

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC143341.5 + phase: 0
(485 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
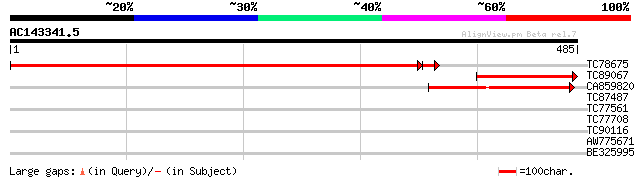
Score E
Sequences producing significant alignments: (bits) Value
TC78675 similar to GP|9858775|gb|AAG01122.1| BAC19.7 {Lycopersic... 711 0.0
TC89067 similar to GP|21537345|gb|AAM61686.1 adenylosuccinate sy... 174 7e-44
CA859820 weakly similar to GP|12836392|db adenylosuccinate synth... 144 1e-34
TC87487 similar to SP|P52780|SYQ_LUPLU Glutaminyl-tRNA synthetas... 30 1.6
TC77561 similar to GP|14423502|gb|AAK62433.1 Unknown protein {Ar... 30 1.6
TC77708 weakly similar to GP|18266654|gb|AAL67600.1 unknown prot... 30 1.6
TC90116 homologue to SP|O50044|KDSA_PEA 2-dehydro-3-deoxyphospho... 30 1.6
AW775671 similar to GP|17065312|gb Unknown protein {Arabidopsis ... 30 2.7
BE325995 similar to GP|15292869|gb| unknown protein {Arabidopsis... 29 4.6
>TC78675 similar to GP|9858775|gb|AAG01122.1| BAC19.7 {Lycopersicon
esculentum}, partial (54%)
Length = 1162
Score = 711 bits (1834), Expect(2) = 0.0
Identities = 352/353 (99%), Positives = 353/353 (99%)
Frame = +2
Query: 1 MNISSLRLDSHAICTHQPQRPFAFHNRFPRNVVVCAAKPVAPPPTKLAAADSSLRRIESL 60
MNISSLRLDSHAICTHQPQRPFAFHNRFPRNVVVCAAKPVAPPPTKLAAADSSLRRIESL
Sbjct: 59 MNISSLRLDSHAICTHQPQRPFAFHNRFPRNVVVCAAKPVAPPPTKLAAADSSLRRIESL 238
Query: 61 SQVSGVLGCQWGDEGKGKLVDILAQHFQIVARCQGGANAGHTIYNAEGKKFALHLVPSGI 120
SQVSGVLGCQWGDEGKGKLVDILAQHFQIVARCQGGANAGHTIYNAEGKKFALHLVPSGI
Sbjct: 239 SQVSGVLGCQWGDEGKGKLVDILAQHFQIVARCQGGANAGHTIYNAEGKKFALHLVPSGI 418
Query: 121 LNEDTLCVIGNGVVVHLPGLFQEIDNLESNGVSCKGRILISDRAHLLFDFHQTVDGLREA 180
LNEDTLCVIGNGVVVHLPGLFQEIDNLESNGVSCKGRILISDRAHLLFDFHQTVDGLREA
Sbjct: 419 LNEDTLCVIGNGVVVHLPGLFQEIDNLESNGVSCKGRILISDRAHLLFDFHQTVDGLREA 598
Query: 181 ELSKSFIGTTKRGIGPCYSSKANRNGIRVGDLRYMETLPQKLDLLLSDAALRFKDFKYGP 240
ELSKSFIGTTKRGIGPCYSSKANRNGIRVGDLRYMETLPQKLDLLLSDAALRFKDFKYGP
Sbjct: 599 ELSKSFIGTTKRGIGPCYSSKANRNGIRVGDLRYMETLPQKLDLLLSDAALRFKDFKYGP 778
Query: 241 DVLREEVEKYKRYAERLEPYIADTVHVMNEAITQKKKVLVEGGQATMLDIDFGTYPFVTS 300
DVLREEVEKYKRYAERLEPYIADTVHVMNEAITQKKKVLVEGGQATMLDIDFGTYPFVTS
Sbjct: 779 DVLREEVEKYKRYAERLEPYIADTVHVMNEAITQKKKVLVEGGQATMLDIDFGTYPFVTS 958
Query: 301 SSPSAGGICTGLGIAPRVIGDLVGVVKAYTTRVGSGPFPTEILGPGGDLLRFA 353
SSPSAGGICTGLGIAPRVIGDLVGVVKAYTTRVGSGPFPTEILGPGGDLL+FA
Sbjct: 959 SSPSAGGICTGLGIAPRVIGDLVGVVKAYTTRVGSGPFPTEILGPGGDLLKFA 1117
Score = 35.4 bits (80), Expect(2) = 0.0
Identities = 14/14 (100%), Positives = 14/14 (100%)
Frame = +3
Query: 354 GQEFGTTTGRPRRC 367
GQEFGTTTGRPRRC
Sbjct: 1119 GQEFGTTTGRPRRC 1160
>TC89067 similar to GP|21537345|gb|AAM61686.1 adenylosuccinate synthetase
{Arabidopsis thaliana}, partial (17%)
Length = 645
Score = 174 bits (441), Expect = 7e-44
Identities = 85/86 (98%), Positives = 86/86 (99%)
Frame = +2
Query: 400 EIQLGVSYKHADGTPVQSFPSDLRLLEQLKVEYESLPGWKTDISSIRNYSDLPRAAQLYV 459
+IQLGVSYKHADGTPVQSFPSDLRLLEQLKVEYESLPGWKTDISSIRNYSDLPRAAQLYV
Sbjct: 5 QIQLGVSYKHADGTPVQSFPSDLRLLEQLKVEYESLPGWKTDISSIRNYSDLPRAAQLYV 184
Query: 460 ERIEELVGVPIHYIGVGPGRDALIFK 485
ERIEELVGVPIHYIGVGPGRDALIFK
Sbjct: 185 ERIEELVGVPIHYIGVGPGRDALIFK 262
>CA859820 weakly similar to GP|12836392|db adenylosuccinate synthetase 2 non
muscle~data source:MGD source key:MGI:87948
evidence:ISS~, partial (27%)
Length = 441
Score = 144 bits (362), Expect = 1e-34
Identities = 70/125 (56%), Positives = 93/125 (74%)
Frame = +3
Query: 359 TTTGRPRRCGWLDIVALSYSCQINGFSSLNLTKLDVLSDLDEIQLGVSYKHADGTPVQSF 418
TTT R RRCGWLD+V L YS ING++SLNLTKLDVL L E+++ +Y DG P++SF
Sbjct: 3 TTTRRLRRCGWLDLVVLKYSTMINGYTSLNLTKLDVLDQLPELKVATAYL-LDGKPLESF 179
Query: 419 PSDLRLLEQLKVEYESLPGWKTDISSIRNYSDLPRAAQLYVERIEELVGVPIHYIGVGPG 478
P+DL +LE++ V+Y++LPGWK DIS R SDLP A+ YV+ IE+ + VPI +IGVG
Sbjct: 180 PADLDILEKVTVQYKTLPGWKIDISKCRRLSDLPENARNYVKMIEDYLNVPIEWIGVGVR 359
Query: 479 RDALI 483
R+ +I
Sbjct: 360 REDMI 374
>TC87487 similar to SP|P52780|SYQ_LUPLU Glutaminyl-tRNA synthetase (EC
6.1.1.18) (Glutamine--tRNA ligase) (GLNRS). [Yellow
lupine], partial (97%)
Length = 2844
Score = 30.4 bits (67), Expect = 1.6
Identities = 18/61 (29%), Positives = 33/61 (53%)
Frame = +1
Query: 422 LRLLEQLKVEYESLPGWKTDISSIRNYSDLPRAAQLYVERIEELVGVPIHYIGVGPGRDA 481
L+ +QL+V+ + P + D SS + +P + LY+ER + + Y G+ PG+ A
Sbjct: 1876 LKPTQQLRVDAKKWPDAQADDSSA--FYKIPFSNVLYIERTDFRMKDSKDYYGLAPGKSA 2049
Query: 482 L 482
+
Sbjct: 2050 I 2052
>TC77561 similar to GP|14423502|gb|AAK62433.1 Unknown protein {Arabidopsis
thaliana}, partial (73%)
Length = 1276
Score = 30.4 bits (67), Expect = 1.6
Identities = 16/54 (29%), Positives = 24/54 (43%)
Frame = +1
Query: 288 LDIDFGTYPFVTSSSPSAGGICTGLGIAPRVIGDLVGVVKAYTTRVGSGPFPTE 341
LD+D T+ +T P + P ++ DLV ++ V SG F TE
Sbjct: 754 LDVDIATHLTITDDGPDRSLLYVETADRPGLLVDLVKIITDINIAVESGEFDTE 915
>TC77708 weakly similar to GP|18266654|gb|AAL67600.1 unknown protein {Oryza
sativa}, partial (30%)
Length = 1593
Score = 30.4 bits (67), Expect = 1.6
Identities = 25/122 (20%), Positives = 47/122 (38%), Gaps = 1/122 (0%)
Frame = +1
Query: 5 SLRLDSHAICTHQPQRPFAFHNRFPRNVVVCAAKPVAPPPTKLAAADSSL-RRIESLSQV 63
SL +S+ H P + + + + C + P++P + L + L RR++S
Sbjct: 388 SLHPESYQALKHLPPTASSLQHHHHNHKLSCTSSPISPSISDLERQNKGLIRRVQS---- 555
Query: 64 SGVLGCQWGDEGKGKLVDILAQHFQIVARCQGGANAGHTIYNAEGKKFALHLVPSGILNE 123
+G L D+ C ++ Y+ + FAL +PS L++
Sbjct: 556 ------------EGNLEDLA-----YATNCNNNMDSSSKRYSVRQRGFALETIPSFSLSK 684
Query: 124 DT 125
T
Sbjct: 685 QT 690
>TC90116 homologue to SP|O50044|KDSA_PEA 2-dehydro-3-deoxyphosphooctonate
aldolase (EC 4.1.2.16), complete
Length = 1255
Score = 30.4 bits (67), Expect = 1.6
Identities = 18/54 (33%), Positives = 25/54 (45%)
Frame = +2
Query: 91 ARCQGGANAGHTIYNAEGKKFALHLVPSGILNEDTLCVIGNGVVVHLPGLFQEI 144
A C A+ H++ GKK V SG L E C+ V V + G+F E+
Sbjct: 644 ANCPVVADITHSLQQPAGKKLDGGGVASGGLRELIPCIARTSVAVGVDGIFMEV 805
>AW775671 similar to GP|17065312|gb Unknown protein {Arabidopsis thaliana},
partial (22%)
Length = 624
Score = 29.6 bits (65), Expect = 2.7
Identities = 12/25 (48%), Positives = 18/25 (72%)
Frame = +2
Query: 227 SDAALRFKDFKYGPDVLREEVEKYK 251
S+ AL K+F +GPD L E+VE ++
Sbjct: 542 SNKALLLKEFNFGPDALLEKVEDFE 616
>BE325995 similar to GP|15292869|gb| unknown protein {Arabidopsis thaliana},
partial (18%)
Length = 463
Score = 28.9 bits (63), Expect = 4.6
Identities = 30/106 (28%), Positives = 45/106 (42%), Gaps = 5/106 (4%)
Frame = -1
Query: 166 LLFDFHQTVDGLREAELSKSFIGTTKRGIGPCYSSKANRNGIRVGDLRYMETLPQ----- 220
L F + T+ G LS S +T+RG C + R I V + T P
Sbjct: 283 LAFPYTTTISGFSIFSLSDSRFNSTERGTE*C----SPRPDI-VPSISSNTTTPSI*ASH 119
Query: 221 KLDLLLSDAALRFKDFKYGPDVLREEVEKYKRYAERLEPYIADTVH 266
+ LL++ + R+K + G VLR YAE L+P + +H
Sbjct: 118 DIVLLITTKS*RWKKTRSGAKVLR--------YAEMLDPVVLQNLH 5
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.140 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,451,751
Number of Sequences: 36976
Number of extensions: 202823
Number of successful extensions: 896
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 889
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 895
length of query: 485
length of database: 9,014,727
effective HSP length: 100
effective length of query: 385
effective length of database: 5,317,127
effective search space: 2047093895
effective search space used: 2047093895
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC143341.5


