

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144345.11 - phase: 0
(339 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
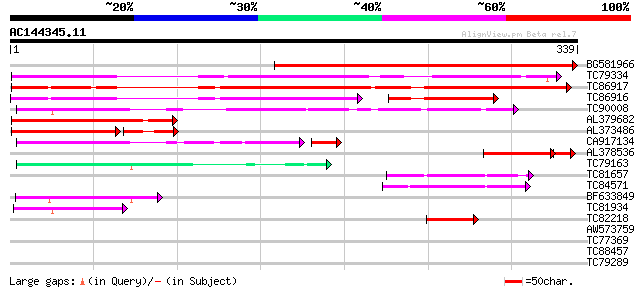
Score E
Sequences producing significant alignments: (bits) Value
BG581966 similar to GP|3776080|emb MtN9 {Medicago truncatula}, p... 369 e-103
TC79334 MtN9 286 6e-78
TC86917 similar to GP|3776080|emb|CAA77093.1 MtN9 {Medicago trun... 284 4e-77
TC86916 similar to GP|3776080|emb|CAA77093.1 MtN9 {Medicago trun... 159 8e-56
TC90008 weakly similar to GP|4249376|gb|AAD14473.1| Strong simil... 137 5e-33
AL379682 weakly similar to SP|P29136|MEP1_ Metalloendoproteinase... 121 4e-28
AL373486 weakly similar to GP|16901508|gb| matrix metalloprotein... 88 2e-23
CA917134 weakly similar to PIR|E96724|E96 hypothetical protein F... 80 7e-19
AL378536 homologue to GP|3776080|emb| MtN9 {Medicago truncatula}... 76 8e-15
TC79163 weakly similar to PIR|E96724|E96724 hypothetical protein... 72 4e-13
TC81657 weakly similar to PIR|T51957|T51957 metalloproteinase (E... 62 3e-10
TC84571 weakly similar to SP|P29136|MEP1_SOYBN Metalloendoprotei... 60 1e-09
BF633849 weakly similar to PIR|E71433|E714 probable metalloprote... 55 3e-08
TC81934 weakly similar to PIR|E71433|E71433 probable metalloprot... 53 2e-07
TC82218 similar to GP|7159629|emb|CAB76364.1 matrix metalloprote... 50 1e-06
AW573759 homologue to PIR|T09387|T09 NMS32/34 protein - alfalfa ... 28 6.6
TC77369 similar to GP|14532824|gb|AAK64094.1 unknown protein {Ar... 28 6.6
TC88457 homologue to PIR|T06493|T06493 1 4-alpha-glucan branchin... 27 8.6
TC79289 weakly similar to GP|7298980|gb|AAF54183.1| CG10903 gene... 27 8.6
>BG581966 similar to GP|3776080|emb MtN9 {Medicago truncatula}, partial (53%)
Length = 683
Score = 369 bits (948), Expect = e-103
Identities = 181/181 (100%), Positives = 181/181 (100%)
Frame = +1
Query: 159 LDMIEGFRNAFTRWSQTTRVLNFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRN 218
LDMIEGFRNAFTRWSQTTRVLNFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRN
Sbjct: 1 LDMIEGFRNAFTRWSQTTRVLNFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRN 180
Query: 219 LDSNVKSGMIRLERSKFWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIM 278
LDSNVKSGMIRLERSKFWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIM
Sbjct: 181 LDSNVKSGMIRLERSKFWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIM 360
Query: 279 YPLIDPLQEKKVQITDSDNQAIQQLYTNTAKANPNSDHSGCFKLFESSMFLLAFCPLVLL 338
YPLIDPLQEKKVQITDSDNQAIQQLYTNTAKANPNSDHSGCFKLFESSMFLLAFCPLVLL
Sbjct: 361 YPLIDPLQEKKVQITDSDNQAIQQLYTNTAKANPNSDHSGCFKLFESSMFLLAFCPLVLL 540
Query: 339 L 339
L
Sbjct: 541 L 543
>TC79334 MtN9
Length = 1241
Score = 286 bits (733), Expect = 6e-78
Identities = 177/335 (52%), Positives = 202/335 (59%), Gaps = 6/335 (1%)
Frame = +1
Query: 2 KIQGLSQIKKHLSTFGYFRQ-FLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQK 60
+IQGLS+IK+HL F Y + +L+ FDD LD +TISAIK YQQFFNLQVTG+L+TETLQ+
Sbjct: 103 EIQGLSKIKQHLYHFKYLQGLYLVGFDDYLDNKTISAIKAYQQFFNLQVTGHLDTETLQQ 282
Query: 61 ISLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMR 120
I LP RCG+PD+
Sbjct: 283 IMLP------------------------------------------------RCGVPDIN 318
Query: 121 YEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVLN 180
+ D G + +GNKWFPKGTK+LTYGFLP K S+D + FRNAFTRWSQTTRVL
Sbjct: 319 PDINPDFG--FARAQGNKWFPKGTKELTYGFLPESKISIDKVNVFRNAFTRWSQTTRVLK 492
Query: 181 FSETTSYDDADIKIGFYHI-YNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFWVSP 239
FSE TSYDDADIKIGFY+I YN+ V DVVV D FI N++S IRLE SK W
Sbjct: 493 FSEATSYDDADIKIGFYNISYNSKEVIDVVVSDFFI------NLRSFTIRLEASKVW--- 645
Query: 240 TTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQA 299
DLETV MHQIGHLLGLDHSSD ESIMYP I PL +KKVQIT SDNQA
Sbjct: 646 -------------DLETVAMHQIGHLLGLDHSSDVESIMYPTIVPLHQKKVQITVSDNQA 786
Query: 300 IQQLYTNTAKANPNSDHSGCF----KLFESSMFLL 330
IQQLYT + N + D G F FESS LL
Sbjct: 787 IQQLYTK--QTNQDRDELGFFDYSGDFFESSSGLL 885
>TC86917 similar to GP|3776080|emb|CAA77093.1 MtN9 {Medicago truncatula},
partial (59%)
Length = 1294
Score = 284 bits (726), Expect = 4e-77
Identities = 167/337 (49%), Positives = 210/337 (61%), Gaps = 2/337 (0%)
Frame = +1
Query: 2 KIQGLSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKI 61
K +G+ QIK++LS FGY Q F++ LD+ET+ A+KTYQ++FN+Q + +E LQ I
Sbjct: 334 KYKGVDQIKQYLSDFGYLEQSG-PFNNTLDQETVLALKTYQRYFNIQQ--DTLSEILQHI 504
Query: 62 SLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRY 121
+LP RCG+PD
Sbjct: 505 ALP------------------------------------------------RCGVPDRIL 540
Query: 122 EYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVLNF 181
+Y ++ +D+SFPKGN+WFPKGTK LTYGF P K LDM FR A T+WS TTRVLNF
Sbjct: 541 KY--NLTNDISFPKGNQWFPKGTKNLTYGFDPRNKIPLDMTNVFRTALTQWSNTTRVLNF 714
Query: 182 SETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRNLDS-NVKSGMIRLERSKFWVSPT 240
+ET SYDDA+IKIGFY+I ++D ++DV VG +FI LDS NVKSG I L+ +K+W PT
Sbjct: 715 TETKSYDDANIKIGFYNITDDDGINDVAVGFTFIV--LDSTNVKSGFITLDATKYWALPT 888
Query: 241 TTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQAI 300
E + FDLET MHQIGHLLGL+HSSD +SIMYP I P +K VQITDSDN AI
Sbjct: 889 -------EHRGFDLETAAMHQIGHLLGLEHSSDNKSIMYPTILPSHQKNVQITDSDNLAI 1047
Query: 301 QQLYTNTAKANPNS-DHSGCFKLFESSMFLLAFCPLV 336
Q+LY+++ KAN NS D SGCFKLF SS LL +V
Sbjct: 1048QKLYSSSTKANANSDDSSGCFKLFGSSSSLLISLSIV 1158
>TC86916 similar to GP|3776080|emb|CAA77093.1 MtN9 {Medicago truncatula},
partial (39%)
Length = 954
Score = 159 bits (403), Expect(2) = 8e-56
Identities = 90/211 (42%), Positives = 119/211 (55%)
Frame = +2
Query: 1 MKIQGLSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQK 60
+K +GL QIK++L FGY Q F++ LD+ET+ A+KTYQ++FN+ + + LQ
Sbjct: 254 LKYKGLDQIKQYLQNFGYLEQSG-PFNNTLDQETVLALKTYQRYFNIYAGQDSLRKILQH 430
Query: 61 ISLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMR 120
I+LP RCG+PDM
Sbjct: 431 IALP------------------------------------------------RCGVPDMN 466
Query: 121 YEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVLN 180
+ Y D +D+S+PKGN+WFPKGTK LTYGF P + L++ FR A TRWSQTTRVLN
Sbjct: 467 FTY--DSTNDISYPKGNQWFPKGTKNLTYGFAPKNEIPLNVTNVFRKALTRWSQTTRVLN 640
Query: 181 FSETTSYDDADIKIGFYHIYNNDIVDDVVVG 211
F+ETTSYDDADIKI F ++ +D + DVVVG
Sbjct: 641 FTETTSYDDADIKIVFNNMTYDDGIYDVVVG 733
Score = 75.5 bits (184), Expect(2) = 8e-56
Identities = 40/66 (60%), Positives = 46/66 (69%)
Frame = +1
Query: 227 MIRLERSKFWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQ 286
+I L+ +K WV PT E E DLET MHQIGHLLGL+HSSD +SIMYP I P Q
Sbjct: 778 LISLDITKHWVFPT-------EDGELDLETAAMHQIGHLLGLEHSSDSKSIMYPTILPSQ 936
Query: 287 EKKVQI 292
+KKVQI
Sbjct: 937 QKKVQI 954
>TC90008 weakly similar to GP|4249376|gb|AAD14473.1| Strong similarity to
GP|2829864 F3I6.6 zinc metalloproteinase homolog from
Arabidopsis thaliana BAC, partial (21%)
Length = 1173
Score = 137 bits (346), Expect = 5e-33
Identities = 99/303 (32%), Positives = 147/303 (47%), Gaps = 3/303 (0%)
Frame = +2
Query: 5 GLSQIKKHLSTFGYFRQFLL--KFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKIS 62
G+S +K++L FGY L F D E SAI +Q FNL TG L+ + + IS
Sbjct: 269 GISDLKEYLQFFGYLNSSTLHSNFTDAFTENLQSAIIEFQTNFNLNTTGQLDQDIYKIIS 448
Query: 63 LPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYE 122
PRCG+PD+ GT + F+
Sbjct: 449 KPRCGVPDII---------------------NGTTTMKNNFI------------------ 511
Query: 123 YGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVLNFS 182
+ + P W + L Y F P + ++ F++AF RWS T LNF
Sbjct: 512 -------NKTMPFKPWWRNVENRSLAYAFHPENNVTDNVKSLFQDAFNRWSNATE-LNFI 667
Query: 183 ETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFWVSPTTT 242
ET S++D+DI+I F + VG S+I+ S+V G + L+ + WV P+
Sbjct: 668 ETMSFNDSDIRIAFLTLDGKG----GTVGGSYIN----SSVNVGSVYLDADEQWVLPSEN 823
Query: 243 YFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDN-QAIQ 301
++ + DLE+VVMHQ+GHLLGL HSS +E+IMYP++ LQEKK+++ + D+ Q IQ
Sbjct: 824 VVEE---DDVDLESVVMHQVGHLLGLGHSSVEEAIMYPIV--LQEKKIELVNVDDLQRIQ 988
Query: 302 QLY 304
++Y
Sbjct: 989 EIY 997
>AL379682 weakly similar to SP|P29136|MEP1_ Metalloendoproteinase 1 precursor
(EC 3.4.24.-) (SMEP1). [Soybean] {Glycine max}, partial
(15%)
Length = 386
Score = 121 bits (303), Expect = 4e-28
Identities = 62/100 (62%), Positives = 78/100 (78%), Gaps = 1/100 (1%)
Frame = +3
Query: 2 KIQGLSQIKKHLSTFGYFRQ-FLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQK 60
+IQGLS+IK+HL F Y + +L+ FDD LD +TISAIK YQQFFNLQVTG+L+TETLQ+
Sbjct: 93 EIQGLSKIKQHLYHFKYLQGLYLVGFDDYLDNKTISAIKAYQQFFNLQVTGHLDTETLQQ 272
Query: 61 ISLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLT 100
I LPRCG+PD+ + D G + +GNKWFPKGTK+LT
Sbjct: 273 IMLPRCGVPDINPDINPDFG--FARAQGNKWFPKGTKELT 386
>AL373486 weakly similar to GP|16901508|gb| matrix metalloproteinase MMP2
{Glycine max}, partial (7%)
Length = 409
Score = 88.2 bits (217), Expect(2) = 2e-23
Identities = 45/66 (68%), Positives = 55/66 (83%), Gaps = 1/66 (1%)
Frame = +2
Query: 2 KIQGLSQIKKHLSTFGYFRQ-FLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQK 60
+IQGLS+IK+HL F Y + +L+ FDD LD +TISAIK YQQFFNLQVTG+L+TETLQ+
Sbjct: 110 EIQGLSKIKQHLYHFKYLQGLYLVGFDDYLDNKTISAIKGYQQFFNLQVTGHLDTETLQQ 289
Query: 61 ISLPRC 66
I LPRC
Sbjct: 290 IMLPRC 307
Score = 38.1 bits (87), Expect = 0.005
Identities = 17/33 (51%), Positives = 22/33 (66%)
Frame = +3
Query: 117 PDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTY 149
PD+ ++GF +GNKWFPKGTK+LTY
Sbjct: 327 PDINPDFGFGRA------QGNKWFPKGTKELTY 407
Score = 38.1 bits (87), Expect(2) = 2e-23
Identities = 17/33 (51%), Positives = 22/33 (66%)
Frame = +3
Query: 69 PDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTY 101
PD+ ++GF +GNKWFPKGTK+LTY
Sbjct: 327 PDINPDFGFGRA------QGNKWFPKGTKELTY 407
>CA917134 weakly similar to PIR|E96724|E96 hypothetical protein F20P5.11
[imported] - Arabidopsis thaliana, partial (26%)
Length = 723
Score = 80.5 bits (197), Expect(2) = 7e-19
Identities = 56/173 (32%), Positives = 78/173 (44%), Gaps = 1/173 (0%)
Frame = +1
Query: 5 GLSQIKKHLSTFGYF-RQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKISL 63
GLS +K + FGY R F D D++ AIKTYQ+ FNL VTG L+ TL+++ L
Sbjct: 136 GLSNLKNYFQHFGYIPRSPKSNFSDDFDDDLQEAIKTYQKNFNLNVTGELDDMTLRQVML 315
Query: 64 PRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYEY 123
PRCG+ D+ GT + GK +R+
Sbjct: 316 PRCGVADII---------------------NGTTTMN----AGKDTETTSNSDSKLRFH- 417
Query: 124 GFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTT 176
V FP +W P+G ++LTY F PG + + + F AF RWS+ T
Sbjct: 418 --TVSHFTVFPGQPRW-PEGKQELTYAFFPGNELTETVKSVFATAFARWSEVT 567
Score = 30.8 bits (68), Expect(2) = 7e-19
Identities = 13/18 (72%), Positives = 14/18 (77%)
Frame = +3
Query: 181 FSETTSYDDADIKIGFYH 198
F+ETT Y ADIKIGF H
Sbjct: 576 FTETTLYSGADIKIGFLH 629
>AL378536 homologue to GP|3776080|emb| MtN9 {Medicago truncatula}, partial
(20%)
Length = 349
Score = 76.3 bits (186), Expect(2) = 8e-15
Identities = 35/43 (81%), Positives = 40/43 (92%)
Frame = +1
Query: 284 PLQEKKVQITDSDNQAIQQLYTNTAKANPNSDHSGCFKLFESS 326
PLQ++KVQIT SDN AIQQLYTN+ KANP+SDHSGCFKLFES+
Sbjct: 4 PLQQRKVQITVSDNLAIQQLYTNSVKANPDSDHSGCFKLFEST 132
Score = 21.2 bits (43), Expect(2) = 8e-15
Identities = 8/13 (61%), Positives = 10/13 (76%)
Frame = +2
Query: 326 SMFLLAFCPLVLL 338
S+ L+ CPLVLL
Sbjct: 125 SLLLIPLCPLVLL 163
>TC79163 weakly similar to PIR|E96724|E96724 hypothetical protein F20P5.11
[imported] - Arabidopsis thaliana, partial (32%)
Length = 762
Score = 71.6 bits (174), Expect = 4e-13
Identities = 57/202 (28%), Positives = 81/202 (39%), Gaps = 14/202 (6%)
Frame = +2
Query: 5 GLSQIKKHLSTFGYFRQFL-LKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKISL 63
GLS++K + + FGY F D DE SA++TYQ+ FNL +TG L+ T+ I
Sbjct: 302 GLSKLKNYFNHFGYIPNGPPSNFTDDFDEALESAVRTYQKNFNLNITGELDDATMNYIVK 481
Query: 64 PRCGIPDM-------------RYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFS 110
PRCG+ D+ F + SF G +P+GT+ LTY F P +
Sbjct: 482 PRCGVADIINGTTSMNSGKFNSSSTNFHTVAHYSFFPGQPRWPEGTQTLTYAFDPSENLD 661
Query: 111 LLRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFT 170
TK++ F NAF
Sbjct: 662 -------------------------------DATKQV-----------------FANAFN 697
Query: 171 RWSQTTRVLNFSETTSYDDADI 192
+WS+ T + F+E TSY +DI
Sbjct: 698 QWSKVT-TITFTEATSYSSSDI 760
>TC81657 weakly similar to PIR|T51957|T51957 metalloproteinase (EC 3.4.24.-)
[imported] - Arabidopsis thaliana, partial (24%)
Length = 537
Score = 62.0 bits (149), Expect = 3e-10
Identities = 37/88 (42%), Positives = 52/88 (59%)
Frame = +2
Query: 226 GMIRLERSKFWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPL 285
G + L++++ WV DLE+VV+H+IGHLLGL HSS +E+IMYP I
Sbjct: 8 GRLHLDKAEDWVVNGDVTESSLS-NAVDLESVVVHEIGHLLGLGHSSIEEAIMYPTISS- 181
Query: 286 QEKKVQITDSDNQAIQQLYTNTAKANPN 313
+ KKV++ D + IQ LY +NPN
Sbjct: 182 RTKKVELESDDIEGIQMLY----GSNPN 253
>TC84571 weakly similar to SP|P29136|MEP1_SOYBN Metalloendoproteinase 1
precursor (EC 3.4.24.-) (SMEP1). [Soybean] {Glycine
max}, partial (28%)
Length = 506
Score = 60.1 bits (144), Expect = 1e-09
Identities = 35/88 (39%), Positives = 51/88 (57%)
Frame = +2
Query: 224 KSGMIRLERSKFWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLID 283
++G L+ ++ WV + K DLE+V +H+IGHLLGL HSS++E+IM+P I
Sbjct: 11 RNGRFHLDAAEDWVV-SGDVSKSSLPTAVDLESVAVHEIGHLLGLGHSSEEEAIMFPTIS 187
Query: 284 PLQEKKVQITDSDNQAIQQLYTNTAKAN 311
+ KKV + D D + IQ LY N
Sbjct: 188 S-RMKKVVLADDDVRGIQYLYGTNPSFN 268
>BF633849 weakly similar to PIR|E71433|E714 probable metalloproteinase (EC
3.4.24.-) - Arabidopsis thaliana, partial (5%)
Length = 544
Score = 55.5 bits (132), Expect = 3e-08
Identities = 31/98 (31%), Positives = 53/98 (53%), Gaps = 10/98 (10%)
Frame = +2
Query: 4 QGLSQIKKHLSTFGYFRQF-----LLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETL 58
+G+ ++K +L FGY +D DE A+K +Q + +L VTG ++T T+
Sbjct: 185 KGIGELKNYLKRFGYLNVQHDATPSTTLNDHFDENLEFALKDFQTYHHLHVTGRVDTTTI 364
Query: 59 QKISLPRCGIPDM----RYEYGFDVGSDVSFPKGN-KW 91
+ +SLPRCG+PD+ + G + S +F + + KW
Sbjct: 365 KILSLPRCGVPDLPKHSHKQNGLQMSSSYAFFQDSPKW 478
>TC81934 weakly similar to PIR|E71433|E71433 probable metalloproteinase (EC
3.4.24.-) - Arabidopsis thaliana, partial (24%)
Length = 619
Score = 52.8 bits (125), Expect = 2e-07
Identities = 25/70 (35%), Positives = 39/70 (55%), Gaps = 2/70 (2%)
Frame = +1
Query: 3 IQGLSQIKKHLSTFGYFRQFLL--KFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQK 60
+ G++++KK+ + FGY L F D D SA+ YQ+ L +TG L++ T+
Sbjct: 211 VSGMAELKKYFNRFGYLSPPLTTANFTDTFDSSFQSAVVLYQKNLGLPITGKLDSNTIST 390
Query: 61 ISLPRCGIPD 70
I PRCG+ D
Sbjct: 391 IVSPRCGVSD 420
>TC82218 similar to GP|7159629|emb|CAB76364.1 matrix metalloproteinase
{Cucumis sativus}, partial (19%)
Length = 688
Score = 50.1 bits (118), Expect = 1e-06
Identities = 20/31 (64%), Positives = 27/31 (86%)
Frame = +1
Query: 250 QEFDLETVVMHQIGHLLGLDHSSDKESIMYP 280
++FDLETV +H++GHLLGL HS+D+ S MYP
Sbjct: 253 KQFDLETVALHEMGHLLGLAHSTDQNSAMYP 345
>AW573759 homologue to PIR|T09387|T09 NMS32/34 protein - alfalfa (fragment),
partial (77%)
Length = 618
Score = 27.7 bits (60), Expect = 6.6
Identities = 16/64 (25%), Positives = 31/64 (48%), Gaps = 6/64 (9%)
Frame = -3
Query: 280 PLIDPLQEKKVQITDSDNQAIQQLYTNTAKANPNSDHSGCFKLF------ESSMFLLAFC 333
PL++ L ++ T S + ++ L+ ++ K N +S CF+ F FL+ F
Sbjct: 550 PLLEALFSLSLKETLSSSSILETLFPSSLKKNLSSSSHKCFRNFRLLGIVSEHFFLIRFA 371
Query: 334 PLVL 337
+V+
Sbjct: 370 RIVI 359
>TC77369 similar to GP|14532824|gb|AAK64094.1 unknown protein {Arabidopsis
thaliana}, complete
Length = 1048
Score = 27.7 bits (60), Expect = 6.6
Identities = 11/29 (37%), Positives = 16/29 (54%)
Frame = +2
Query: 304 YTNTAKANPNSDHSGCFKLFESSMFLLAF 332
+TN K+NP+ +S F LF +F F
Sbjct: 128 FTNNPKSNPSKSNSSSFNLFYLDLFYSTF 214
>TC88457 homologue to PIR|T06493|T06493 1 4-alpha-glucan branching enzyme
(EC 2.4.1.18) I - garden pea, partial (27%)
Length = 1201
Score = 27.3 bits (59), Expect = 8.6
Identities = 32/118 (27%), Positives = 50/118 (42%), Gaps = 5/118 (4%)
Frame = +3
Query: 83 VSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYEYGFDVGSDVSFPK--GNKWF 140
+ FP+G++ P GT +PG S +C FD+G D + + G + F
Sbjct: 195 IDFPRGDQHLPNGT------VVPGNNNSYDKC-------RRRFDLG-DAEYLRYHGMQEF 332
Query: 141 PKGTKKL--TYGFLPGK-KFSLDMIEGFRNAFTRWSQTTRVLNFSETTSYDDADIKIG 195
+ + L YGF+ + ++ EG R V NF T SY +D K+G
Sbjct: 333 DRAMQHLEERYGFMISEHQYISRKNEGDRVIIFERDNLVFVFNFHWTNSY--SDYKVG 500
>TC79289 weakly similar to GP|7298980|gb|AAF54183.1| CG10903 gene product
{Drosophila melanogaster}, partial (83%)
Length = 1098
Score = 27.3 bits (59), Expect = 8.6
Identities = 12/48 (25%), Positives = 25/48 (52%), Gaps = 2/48 (4%)
Frame = +2
Query: 46 NLQVTGNLNTETLQKISLPRCGIPDMRYEYGFDVG--SDVSFPKGNKW 91
N+Q+ ++ L+ ++LP+ G+P + + G G +V G+ W
Sbjct: 101 NIQIQTSMTERALELLNLPKDGVPKLLLDIGCGSGLSGEVITESGHHW 244
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,488,953
Number of Sequences: 36976
Number of extensions: 136710
Number of successful extensions: 684
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 581
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 659
length of query: 339
length of database: 9,014,727
effective HSP length: 97
effective length of query: 242
effective length of database: 5,428,055
effective search space: 1313589310
effective search space used: 1313589310
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144345.11


