

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142143.13 - phase: 0
(259 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
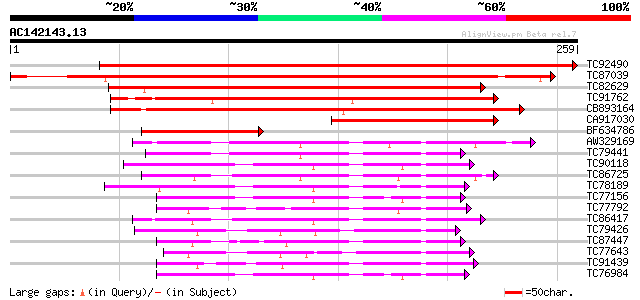
Sequences producing significant alignments: (bits) Value
TC92490 similar to GP|7269704|emb|CAB79652.1 predicted protein {... 449 e-127
TC87039 similar to GP|7269704|emb|CAB79652.1 predicted protein {... 327 2e-90
TC82629 similar to GP|7269704|emb|CAB79652.1 predicted protein {... 299 6e-82
TC91762 homologue to PIR|T13434|T13434 hypothetical protein T17A... 269 9e-73
CB893164 weakly similar to GP|6016718|gb| hypothetical protein {... 203 5e-53
CA917030 similar to GP|18461166|dbj contains ESTs AU090550(E2560... 160 6e-40
BF634786 similar to GP|7269704|emb| predicted protein {Arabidops... 100 6e-22
AW329169 similar to PIR|E84636|E84 NAM (no apical meristem)-like... 68 3e-12
TC79441 similar to GP|9757865|dbj|BAB08499.1 NAM (no apical meri... 67 7e-12
TC90118 homologue to GP|15795116|dbj|BAB02380. jasmonic acid reg... 67 9e-12
TC86725 similar to GP|9758909|dbj|BAB09485.1 NAM (no apical meri... 63 1e-10
TC78189 homologue to GP|14485513|emb|CAC42087. putative NAC doma... 63 1e-10
TC77156 homologue to GP|13877717|gb|AAK43936.1 OsNAC6 protein-li... 62 2e-10
TC77792 similar to GP|1418990|emb|CAA99760.1 unknown {Lycopersic... 60 8e-10
TC86417 similar to GP|16226943|gb|AAL16305.1 AT4g27410/F27G19_10... 57 9e-09
TC79426 similar to GP|7270509|emb|CAB80274.1 NAM / CUC2-like pro... 56 2e-08
TC87447 homologue to GP|15148912|gb|AAK84883.1 NAC domain protei... 55 2e-08
TC77643 similar to GP|21105742|gb|AAM34770.1 nam-like protein 7 ... 55 3e-08
TC91439 similar to PIR|D96683|D96683 hypothetical protein F12P19... 55 4e-08
TC76984 similar to GP|13877717|gb|AAK43936.1 OsNAC6 protein-like... 54 6e-08
>TC92490 similar to GP|7269704|emb|CAB79652.1 predicted protein {Arabidopsis
thaliana}, partial (57%)
Length = 1236
Score = 449 bits (1155), Expect = e-127
Identities = 217/218 (99%), Positives = 217/218 (99%)
Frame = +2
Query: 42 VRTLTCPSCGHNIEFQDQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLID 101
VRTLTCPSCGHNIEFQDQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLID
Sbjct: 2 VRTLTCPSCGHNIEFQDQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLID 181
Query: 102 EFIPTLEGENGICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETR 161
EFIPTLEGENGICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETR
Sbjct: 182 EFIPTLEGENGICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETR 361
Query: 162 WHKTGKTRPVIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVV 221
WHKTGKTRPVIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVV
Sbjct: 362 WHKTGKTRPVIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVV 541
Query: 222 SKKILEYLSSSERRLACNSVIRFMYSSVNFFLSLMASS 259
SKKILEYLSSS RRLACNSVIRFMYSSVNFFLSLMASS
Sbjct: 542 SKKILEYLSSS*RRLACNSVIRFMYSSVNFFLSLMASS 655
>TC87039 similar to GP|7269704|emb|CAB79652.1 predicted protein {Arabidopsis
thaliana}, partial (57%)
Length = 1150
Score = 327 bits (839), Expect = 2e-90
Identities = 162/254 (63%), Positives = 188/254 (73%), Gaps = 5/254 (1%)
Frame = +1
Query: 1 MTWCNDSDDQDRGIELITSTVIPGTSNESPLVTDDETKPTEV---RTLTCPSCGHNIEFQ 57
M WCND D N + VTD+ E+ +T+ CPSCGHNIE +
Sbjct: 52 MAWCNDKD------------------NNNSDVTDNIKLNEEIDQNQTIECPSCGHNIEVK 177
Query: 58 DQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTH 117
+QGG+NELPGLPAGVKFDP+D EIL+HLEAKV+ +V KLHPLIDEFIPT++GENGICYTH
Sbjct: 178 NQGGVNELPGLPAGVKFDPSDVEILEHLEAKVMSNVSKLHPLIDEFIPTIQGENGICYTH 357
Query: 118 PEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVM 177
P +LPGV KDGQIRHFFHRPSKAYTTGTRKRRKV TDEEG ETRWHKTGKTR + +GV+
Sbjct: 358 PAQLPGVSKDGQIRHFFHRPSKAYTTGTRKRRKVHTDEEGIETRWHKTGKTRAIFSNGVV 537
Query: 178 KGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVVSKKILEYLSSSERRLA 237
KGFKKILVLYTNY + KPEKTNWVMHQYHLG NEEE+DGE V SK + + R+
Sbjct: 538 KGFKKILVLYTNYCKDGKPEKTNWVMHQYHLGMNEEERDGEFVASK---IFYQTQPRQHG 708
Query: 238 CNSV--IRFMYSSV 249
NS+ +R Y +
Sbjct: 709 SNSINNVRDSYEKI 750
>TC82629 similar to GP|7269704|emb|CAB79652.1 predicted protein {Arabidopsis
thaliana}, partial (52%)
Length = 790
Score = 299 bits (766), Expect = 6e-82
Identities = 139/173 (80%), Positives = 153/173 (88%), Gaps = 1/173 (0%)
Frame = +3
Query: 46 TCPSCGHNIEFQDQG-GINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFI 104
TCP+CGH+I+ QDQG GI+ELPGLPAGVKFDP DQEIL+HLEAKV D+ LHPLIDEFI
Sbjct: 258 TCPTCGHHIKCQDQGAGIHELPGLPAGVKFDPTDQEILEHLEAKVRSDIQMLHPLIDEFI 437
Query: 105 PTLEGENGICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHK 164
PTLEGENGIC THPEKLPGV KDG IRHFFHRPSKAYTTGTRKRRKV TD +GSETRWHK
Sbjct: 438 PTLEGENGICCTHPEKLPGVGKDGLIRHFFHRPSKAYTTGTRKRRKVHTDADGSETRWHK 617
Query: 165 TGKTRPVIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDG 217
TGKTRPV + G +KG+KKILVLYTNY +Q+KPEKTNWVMHQYHLG+ + K G
Sbjct: 618 TGKTRPVFVSGKLKGYKKILVLYTNYRKQRKPEKTNWVMHQYHLGNMKRRKKG 776
>TC91762 homologue to PIR|T13434|T13434 hypothetical protein T17A13.50 -
Arabidopsis thaliana, partial (40%)
Length = 1002
Score = 269 bits (687), Expect = 9e-73
Identities = 132/184 (71%), Positives = 151/184 (81%), Gaps = 7/184 (3%)
Frame = +3
Query: 47 CPSCGHNIEFQDQGGINELPGLPAGVKFDPNDQEILQHLEAKVLF-DVPKLHPLIDEFIP 105
CP CGH DQG + L GLPAGVKFDP DQE+++HLEAKV ++ K HPLIDEFI
Sbjct: 303 CPRCGHKF---DQGKPDWL-GLPAGVKFDPTDQELIEHLEAKVESKNMQKSHPLIDEFIL 470
Query: 106 TLEGENGICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDE------EGSE 159
T+EGE+GICYTHPEKLPGV +DG RHFFHRPSKAYTTGTRKRRK+QT+E G E
Sbjct: 471 TIEGEDGICYTHPEKLPGVTRDGLSRHFFHRPSKAYTTGTRKRRKIQTEECDLQGSTGGE 650
Query: 160 TRWHKTGKTRPVIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGEL 219
TRWHKTGKTRPV+++G KG KKILVLYTN+G+ +KPEKTNWVMHQYHLG +EEEK+GEL
Sbjct: 651 TRWHKTGKTRPVMVNGKQKGCKKILVLYTNFGKNRKPEKTNWVMHQYHLGQHEEEKEGEL 830
Query: 220 VVSK 223
VVSK
Sbjct: 831 VVSK 842
>CB893164 weakly similar to GP|6016718|gb| hypothetical protein {Arabidopsis
thaliana}, partial (45%)
Length = 818
Score = 203 bits (517), Expect = 5e-53
Identities = 101/190 (53%), Positives = 125/190 (65%), Gaps = 1/190 (0%)
Frame = +3
Query: 47 CPSCGHNIEFQDQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPT 106
CP+C I+ D +E PG P G+KFDP+D EIL+HL AK K H I+EFIPT
Sbjct: 159 CPNCHSRIDNSDVS--SEWPGFPLGIKFDPSDVEILEHLAAKCGAGNIKPHMFIEEFIPT 332
Query: 107 LEGENGICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKV-QTDEEGSETRWHKT 165
LEGE GICYTHPE LPG KKDG HFFHR AY++G RKRRK+ RWHKT
Sbjct: 333 LEGEQGICYTHPENLPGAKKDGSSVHFFHRTMNAYSSGQRKRRKIHHRGSTEDHVRWHKT 512
Query: 166 GKTRPVIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVVSKKI 225
GKT+ +I G KGFKKI+VLY ++ K K++W MHQYHLG++E+EK+GE VVSK
Sbjct: 513 GKTKAIIEHGEHKGFKKIMVLYIRSEKESKSYKSDWKMHQYHLGTDEDEKNGEYVVSKIF 692
Query: 226 LEYLSSSERR 235
+ +E +
Sbjct: 693 YKQNEKNEEK 722
>CA917030 similar to GP|18461166|dbj contains ESTs AU090550(E2560)
AU063972(E2560)~similar to Arabidopsis thaliana
chromosome 4 F20O9., partial (25%)
Length = 724
Score = 160 bits (404), Expect = 6e-40
Identities = 75/76 (98%), Positives = 75/76 (98%)
Frame = +2
Query: 148 RRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYH 207
RRKVQTDEEGSETRWHKTGKTRPV IDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYH
Sbjct: 2 RRKVQTDEEGSETRWHKTGKTRPVFIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYH 181
Query: 208 LGSNEEEKDGELVVSK 223
LGSNEEEKDGELVVSK
Sbjct: 182 LGSNEEEKDGELVVSK 229
>BF634786 similar to GP|7269704|emb| predicted protein {Arabidopsis
thaliana}, partial (18%)
Length = 673
Score = 100 bits (249), Expect = 6e-22
Identities = 47/56 (83%), Positives = 50/56 (88%)
Frame = +1
Query: 61 GINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYT 116
GI+ELPGLPAGVKFDP DQEIL+HLEAKV D+ LHPLIDEFIPTLEGENGIC T
Sbjct: 505 GIHELPGLPAGVKFDPTDQEILEHLEAKVRSDIQMLHPLIDEFIPTLEGENGICCT 672
Score = 29.6 bits (65), Expect = 1.2
Identities = 10/14 (71%), Positives = 13/14 (92%)
Frame = +1
Query: 46 TCPSCGHNIEFQDQ 59
TCP+CGH+I+ QDQ
Sbjct: 265 TCPTCGHHIKCQDQ 306
>AW329169 similar to PIR|E84636|E84 NAM (no apical meristem)-like protein
[imported] - Arabidopsis thaliana, partial (52%)
Length = 631
Score = 68.2 bits (165), Expect = 3e-12
Identities = 56/190 (29%), Positives = 83/190 (43%), Gaps = 6/190 (3%)
Frame = +3
Query: 57 QDQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYT 116
++ GG E LP G +F P D+E++ + D F + + +
Sbjct: 6 RESGGKEET--LPPGFRFHPTDEELITCYLINKISD--------SSFTGKAITDIDLNKS 155
Query: 117 HPEKLPGVKKDGQIR-HFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVI--I 173
P +LPG K G+ +FF+ + Y TG R R T W TGK R + +
Sbjct: 156 EPWELPGKAKMGEKEWYFFNMRDRKYPTGVRTNRATNTGY------WKTTGKDREIFDSV 317
Query: 174 DGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSN---EEEKDGELVVSKKILEYLS 230
+ G KK LV Y GR + EK+NWVMH+Y + S K E V+ + + S
Sbjct: 318 TSELVGMKKTLVFYK--GRAPRGEKSNWVMHEYRIHSKSTFRTNKQDEWVICRIFKK--S 485
Query: 231 SSERRLACNS 240
S ++ CNS
Sbjct: 486 GSGKKYPCNS 515
>TC79441 similar to GP|9757865|dbj|BAB08499.1 NAM (no apical meristem)-like
protein {Arabidopsis thaliana}, partial (61%)
Length = 1247
Score = 67.0 bits (162), Expect = 7e-12
Identities = 47/147 (31%), Positives = 65/147 (43%), Gaps = 1/147 (0%)
Frame = +3
Query: 63 NELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLP 122
+E LP G +F P D+E++ H K + D F G+ + + P LP
Sbjct: 174 DEKMDLPPGFRFHPTDEELISHYLYKKVID--------SNFSARAIGDVDLNKSEPWDLP 329
Query: 123 GVKKDGQIR-HFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFK 181
K G+ +FF + Y TG R R + W TGK + + + G K
Sbjct: 330 FKAKMGEKEWYFFCVRDRKYPTGLRTNRATEAGY------WKATGKDKEIFKGKSLVGMK 491
Query: 182 KILVLYTNYGRQKKPEKTNWVMHQYHL 208
K LV Y GR K EK+NWVMH+Y L
Sbjct: 492 KTLVFYK--GRAPKGEKSNWVMHEYRL 566
>TC90118 homologue to GP|15795116|dbj|BAB02380. jasmonic acid regulatory
protein-like {Arabidopsis thaliana}, partial (43%)
Length = 612
Score = 66.6 bits (161), Expect = 9e-12
Identities = 48/163 (29%), Positives = 70/163 (42%), Gaps = 3/163 (1%)
Frame = +3
Query: 53 NIEFQDQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENG 112
N+E D ++ P LP G +F P D+E++ H K P +I E
Sbjct: 96 NMESTDSSTGSQQPNLPPGFRFHPTDEELVVHYLKKKAASAPLPVAIIAEV--------D 251
Query: 113 ICYTHPEKLPGVKKDGQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPV 171
+ P +LP G+ +F P + Y G R R + W TG +PV
Sbjct: 252 LYKFDPWELPAKATFGEQEWYFFSPRDRKYPNGARPNRAATSGY------WKATGTDKPV 413
Query: 172 IIDGVMK--GFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNE 212
+ G + G KK LV Y G+ + KTNW+MH+Y L N+
Sbjct: 414 LTSGGTQKVGVKKALVFYG--GKPPRGIKTNWIMHEYRLADNK 536
>TC86725 similar to GP|9758909|dbj|BAB09485.1 NAM (no apical meristem)-like
protein {Arabidopsis thaliana}, partial (50%)
Length = 1645
Score = 63.2 bits (152), Expect = 1e-10
Identities = 58/176 (32%), Positives = 78/176 (43%), Gaps = 13/176 (7%)
Frame = +3
Query: 61 GINELPGLPAGVKFDPNDQEILQ-HLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPE 119
G ++ LP G +F P D EI+ +L KV +F T GE + P
Sbjct: 156 GEDQTLDLPPGFRFHPTDAEIIVCYLTEKVKNS---------KFSATAIGEADLNKCEPW 308
Query: 120 KLPGVKKDGQIR-HFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGV-- 176
LP K G+ +FF + + Y TG R R ++ W TGK + + G
Sbjct: 309 DLPKKAKMGEKEWYFFCQKDRKYPTGMRTNRATESGY------WKATGKDKEIYHKGKGI 470
Query: 177 --MKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSN-------EEEKDGELVVSK 223
+ G KK LV Y GR K EKTNWVMH++ L +EKD E VVS+
Sbjct: 471 QNLVGMKKTLVFYK--GRAPKGEKTNWVMHEFRLEGKFATHNLPNKEKD-EWVVSR 629
>TC78189 homologue to GP|14485513|emb|CAC42087. putative NAC domain protein
{Solanum tuberosum}, partial (56%)
Length = 1556
Score = 62.8 bits (151), Expect = 1e-10
Identities = 49/171 (28%), Positives = 72/171 (41%), Gaps = 4/171 (2%)
Frame = +2
Query: 44 TLTCPSCGHNIEFQDQGGINELPG---LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLI 100
+L S HN E + + + G LP G +F P D E++ H + +P P+I
Sbjct: 44 SLQYQSTQHNTEEEKEEYTTRMQGALELPPGFRFHPTDDELVNHYLCRKCASLPIAVPII 223
Query: 101 DEFIPTLEGENGICYTHPEKLPGVKKDGQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSE 159
E + P LP + G+ +F P + Y G+R R T
Sbjct: 224 KEI--------DLYKFDPWHLPEMALYGEKEWYFFSPRDRKYPNGSRPNRAAGTGY---- 367
Query: 160 TRWHKTGKTRPVIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGS 210
W TG +P+ + G KK LV Y G+ K KTNW+MH+Y L +
Sbjct: 368 --WKATGADKPIGKPKAL-GIKKALVFYA--GKAPKGIKTNWIMHEYRLAN 505
>TC77156 homologue to GP|13877717|gb|AAK43936.1 OsNAC6 protein-like protein
{Arabidopsis thaliana}, partial (66%)
Length = 1096
Score = 62.4 bits (150), Expect = 2e-10
Identities = 45/144 (31%), Positives = 64/144 (44%), Gaps = 3/144 (2%)
Frame = +3
Query: 68 LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKD 127
LP G +F P D+E++ H + P P+I E + P LPG+
Sbjct: 258 LPPGFRFHPTDEELVMHYLCRKCTSQPIAVPIIAEI--------DLYKYDPWDLPGMATY 413
Query: 128 GQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMK--GFKKIL 184
G+ +F P + Y G+R R + W TG +P+ G K G KK L
Sbjct: 414 GEKEWYFFSPRDRKYPNGSRPNRAAGSGY------WKATGADKPI---GHPKPVGIKKAL 566
Query: 185 VLYTNYGRQKKPEKTNWVMHQYHL 208
V Y G+ K +KTNW+MH+Y L
Sbjct: 567 VFYA--GKAPKGDKTNWIMHEYRL 632
>TC77792 similar to GP|1418990|emb|CAA99760.1 unknown {Lycopersicon
esculentum}, partial (52%)
Length = 1022
Score = 60.1 bits (144), Expect = 8e-10
Identities = 45/147 (30%), Positives = 68/147 (45%), Gaps = 3/147 (2%)
Frame = +1
Query: 68 LPAGVKFDPNDQE-ILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKK 126
LP G +F P D+E ++Q+L+ KV PL IP ++ +C + P LPG +
Sbjct: 436 LPPGFRFHPTDEELVVQYLKRKVFS-----FPLPASIIPEVD----LCKSDPWDLPGDME 588
Query: 127 DGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGV--MKGFKKIL 184
Q R+FF Y G R R + W TG + ++ + G KK L
Sbjct: 589 --QERYFFSTKEAKYPNGNRSNRATNSGY------WKATGLDKQIMNSKTHEVAGMKKTL 744
Query: 185 VLYTNYGRQKKPEKTNWVMHQYHLGSN 211
V Y G+ +T+W+MH+Y L S+
Sbjct: 745 VFYR--GKPPHGSRTDWIMHEYRLTSS 819
>TC86417 similar to GP|16226943|gb|AAL16305.1 AT4g27410/F27G19_10
{Arabidopsis thaliana}, partial (67%)
Length = 1628
Score = 56.6 bits (135), Expect = 9e-09
Identities = 49/163 (30%), Positives = 69/163 (42%), Gaps = 2/163 (1%)
Frame = +2
Query: 57 QDQGGINELPGLPAGVKFDPNDQEIL-QHLEAKVLFDVPKLHPLIDEFIPTLEGENGICY 115
QD+ + +L LP G +F P D+E+L Q+L KV F + E +
Sbjct: 146 QDKDPLAQL-SLPPGFRFYPTDEELLVQYLCRKVAGH---------HFSLQIIAEIDLYK 295
Query: 116 THPEKLPGVKKDGQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIID 174
P LP G+ +F P + Y GTR R + W TG + + +
Sbjct: 296 FDPWILPSKAIFGEKEWYFFSPRDRKYPNGTRPNRVAGSGY------WKATGTDKIITNE 457
Query: 175 GVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDG 217
G G KK LV Y G+ K KTNW+MH+Y L + G
Sbjct: 458 GRKVGIKKALVFYV--GKAPKGTKTNWIMHEYRLLDSSRNNGG 580
>TC79426 similar to GP|7270509|emb|CAB80274.1 NAM / CUC2-like protein
{Arabidopsis thaliana}, partial (28%)
Length = 1011
Score = 55.8 bits (133), Expect = 2e-08
Identities = 44/153 (28%), Positives = 69/153 (44%), Gaps = 4/153 (2%)
Frame = +1
Query: 58 DQGGINELPGLPAGVKFDPNDQEILQH-LEAKVLFDVPKLHPLIDEFIPTLEGENGICYT 116
D + L LP G +F P D+E++ + L +K+ + ++ + E +C
Sbjct: 277 DGMAVLSLNSLPLGFRFRPTDEELIDYYLRSKINGNGDEVWVI---------REIDVCKW 429
Query: 117 HPEKLP--GVKKDGQIRHFFHRPS-KAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVII 173
P +P V ++ FF P + Y G R R + W TGK R +
Sbjct: 430 EPWDMPDLSVVRNKDPEWFFFCPQDRKYPNGHRLNRAT------NHGYWKATGKDRKIKS 591
Query: 174 DGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQY 206
++ G KK LV Y+ GR K ++TNWVMH+Y
Sbjct: 592 GTILIGMKKTLVFYS--GRAPKGKRTNWVMHEY 684
>TC87447 homologue to GP|15148912|gb|AAK84883.1 NAC domain protein NAC1
{Phaseolus vulgaris}, partial (92%)
Length = 1496
Score = 55.5 bits (132), Expect = 2e-08
Identities = 42/143 (29%), Positives = 65/143 (45%), Gaps = 2/143 (1%)
Frame = +2
Query: 68 LPAGVKFDPNDQEIL-QHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVK- 125
LP G +F P D+E++ +L KV D F+ ++ + C P +P
Sbjct: 110 LPPGFRFHPRDEELVCDYLMKKVTHS--------DSFL-MIDVDLNKC--EPWDIPEAAC 256
Query: 126 KDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKILV 185
G+ +F+ + + Y TG R R + W TGK R ++ G + G +K LV
Sbjct: 257 VGGKEWYFYTQRDRKYATGLRTNRATASGY------WKATGKDRAILRKGTLVGMRKTLV 418
Query: 186 LYTNYGRQKKPEKTNWVMHQYHL 208
Y GR K KT WVMH++ +
Sbjct: 419 FYQ--GRAPKGRKTEWVMHEFRI 481
>TC77643 similar to GP|21105742|gb|AAM34770.1 nam-like protein 7 {Petunia x
hybrida}, partial (27%)
Length = 1611
Score = 55.1 bits (131), Expect = 3e-08
Identities = 47/147 (31%), Positives = 65/147 (43%), Gaps = 5/147 (3%)
Frame = +2
Query: 71 GVKFDPNDQE-ILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLP--GVKKD 127
G +F P D+E ++ +L+ K+ K + + E + PE LP K
Sbjct: 632 GFRFHPTDEELVMYYLKRKICGKKLKFNVI---------KETDVYKWDPEDLPEQSFLKT 784
Query: 128 GQIRHFF--HRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKILV 185
G + FF HR K Y G R R + W TGK R VI + G KK LV
Sbjct: 785 GDRQWFFFCHRDRK-YPNGGRSSRATR------HGYWKATGKDRNVIYNSRSVGVKKTLV 943
Query: 186 LYTNYGRQKKPEKTNWVMHQYHLGSNE 212
Y GR E+T+WVMH+Y + +E
Sbjct: 944 FYL--GRAPSGERTDWVMHEYTMDEDE 1018
>TC91439 similar to PIR|D96683|D96683 hypothetical protein F12P19.8
[imported] - Arabidopsis thaliana, partial (28%)
Length = 1154
Score = 54.7 bits (130), Expect = 4e-08
Identities = 46/151 (30%), Positives = 65/151 (42%), Gaps = 4/151 (2%)
Frame = +3
Query: 68 LPAGVKFDPNDQEILQH-LEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPG--- 123
LP G +F P D+E++ + L K+ H + E I ++ + P LPG
Sbjct: 51 LPPGFRFHPTDEELVAYYLNRKI-----NGHKIELEIIAEVD----LYKCEPWDLPGKSL 203
Query: 124 VKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKI 183
+ + +FF + Y G+R R ++ W TGK R V G KK
Sbjct: 204 LPGNDMEWYFFSPRDRKYPNGSRTNRATKSGY------WKATGKDRKVNSQARAVGVKKT 365
Query: 184 LVLYTNYGRQKKPEKTNWVMHQYHLGSNEEE 214
LV Y GR +TNWVMH+Y L E E
Sbjct: 366 LVYYR--GRAPHGSRTNWVMHEYRLDERECE 452
>TC76984 similar to GP|13877717|gb|AAK43936.1 OsNAC6 protein-like protein
{Arabidopsis thaliana}, partial (71%)
Length = 1450
Score = 53.9 bits (128), Expect = 6e-08
Identities = 41/146 (28%), Positives = 63/146 (43%), Gaps = 3/146 (2%)
Frame = +2
Query: 68 LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKD 127
LP G +F P D+E++ H + P++ E + P +LP +
Sbjct: 155 LPPGFRFHPTDEELVNHYLCRKCAGQSISVPVVKEV--------DLYKFDPWQLPEMGYH 310
Query: 128 GQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMK--GFKKIL 184
+ +F P + Y G+R R + W TG +P+ G +K G KK L
Sbjct: 311 SEKEWYFFSPRDRKYPNGSRPNRAAGSGY------WKATGADKPI---GKLKPMGIKKAL 463
Query: 185 VLYTNYGRQKKPEKTNWVMHQYHLGS 210
V Y G+ K KTNW+MH+Y L +
Sbjct: 464 VFYA--GKAPKGVKTNWIMHEYRLAN 535
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,211,574
Number of Sequences: 36976
Number of extensions: 109710
Number of successful extensions: 631
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 618
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 621
length of query: 259
length of database: 9,014,727
effective HSP length: 94
effective length of query: 165
effective length of database: 5,538,983
effective search space: 913932195
effective search space used: 913932195
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC142143.13


