

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144431.6 - phase: 0
(377 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
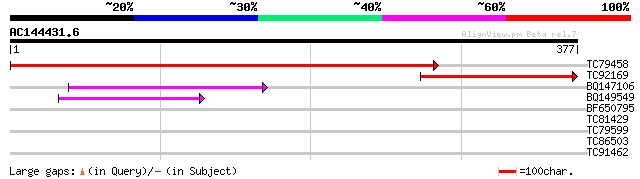
Sequences producing significant alignments: (bits) Value
TC79458 weakly similar to PIR|D85436|D85436 MAP3K-like protein k... 545 e-155
TC92169 similar to PIR|D85436|D85436 MAP3K-like protein kinase [... 202 2e-52
BQ147106 similar to GP|15451188|gb Unknown protein {Arabidopsis ... 43 2e-04
BQ149549 similar to PIR|B86270|B86 hypothetical protein AAF81295... 41 9e-04
BF650795 similar to GP|20161490|dbj beta-1 3 glucanase-like prot... 29 3.4
TC81429 similar to GP|9759108|dbj|BAB09677.1 gb|AAD32776.1~gene_... 28 5.8
TC79599 similar to GP|9757867|dbj|BAB08454.1 gb|AAD30228.1~gene_... 28 5.8
TC86503 similar to GP|15028127|gb|AAK76687.1 unknown protein {Ar... 28 7.6
TC91462 similar to GP|7769858|gb|AAF69536.1| F12M16.20 {Arabidop... 27 9.9
>TC79458 weakly similar to PIR|D85436|D85436 MAP3K-like protein kinase
[imported] - Arabidopsis thaliana, partial (35%)
Length = 1088
Score = 545 bits (1403), Expect = e-155
Identities = 285/285 (100%), Positives = 285/285 (100%)
Frame = +2
Query: 1 MPVLHVKHVLQMEIHDPDVAGYYLQIPLTK*RVYRLIGIHG*RHIIHLLWLVQDQPQVQL 60
MPVLHVKHVLQMEIHDPDVAGYYLQIPLTK*RVYRLIGIHG*RHIIHLLWLVQDQPQVQL
Sbjct: 233 MPVLHVKHVLQMEIHDPDVAGYYLQIPLTK*RVYRLIGIHG*RHIIHLLWLVQDQPQVQL 412
Query: 61 SLHL*TRMTLSLINSKMVFEDLC*ICMIFKTMYGCVILLEANVSTSVHSYLQ*TRLGTCV 120
SLHL*TRMTLSLINSKMVFEDLC*ICMIFKTMYGCVILLEANVSTSVHSYLQ*TRLGTCV
Sbjct: 413 SLHL*TRMTLSLINSKMVFEDLC*ICMIFKTMYGCVILLEANVSTSVHSYLQ*TRLGTCV 592
Query: 121 HFWTQIHLK*SPSLLRIMLELLQP*LRSSKLQGLTSTCSRLVGCPRTVAIGLLLMT*S*T 180
HFWTQIHLK*SPSLLRIMLELLQP*LRSSKLQGLTSTCSRLVGCPRTVAIGLLLMT*S*T
Sbjct: 593 HFWTQIHLK*SPSLLRIMLELLQP*LRSSKLQGLTSTCSRLVGCPRTVAIGLLLMT*S*T 772
Query: 181 INDLSHSPRDLPRKQLKAFLSRGNTS*KTNMEMKVCSRVLVRIEMNRHQ*TQNLGH*F** 240
INDLSHSPRDLPRKQLKAFLSRGNTS*KTNMEMKVCSRVLVRIEMNRHQ*TQNLGH*F**
Sbjct: 773 INDLSHSPRDLPRKQLKAFLSRGNTS*KTNMEMKVCSRVLVRIEMNRHQ*TQNLGH*F** 952
Query: 241 TFFILHRIVLRPVATIQLHYSAC*RLAMKRPATDGQISLLLIII* 285
TFFILHRIVLRPVATIQLHYSAC*RLAMKRPATDGQISLLLIII*
Sbjct: 953 TFFILHRIVLRPVATIQLHYSAC*RLAMKRPATDGQISLLLIII* 1087
>TC92169 similar to PIR|D85436|D85436 MAP3K-like protein kinase [imported] -
Arabidopsis thaliana, partial (6%)
Length = 505
Score = 202 bits (513), Expect = 2e-52
Identities = 104/104 (100%), Positives = 104/104 (100%)
Frame = +3
Query: 274 DGQISLLLIII*GVMVEEYHKLLMQQMVVSLVDVTALLIARQMVRLVHVMSLQFLHLHLL 333
DGQISLLLIII*GVMVEEYHKLLMQQMVVSLVDVTALLIARQMVRLVHVMSLQFLHLHLL
Sbjct: 3 DGQISLLLIII*GVMVEEYHKLLMQQMVVSLVDVTALLIARQMVRLVHVMSLQFLHLHLL 182
Query: 334 QKLHLMETRNLLIIVKQAMHIVVEQL**CNWLC*LWLPQLSWHG 377
QKLHLMETRNLLIIVKQAMHIVVEQL**CNWLC*LWLPQLSWHG
Sbjct: 183 QKLHLMETRNLLIIVKQAMHIVVEQL**CNWLC*LWLPQLSWHG 314
>BQ147106 similar to GP|15451188|gb Unknown protein {Arabidopsis thaliana},
partial (36%)
Length = 662
Score = 43.1 bits (100), Expect = 2e-04
Identities = 43/132 (32%), Positives = 59/132 (44%)
Frame = +3
Query: 40 HG*RHIIHLLWLVQDQPQVQLSLHL*TRMTLSLINSKMVFEDLC*ICMIFKTMYGCVILL 99
HG HIIH + + L + TL L N M ED C ICM + ++G VI
Sbjct: 267 HGS*HIIHSV*WMHPL*MEFKDLLFIIKKTLLLTN*GME*ED*CWICMTSRMIFGYVIHF 446
Query: 100 EANVSTSVHSYLQ*TRLGTCVHFWTQIHLK*SPSLLRIMLELLQP*LRSSKLQGLTSTCS 159
+ N +TS LQ*T HF +I + P L+ M + *L S + + CS
Sbjct: 447 KDNATTSPPFNLQ*TL*KKWKHF*LKIQWRLLPL*LKTMCVHQRR*LICS*MLVWINICS 626
Query: 160 RLVGCPRTVAIG 171
+ C + V IG
Sbjct: 627 PCLICQKMVKIG 662
>BQ149549 similar to PIR|B86270|B86 hypothetical protein AAF81295.1
[imported] - Arabidopsis thaliana, partial (29%)
Length = 635
Score = 40.8 bits (94), Expect = 9e-04
Identities = 36/97 (37%), Positives = 43/97 (44%)
Frame = +1
Query: 33 VYRLIGIHG*RHIIHLLWLVQDQPQVQLSLHL*TRMTLSLINSKMVFEDLC*ICMIFKTM 92
+Y I +H *+ I+HL QV L L T+ T S N M F C*I MI M
Sbjct: 334 LYHSINMHF*QRIMHLQLKGNHLTQVCLELLSPTKKTPSPNNLTMEFGH*C*IHMILMMM 513
Query: 93 YGCVILLEANVSTSVHSYLQ*TRLGTCVHFWTQIHLK 129
G VI + V TS H T HF QI +K
Sbjct: 514 CGYVIHFKVIVMTSPHLNQLLTH*RKLKHFCQQIQMK 624
>BF650795 similar to GP|20161490|dbj beta-1 3 glucanase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (30%)
Length = 668
Score = 28.9 bits (63), Expect = 3.4
Identities = 16/47 (34%), Positives = 28/47 (59%)
Frame = +1
Query: 287 VMVEEYHKLLMQQMVVSLVDVTALLIARQMVRLVHVMSLQFLHLHLL 333
+ V+ +HKL+ + LVD+ AL++ Q LV +++Q L + LL
Sbjct: 271 LQVQVHHKLVYKLHWTMLVDMVALIV--QQFNLVEAVTIQILFMILL 405
>TC81429 similar to GP|9759108|dbj|BAB09677.1
gb|AAD32776.1~gene_id:MJJ3.25~similar to unknown protein
{Arabidopsis thaliana}, partial (35%)
Length = 945
Score = 28.1 bits (61), Expect = 5.8
Identities = 26/76 (34%), Positives = 42/76 (55%), Gaps = 4/76 (5%)
Frame = +2
Query: 277 ISLLLIII*GVM--VEEYHKLLMQQMVVS--LVDVTALLIARQMVRLVHVMSLQFLHLHL 332
I + +III VM V+E + LMQ MVV L+ + +++ M+ L+ L FL L+L
Sbjct: 455 IFIAIIIIMEVM*VVKEVYVSLMQTMVVQAILISIQQMVVQAMMITLL----LVFLILNL 622
Query: 333 LQKLHLMETRNLLIIV 348
+ + L E LI++
Sbjct: 623 VVFMILEEFLLFLIVL 670
>TC79599 similar to GP|9757867|dbj|BAB08454.1
gb|AAD30228.1~gene_id:K9I9.2~similar to unknown protein
{Arabidopsis thaliana}, partial (23%)
Length = 1530
Score = 28.1 bits (61), Expect = 5.8
Identities = 17/54 (31%), Positives = 26/54 (47%)
Frame = +2
Query: 292 YHKLLMQQMVVSLVDVTALLIARQMVRLVHVMSLQFLHLHLLQKLHLMETRNLL 345
+H L +Q ++ +V + L I + VRL LHL+ KLHL +L
Sbjct: 866 HHSLFLQLTLLKMVSLVHLAIQ*KTVRL---------RLHLIHKLHLFRNFGVL 1000
>TC86503 similar to GP|15028127|gb|AAK76687.1 unknown protein {Arabidopsis
thaliana}, partial (54%)
Length = 1915
Score = 27.7 bits (60), Expect = 7.6
Identities = 9/9 (100%), Positives = 9/9 (100%)
Frame = -2
Query: 369 WLPQLSWHG 377
WLPQLSWHG
Sbjct: 1119 WLPQLSWHG 1093
>TC91462 similar to GP|7769858|gb|AAF69536.1| F12M16.20 {Arabidopsis
thaliana}, partial (21%)
Length = 1134
Score = 27.3 bits (59), Expect = 9.9
Identities = 16/45 (35%), Positives = 27/45 (59%)
Frame = -1
Query: 304 LVDVTALLIARQMVRLVHVMSLQFLHLHLLQKLHLMETRNLLIIV 348
+++ LL +V+LVH LQ LH HL ++ + RNLL+++
Sbjct: 156 ILNYILLLHEISLVQLVHEKELQQLHCHL--EVLSVRPRNLLLLL 28
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.358 0.158 0.540
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,912,724
Number of Sequences: 36976
Number of extensions: 172901
Number of successful extensions: 2026
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 1490
Number of HSP's successfully gapped in prelim test: 68
Number of HSP's that attempted gapping in prelim test: 501
Number of HSP's gapped (non-prelim): 1603
length of query: 377
length of database: 9,014,727
effective HSP length: 98
effective length of query: 279
effective length of database: 5,391,079
effective search space: 1504111041
effective search space used: 1504111041
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144431.6


