

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142143.3 + phase: 0 /pseudo
(212 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
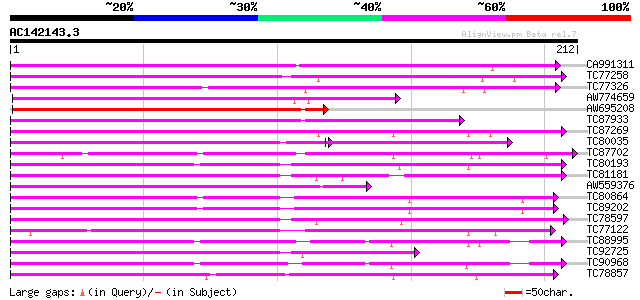
Score E
Sequences producing significant alignments: (bits) Value
CA991311 weakly similar to SP|O81973|C933 Cytochrome P450 93A3 (... 137 3e-33
TC77258 similar to GP|5059126|gb|AAD38930.1| cytochrome P450 mon... 125 2e-29
TC77326 similar to GP|12004682|gb|AAG44132.1 cytochrome P450 {Pi... 113 6e-26
AW774659 weakly similar to PIR|T00934|T009 probable cytochrome P... 111 2e-25
AW695208 similar to GP|9293968|dbj| cytochrome P450-like protein... 107 3e-24
TC87933 weakly similar to GP|5832707|dbj|BAA84071.1 cytochrome P... 101 2e-22
TC87269 similar to PIR|T05940|T05940 cytochrome P450 83D1p - soy... 101 2e-22
TC80035 weakly similar to GP|18252325|gb|AAL66194.1 cytochrome P... 72 7e-21
TC87702 weakly similar to SP|O48923|C7DA_SOYBN Cytochrome P450 7... 92 1e-19
TC80193 weakly similar to SP|O22307|C7DB_LOTJA Cytochrome P450 7... 89 1e-18
TC81181 similar to PIR|T05940|T05940 cytochrome P450 83D1p - soy... 88 3e-18
AW559376 weakly similar to GP|9759545|dbj cytochrome P450 {Arabi... 86 8e-18
TC80864 similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (E... 86 1e-17
TC89202 weakly similar to SP|O48923|C7DA_SOYBN Cytochrome P450 7... 85 2e-17
TC78597 similar to GP|166949|gb|AAA32913.1|| cytochrome P-450LXX... 83 9e-17
TC77122 similar to GP|5514645|emb|CAB50768.1 cytochrome P450 {Ci... 82 1e-16
TC88995 similar to SP|O81971|C7D9_SOYBN Cytochrome P450 71D9 (EC... 82 2e-16
TC92725 weakly similar to GP|5713172|gb|AAD47832.1| cytochrome P... 82 2e-16
TC90968 similar to SP|O81971|C7D9_SOYBN Cytochrome P450 71D9 (EC... 79 1e-15
TC78857 weakly similar to SP|O81974|C7D8_SOYBN Cytochrome P450 7... 77 4e-15
>CA991311 weakly similar to SP|O81973|C933 Cytochrome P450 93A3 (EC 1.14.-.-)
(P450 CP5). [Soybean] {Glycine max}, partial (33%)
Length = 768
Score = 137 bits (345), Expect = 3e-33
Identities = 71/212 (33%), Positives = 125/212 (58%), Gaps = 6/212 (2%)
Frame = +3
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTH+ +FS R G Y + F+ APYG YW+FMKKIC+++LL L +F +R
Sbjct: 3 LKTHETSFSNRLINGVVRYLSYGANDFMFAPYGDYWKFMKKICMSELLGGRTLDQFRPLR 182
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
+QE +LL L + D+G + LTN I+ R ++ TC + D E I+ LV+
Sbjct: 183 QQETLRLLNVLKKNGETGEAVDVGAELLRLTNRIMTRTTITKTCCEN-GFDVEDIRKLVK 359
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQG- 179
+ +G + ++ + + +DL G+ K+L++++ FD ++E ++EH+E + G
Sbjct: 360 DCSELGGQFNLSDFIWFFKNWDLQGFNKRLKELMNMFDTMIESVIREHQEEMKKRKENGE 539
Query: 180 -----DMMDILLQVYRNENAEVRLTRNDIKAF 206
D++DILL+++ +E++E++LTR ++KAF
Sbjct: 540 GAHVKDLLDILLEIHEDESSEIKLTRKNVKAF 635
>TC77258 similar to GP|5059126|gb|AAD38930.1| cytochrome P450
monooxygenaseCYP93D1 {Glycine max}, partial (95%)
Length = 1823
Score = 125 bits (313), Expect = 2e-29
Identities = 76/215 (35%), Positives = 123/215 (56%), Gaps = 7/215 (3%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KT + +F+ RP +S + Y + + PYG YWRF+KK+C+T+LLS L F+HIR
Sbjct: 349 LKTCEESFTNRPIMIASENLTYGAADYFFIPYGNYWRFLKKLCMTELLSGKTLEHFVHIR 528
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEV--IQCL 118
E E++ + +LL S + ++ + TNNI+ RM+MG K ++ EV ++ +
Sbjct: 529 EDEIKCFMGTLLEISKNGKPIEMRHELIRHTNNIISRMTMGK---KSSGMNDEVGQLRKV 699
Query: 119 VREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTE--- 175
VRE + ++G+++G + DL G+GK+ + + D ++E LKEHEE E
Sbjct: 700 VREIGELLGAFNLGDIIGFMRPLDLQGFGKRNKDTHHKMDVMMEKVLKEHEEARAKEGAG 879
Query: 176 -DCQGDMMDILLQ-VYRNENAEVRLTRNDIKAFFL 208
D + D+ DILL + ++ AE +LTR KAF L
Sbjct: 880 SDRKKDLFDILLNLIEADDGAESKLTRQSAKAFAL 984
>TC77326 similar to GP|12004682|gb|AAG44132.1 cytochrome P450 {Pisum
sativum}, complete
Length = 1864
Score = 113 bits (282), Expect = 6e-26
Identities = 66/211 (31%), Positives = 121/211 (57%), Gaps = 5/211 (2%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHD + RP+ + Y S + YGPYWR +++C+ +L S+ +L + +IR
Sbjct: 328 LKTHDATLAGRPKLSAGKYTTYNYSDITWSQYGPYWRQARRMCLLELFSAKRLESYEYIR 507
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYN---IDPEVIQC 117
+QE+ L L +++N++ + +TL+ N++ RM +G +K + I P+ +
Sbjct: 508 KQEMHDFLHKLF--NSKNKTILVKDHLSTLSLNVISRMVLGKKYLEKTDNAVISPDEFKK 681
Query: 118 LVREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEH-EERINTED 176
++ E + L++G+ + + DL GY K+++ + +FD+ +E L+EH E R N +D
Sbjct: 682 MLDELFLLNGILNIGDFIPWIHFLDLQGYVKRMKTLSKKFDRFMEHVLEEHIERRKNVKD 861
Query: 177 -CQGDMMDILLQVYRNENAEVRLTRNDIKAF 206
DM+D+LLQ+ + N EV+L R+ +KAF
Sbjct: 862 YVAKDMVDVLLQLAEDPNLEVKLERHGVKAF 954
>AW774659 weakly similar to PIR|T00934|T009 probable cytochrome P450
[imported] - Arabidopsis thaliana, partial (13%)
Length = 631
Score = 111 bits (278), Expect = 2e-25
Identities = 71/184 (38%), Positives = 90/184 (48%), Gaps = 39/184 (21%)
Frame = +3
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
KT+DL FS RP F + Y S FITAPYGPYWRFMKK+CVT+LLS QL R IR
Sbjct: 78 KTNDLAFSSRPAFTFADRLPYGTSGFITAPYGPYWRFMKKLCVTELLSGRQLERSRSIRR 257
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCF-------KKYNI---- 110
E+E +K +L + + DLG + LTNN+ CRM M T C +K I
Sbjct: 258 DEIELSVKRVLENAGSAAAVDLGAELMKLTNNVTCRMVMSTRCVIRS**ENRKSKIFVFL 437
Query: 111 ----------------------------DPEVIQCLVREFMHVGAKLSMGEVLGPLGKFD 142
D + I+ LV+E + AKL G+VLGPL +
Sbjct: 438 IYVSHNTCVLVVFNSLSFCGTRCSEKCDDADKIRKLVKESFELAAKLCFGDVLGPLKELS 617
Query: 143 LFGY 146
+ Y
Sbjct: 618 FWMY 629
>AW695208 similar to GP|9293968|dbj| cytochrome P450-like protein
{Arabidopsis thaliana}, partial (13%)
Length = 431
Score = 107 bits (267), Expect = 3e-24
Identities = 59/118 (50%), Positives = 74/118 (62%)
Frame = +3
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
KT+DL FS RP F + Y S FITAPYGPYWRFMKK+CVT+LLS QL R IR
Sbjct: 81 KTNDLAFSSRPAFTFADRLPYGTSGFITAPYGPYWRFMKKLCVTELLSGRQLERSRSIRR 260
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLV 119
E+E +K +L + + DLG + LTNN+ CRM M T C +K + D + I+ LV
Sbjct: 261 DEIELSVKRVLENAGSAAAVDLGAELMKLTNNVTCRMVMSTRCSEKCD-DADKIRKLV 431
>TC87933 weakly similar to GP|5832707|dbj|BAA84071.1 cytochrome P450
{Antirrhinum majus}, partial (68%)
Length = 1614
Score = 101 bits (251), Expect = 2e-22
Identities = 54/170 (31%), Positives = 92/170 (53%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTH+L +S+R + + Y + F APYG YW+F+KK+ T+LL + + +F+ IR
Sbjct: 313 LKTHELTYSHRKMSIAINVVAYDDATFAFAPYGTYWKFIKKLSTTELLGNRTMAQFLPIR 492
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
QEL + +++L S S +L L+NNI+ RM + C N E ++ LVR
Sbjct: 493 TQELHEFIQTLANKSKAEESVNLTQALIKLSNNIISRMMLSIDCSGTDN-QAEQVRALVR 669
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEE 170
E + + ++ + +G FDL G+ K+ I +D +L+ + + EE
Sbjct: 670 EVTQIFGEFNVSDFIGIFKNFDLQGFKKRALDIHKRYDALLDKIISDREE 819
>TC87269 similar to PIR|T05940|T05940 cytochrome P450 83D1p - soybean
(fragment), partial (79%)
Length = 1878
Score = 101 bits (251), Expect = 2e-22
Identities = 69/221 (31%), Positives = 110/221 (49%), Gaps = 13/221 (5%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHDL F+ RP F Y G APY PYWR M+K+CV L SS ++ F +R
Sbjct: 490 LKTHDLKFASRPSFLGLRKLSYNGLDLAFAPYSPYWREMRKLCVQHLFSSQRVHSFRPVR 669
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEV------ 114
E E+ +L++ L + + +L +LTN I+C+++ G T Y + E+
Sbjct: 670 ENEVAQLIQKLSQYGGDEKGANLSEILMSLTNTIICKIAFGKTYVCDYEEEVELGSGQKR 849
Query: 115 --IQCLVREFMHVGAKLSMGEVLGPLGKFD-LFGYGKKLRKIVGEFDKILEGFLKEHEE- 170
+Q L+ E + A+ + LG D + G KL K E D I + + +H +
Sbjct: 850 SRLQVLLNEAQALLAEFYFSDNFPLLGWIDRVKGTLGKLDKTFKELDLIYQRVIDDHMDN 1029
Query: 171 --RINTEDCQ-GDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
R T++ + D++DILLQ+ + + LT + IKA +
Sbjct: 1030SARPKTKEQEVDDIIDILLQMMNDHSLSFDLTLDHIKAVLM 1152
>TC80035 weakly similar to GP|18252325|gb|AAL66194.1 cytochrome P450 {Pyrus
communis}, partial (49%)
Length = 1342
Score = 72.0 bits (175), Expect(2) = 7e-21
Identities = 45/121 (37%), Positives = 63/121 (51%)
Frame = +3
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHD FS RP+ +S Y G APYG YWR MKK+C LLS S++ F +R
Sbjct: 276 LKTHDAIFSSRPKLQASEHMFYGGKGLGLAPYGSYWRNMKKLCTLHLLSGSKVEMFASMR 455
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
+EL L+KS+ + +L + NI +M +G C K ++D +VI V
Sbjct: 456 SEELGMLIKSVDKAAVLGEVVNLSEIVGEVIANITYKMVLG--CNKDSDLDLKVIYSRVY 629
Query: 121 E 121
E
Sbjct: 630 E 632
Score = 45.1 bits (105), Expect(2) = 7e-21
Identities = 21/70 (30%), Positives = 40/70 (57%)
Frame = +1
Query: 119 VREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQ 178
+RE M++ ++ + L L FD+ G +++K FD+I+E +K+HE+ + +
Sbjct: 616 IRECMNLAGSFNLADFLPWLSIFDMQGLTSRMKKTSKAFDQIVEKIIKDHEQDMKKDPHH 795
Query: 179 GDMMDILLQV 188
D +DILL +
Sbjct: 796 KDFIDILLSL 825
>TC87702 weakly similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (EC
1.14.-.-). [Soybean] {Glycine max}, partial (51%)
Length = 1732
Score = 92.4 bits (228), Expect = 1e-19
Identities = 68/221 (30%), Positives = 115/221 (51%), Gaps = 9/221 (4%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSH--DYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMH 58
MKTHDLNF+ RP S+ Y YKG F +P+G YWR M+KIC +LLS +++ F
Sbjct: 301 MKTHDLNFANRPPLLSAEIVTYGYKGMTF--SPHGSYWRQMRKICTMELLSQNRVESFRL 474
Query: 59 IREQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCL 118
RE+EL +K I S+E +L +L + R + G + D E + +
Sbjct: 475 QREEELANFVKD--INSSEGSPINLSEKLDSLAYGLTSRTAFGANVEDE---DKEKYRKI 639
Query: 119 VREFMHVGAKLSMGEVLGPLGKFD-LFGYGKKLRKIVGEFDKILEGFLKEHEER-INT-- 174
+++ + V S+ ++ + L G +++ K+ GE DKILE ++ H ++ + T
Sbjct: 640 IKDVLKVAGGFSLADLYPSIRILQVLTGLRQRIEKLHGETDKILENIVRSHRKKNLETKA 819
Query: 175 -EDCQGDMMDILLQVYRNENAEVRLT--RNDIKAFFLVSCF 212
E+ D++D+LL++ N + E + R+ K F + C+
Sbjct: 820 KEEKWEDLVDVLLKLQTNSDLEHPIIK*RSQSKNFGYLRCW 942
>TC80193 weakly similar to SP|O22307|C7DB_LOTJA Cytochrome P450 71D11 (EC
1.14.-.-) (Fragment). {Lotus japonicus}, partial (32%)
Length = 941
Score = 89.0 bits (219), Expect = 1e-18
Identities = 61/214 (28%), Positives = 104/214 (48%), Gaps = 6/214 (2%)
Frame = +2
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MKTHD+NF+ RPQ SS Y + +++PYG YWR ++KIC +LLS ++ + IR
Sbjct: 257 MKTHDINFATRPQGLSSDLIAYNSTGIVSSPYGNYWRQLRKICTLELLSLKRVNSYQPIR 436
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
E+E L+K I S E +L + I+ + + G KK+ + + + +
Sbjct: 437 EEEFSNLVK--WIASKEGSPINLTQAVLSSIYTIVSKSAFG----KKFKVPQKNLYQQKK 598
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLF-GYGKKLRKIVGEFDKILEGFLKEHEE-----RINT 174
+ + L+ + F G KL ++ D+I+E + EH+E +
Sbjct: 599 RY*RLQQVLT*QSYFPSITWMHFFTGLRPKLERVHRVVDQIMENIINEHKEAKSKGEFDQ 778
Query: 175 EDCQGDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
+ D++D+LL+ N E LT ++IKA +
Sbjct: 779 VESDEDLVDVLLKYQDGNNKEFFLTIDNIKAIIM 880
>TC81181 similar to PIR|T05940|T05940 cytochrome P450 83D1p - soybean
(fragment), partial (37%)
Length = 905
Score = 87.8 bits (216), Expect = 3e-18
Identities = 64/230 (27%), Positives = 101/230 (43%), Gaps = 22/230 (9%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHDL F+ RP F Y G APY YWR MKK+C L S L F IR
Sbjct: 235 LKTHDLKFASRPSFLGLRKLSYNGLDLGFAPYSSYWRDMKKLCALHLFSPKSLHSFRPIR 414
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPE------- 113
E E+ +L++ L + + +L + TN I+C+++ G KKY D E
Sbjct: 415 ENEVAELIQKLSQYDGDEKGVNLSEILISFTNAIICKIAFG----KKYVFDYEEEVELGS 582
Query: 114 ------------VIQCLVREFM---HVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFD 158
Q L+ EF H + + G LG+ D KKL+++ +
Sbjct: 583 GERRSRLQVLLNEAQALLTEFYFSDHFPLLGWVDRIKGTLGRLD-----KKLKELDLIYQ 747
Query: 159 KILEGFLKEHEERINTEDCQGDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
++++ + + E D++DI LQ+ + + LT + +KA +
Sbjct: 748 QVIDDHMDNSTKPKTKEQEVADIIDIFLQMMNDNSLSFDLTLDHVKAVLM 897
>AW559376 weakly similar to GP|9759545|dbj cytochrome P450 {Arabidopsis
thaliana}, partial (24%)
Length = 636
Score = 86.3 bits (212), Expect = 8e-18
Identities = 45/135 (33%), Positives = 79/135 (58%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KT++ F RP+ + Y S F APYG YW+FMKK+C+T+LL L +++ IR
Sbjct: 232 LKTNETCFLNRPKQSNLEYITYGSSDFALAPYGTYWKFMKKLCMTELLGGRILHQYLPIR 411
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
+E+ LK ++ ++ + ++G + T L+NNI+ RM+ C ++I+ LV+
Sbjct: 412 AEEITLFLKGMMEKADLRKEVNVGEELTMLSNNIITRMAXKRRCSDVEGEGHQLIE-LVK 588
Query: 121 EFMHVGAKLSMGEVL 135
E +G K ++G++L
Sbjct: 589 EMAVLGGKFNLGDML 633
>TC80864 similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (EC
1.14.-.-). [Soybean] {Glycine max}, partial (46%)
Length = 1083
Score = 85.9 bits (211), Expect = 1e-17
Identities = 60/207 (28%), Positives = 104/207 (49%), Gaps = 2/207 (0%)
Frame = +3
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MKTHDLNF RP S+ Y Y + A YG +WR ++KICV +LLS+ ++ F IR
Sbjct: 351 MKTHDLNFCDRPNLLLSNIYSYNATDIAFAAYGEHWRQLRKICVIELLSAKRVQSFRSIR 530
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
E E+ L+KS I ++E +L ++TN I R + G K N +V +
Sbjct: 531 EDEVSNLVKS--ITASEGSVVNLTRKIFSMTNGITARAAFG-----KRNRHQDVFIPALE 689
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLFGYGK-KLRKIVGEFDKILEGFLKEHEERINTEDCQG 179
+ + + + + ++ + K K+ K+ E D I + + +H
Sbjct: 690 KVVVLLGRFEIADLYPSIKLLQWMTREKTKMEKLHTEIDMIAQDIIDDHRSIHKEASNDE 869
Query: 180 DMMDILLQVYR-NENAEVRLTRNDIKA 205
D++D+LL++ + N ++E LT +++K+
Sbjct: 870 DLVDVLLKIQQENYHSEHPLTDDNMKS 950
>TC89202 weakly similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (EC
1.14.-.-). [Soybean] {Glycine max}, partial (38%)
Length = 1040
Score = 84.7 bits (208), Expect = 2e-17
Identities = 61/207 (29%), Positives = 104/207 (49%), Gaps = 2/207 (0%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MKTHDLNFS RP S + Y + I + YG WR ++KICV +LLS+ ++ F IR
Sbjct: 298 MKTHDLNFSGRPNLLLSTIWSYNATDVIFSIYGERWRQLRKICVIELLSAKRVQSFRSIR 477
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
E E+ L+KS I ++E +L + T I R + G K + EV + +
Sbjct: 478 EDEVTNLVKS--ITASEGSVVNLTQKILSTTYGITARAAFG-----KRSKHQEVFRSAIE 636
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLFGYGK-KLRKIVGEFDKILEGFLKEHEERINTEDCQG 179
E + + + ++ + K K+ K+ E D L+ + +H+ E
Sbjct: 637 EVASLLGGVCIVDLFPSIKLLQCLSRAKTKMEKLHKELDMTLQDIIDDHKSIHKEESNDE 816
Query: 180 DMMDILLQVYR-NENAEVRLTRNDIKA 205
D++D+LL++ + N +++ LT ++IK+
Sbjct: 817 DLVDVLLKIQQENYHSQHPLTDDNIKS 897
>TC78597 similar to GP|166949|gb|AAA32913.1|| cytochrome P-450LXXIA1
(cyp71A1) {Persea americana}, partial (18%)
Length = 1170
Score = 82.8 bits (203), Expect = 9e-17
Identities = 60/215 (27%), Positives = 105/215 (47%), Gaps = 6/215 (2%)
Frame = +2
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MK HD FS RPQ + LY YG W+ +K+CV++LL+ + + +R
Sbjct: 365 MKNHDRIFSNRPQHIAPKILLYGCDDVGFGLYGENWKQKRKLCVSELLNMKSVQSYHFVR 544
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPE-VIQCLV 119
E+E+++L+ L S + +L + +NNI+C+ ++G +KY D E ++ L
Sbjct: 545 EEEVDELVNKLREASLNDECVNLSEMIISTSNNIVCKCTLG----RKYEGDNEGNVKELA 712
Query: 120 REFMHVGAKLSMGEVLGPLGKFDLFG-----YGKKLRKIVGEFDKILEGFLKEHEERINT 174
R+ M ++G+ LG D+ + + R++ FD+++E L ++ N
Sbjct: 713 RKVMIYLQAFAVGDYFPSLGWIDVLSGKIREFKETFRELDSLFDQVIEERLALKKKMEND 892
Query: 175 EDCQGDMMDILLQVYRNENAEVRLTRNDIKAFFLV 209
+ + +DILLQ+ L+ NDIKA V
Sbjct: 893 QFKKKGFVDILLQLQEGGMLGFELSNNDIKALLTV 997
>TC77122 similar to GP|5514645|emb|CAB50768.1 cytochrome P450 {Cicer
arietinum}, partial (97%)
Length = 2308
Score = 82.4 bits (202), Expect = 1e-16
Identities = 51/213 (23%), Positives = 110/213 (50%), Gaps = 9/213 (4%)
Frame = +1
Query: 1 MKTHDL-NFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHI 59
++TH+ +F+ R Q + Y S + P+ PYW+F++K+ + LL+++ + + +
Sbjct: 409 LQTHEATSFNTRFQTSAISRLTYDNSVAMV-PFAPYWKFIRKLIMNDLLNATTVNKLRPL 585
Query: 60 REQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLV 119
R +E+ K+LK + + + D+ + TN+ + M +G + E ++ +
Sbjct: 586 RSREILKVLKVMANSAETQQPLDVTEELLKWTNSTISTMMLG---------EAEEVRDIA 738
Query: 120 REFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEE----RINTE 175
R+ + + + S+ + PL KF Y K+ +I ++D I+E +K+ +E R N E
Sbjct: 739 RDVLKIFGEYSVTNFIWPLNKFKFGNYEKRTDEIFNKYDPIIEKVIKKRQEIVNKRKNGE 918
Query: 176 DCQGD----MMDILLQVYRNENAEVRLTRNDIK 204
+G+ +D LL+ ++E ++++T+ IK
Sbjct: 919 IIEGEQSVVFLDTLLEYAQDETMDIKITKEQIK 1017
>TC88995 similar to SP|O81971|C7D9_SOYBN Cytochrome P450 71D9 (EC 1.14.-.-)
(P450 CP3). [Soybean] {Glycine max}, partial (71%)
Length = 1253
Score = 82.0 bits (201), Expect = 2e-16
Identities = 63/217 (29%), Positives = 103/217 (47%), Gaps = 9/217 (4%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MK HDL F+ RP +S Y APYG YWR ++KIC +LLSS ++ F IR
Sbjct: 280 MKNHDLVFASRPPIQASKIMSYDSLGLAFAPYGDYWRNLRKICTLELLSSKRVQSFQPIR 459
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
+E+ L+K I S E + + + + I R + G C + + +VR
Sbjct: 460 SEEVTNLIK--WISSKEGSQINFTKEVFSTISTITSRTAFGKKCKEN-----QKFISIVR 618
Query: 121 EFMHVGAKLSMGEVLGPLGKF--DLFGYGKKLRKIVGEFDKILEGFLKEHEE----RINT 174
+ + + +G+ L P + ++ G KL K+ + D I++ + EH E R+N
Sbjct: 619 DAIKIAGGFDLGD-LYPSCRLLQNISGLKPKLEKLHKQADLIMQNIIDEHREVNKSRVNE 795
Query: 175 ---EDCQGDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
E+ + D++D+LL+ + L N +KA L
Sbjct: 796 NQGEEVEEDLVDVLLK-------QDGLNDNSVKAVIL 885
>TC92725 weakly similar to GP|5713172|gb|AAD47832.1| cytochrome P450
{Nicotiana tabacum}, partial (21%)
Length = 895
Score = 82.0 bits (201), Expect = 2e-16
Identities = 46/156 (29%), Positives = 83/156 (52%), Gaps = 3/156 (1%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHDL F+ RP + Y + YG YWR M+K+C +LLS S++ F + R
Sbjct: 427 LKTHDLVFASRPPIEAGKVMFYNQKDVSFSVYGSYWRNMRKMCTLELLSHSKINSFRNTR 606
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKY---NIDPEVIQC 117
EQEL+ L+K + +N+ + D+ LT ++ C + G KKY +++ + +
Sbjct: 607 EQELDLLIKFIREAANDGTTVDISAKVAALTVDMTCIIVFG----KKYSDKDLNEKGFKA 774
Query: 118 LVREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKI 153
++E M + A ++ + + +G DL G ++++ I
Sbjct: 775 SMKELMSLAATSNIADFIPYIGALDLNGLTRRMKAI 882
>TC90968 similar to SP|O81971|C7D9_SOYBN Cytochrome P450 71D9 (EC 1.14.-.-)
(P450 CP3). [Soybean] {Glycine max}, partial (64%)
Length = 1112
Score = 79.3 bits (194), Expect = 1e-15
Identities = 65/217 (29%), Positives = 103/217 (46%), Gaps = 9/217 (4%)
Frame = +3
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHDL F+ RP +S Y +PYG YWR ++KIC +LLSS ++ F IR
Sbjct: 66 LKTHDLVFASRPPIQASKIMSYNSIGLSFSPYGDYWRQLRKICALELLSSKRVQSFQPIR 245
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
+E+ L+K I S E +L + + I R++ G C D + LV
Sbjct: 246 SEEMTNLIK--WIASKEGSEINLTKEVNSRIFLITSRVAFGKEC-----KDNKKFISLVW 404
Query: 121 EFMHVGAKLSMGEVLGPLGKF--DLFGYGKKLRKIVGEFDKILEGFLKEHE-------ER 171
E V ++G+ L P K+ ++ G KL K+ + D IL+ + EH
Sbjct: 405 EATRVAGGFNLGD-LYPSYKWLQNISGLKPKLEKLHKQTDVILQNIIDEHRLVDKSRAIE 581
Query: 172 INTEDCQGDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
++E+ D++D+LL+ + L+ N +KA L
Sbjct: 582 DHSEEVAEDLVDVLLK-------QDCLSDNSVKAVIL 671
>TC78857 weakly similar to SP|O81974|C7D8_SOYBN Cytochrome P450 71D8 (EC
1.14.-.-) (P450 CP7). [Soybean] {Glycine max}, partial
(38%)
Length = 1724
Score = 77.4 bits (189), Expect = 4e-15
Identities = 54/210 (25%), Positives = 103/210 (48%), Gaps = 5/210 (2%)
Frame = +3
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+K HD + S RP Y G I +PY WR ++KICV SS ++ F H+R
Sbjct: 354 LKDHDHDVSSRPPSHGPKTLSYNGIDMIFSPYNDCWREIRKICVVHFFSSKKISSFAHVR 533
Query: 61 EQELEKLLKSLL--ICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCL 118
+ E++ +++ + +CS ++ ++L +++++I+CR++ G + ++ + L
Sbjct: 534 KSEVKLMIEKISNHVCS--SKISNLSEVLMSVSSSIVCRIAFGKS-YEHEGGEKSRFHGL 704
Query: 119 VREFMHVGAKLSMGEVLGPLGKFD-LFGYGKKLRKIVGEFDKILEGFLKEHEERIN--TE 175
+ E + + + + +G D L G ++ D+ E LKEH N +
Sbjct: 705 LNETQAIFLSFFVSDYIPFMGWVDKLTGAIARVDNTFKALDEFFEQVLKEHLNPNNRKKD 884
Query: 176 DCQGDMMDILLQVYRNENAEVRLTRNDIKA 205
D + D++D+LL++ + LT N IKA
Sbjct: 885 DEEKDIVDVLLELKNQGRLSIDLTNNHIKA 974
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.142 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,146,985
Number of Sequences: 36976
Number of extensions: 105917
Number of successful extensions: 688
Number of sequences better than 10.0: 121
Number of HSP's better than 10.0 without gapping: 670
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 672
length of query: 212
length of database: 9,014,727
effective HSP length: 92
effective length of query: 120
effective length of database: 5,612,935
effective search space: 673552200
effective search space used: 673552200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC142143.3


