL---------SLSWPGLLTGLS 599
Query: 396 HIASKPLFGWLRTSAFLQISKGVNGVVEYLLIARIVEMKMKLFFMCFAIASTLP 449
+AS P+FG L S S +NG++ L + + F AIA+T+P
Sbjct: 600 KVASLPVFGSLHMSVATMHSVTINGLMIQLALFSTLPFLCLTFHGNTAIAATIP 761
>TC87587 similar to GP|20259233|gb|AAM14332.1 putative ankyrin protein
{Arabidopsis thaliana}, partial (70%)
Length = 1641
Score = 28.9 bits (63), Expect = 6.6
Identities = 9/21 (42%), Positives = 15/21 (70%)
Frame = +2
Query: 20 CAQAGMVHKYLIYFLLMIYFF 40
C++ G H Y++YF L+ +FF
Sbjct: 1316 CSRIGENHNYMVYFYLIFFFF 1378
>TC85921 weakly similar to GP|11908074|gb|AAG41466.1 unknown protein
{Arabidopsis thaliana}, partial (38%)
Length = 1032
Score = 28.9 bits (63), Expect = 6.6
Identities = 18/57 (31%), Positives = 27/57 (46%)
Frame = +3
Query: 278 GILGMVVILTFG*ISGFLTMNHLCL*VTRPTWI*LYRLGMFLVLLEIGTLISS*ITC 334
G+ G +V L IS + + *+ R WI L M L + + L+S+*I C
Sbjct: 684 GLFGSIVTLLLIWISAHSIHSRVPK*IQRKLWIILLHTYMHLFIFVVNCLLSN*IVC 854
>BI270042 similar to GP|17979097|gb putative N-acetylornithine deacetylase
{Arabidopsis thaliana}, partial (26%)
Length = 438
Score = 28.5 bits (62), Expect = 8.6
Identities = 14/22 (63%), Positives = 16/22 (72%), Gaps = 2/22 (9%)
Frame = +2
Query: 383 WK--VTGKVFGSGMAHIASKPL 402
WK VTGK+F SG+AH A PL
Sbjct: 323 WKLHVTGKMFHSGLAHKAINPL 388
>TC79206 similar to GP|20339364|gb|AAM19355.1 UOS1 {Pisum sativum}, partial
(40%)
Length = 842
Score = 28.5 bits (62), Expect = 8.6
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = -3
Query: 27 HKYLIYFLLMIYFFLQRPQLNRLIVFC 53
H Y++Y I F+L +LN L VFC
Sbjct: 723 HSYVLYLFW*ITFYLWLKELNSLTVFC 643
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.356 0.161 0.537
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,191,905
Number of Sequences: 36976
Number of extensions: 294279
Number of successful extensions: 3770
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1655
Number of HSP's successfully gapped in prelim test: 181
Number of HSP's that attempted gapping in prelim test: 2015
Number of HSP's gapped (non-prelim): 1970
length of query: 656
length of database: 9,014,727
effective HSP length: 102
effective length of query: 554
effective length of database: 5,243,175
effective search space: 2904718950
effective search space used: 2904718950
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC144475.2


