

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
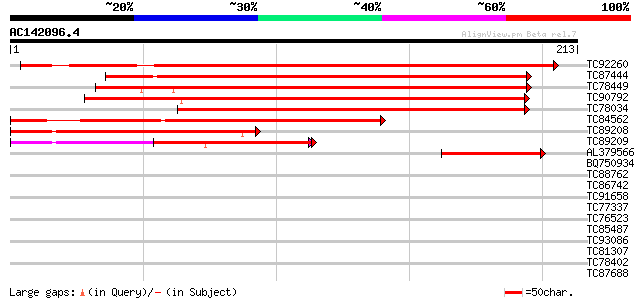
Query= AC142096.4 - phase: 0 /pseudo
(213 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC92260 homologue to GP|15912211|gb|AAL08239.1 At2g31160/T16B12.... 306 4e-84
TC87444 similar to GP|21555695|gb|AAM63916.1 unknown {Arabidopsi... 256 6e-69
TC78449 similar to GP|21555695|gb|AAM63916.1 unknown {Arabidopsi... 252 9e-68
TC90792 similar to GP|4559331|gb|AAD22993.1| expressed protein {... 222 7e-59
TC78034 similar to GP|21592379|gb|AAM64330.1 unknown {Arabidopsi... 218 1e-57
TC84562 homologue to GP|15912211|gb|AAL08239.1 At2g31160/T16B12.... 188 2e-48
TC89208 homologue to GP|15912211|gb|AAL08239.1 At2g31160/T16B12.... 90 7e-19
TC89209 homologue to GP|15912211|gb|AAL08239.1 At2g31160/T16B12.... 87 4e-18
AL379566 homologue to GP|15912211|gb| At2g31160/T16B12.3 {Arabid... 74 5e-14
BQ750934 similar to PIR|T19673|T19 hypothetical protein C33B4.3 ... 37 0.004
TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Ar... 37 0.004
TC86742 similar to PIR|T01541|T01541 hypothetical protein A_IG00... 37 0.006
TC91658 similar to GP|14329812|emb|CAC40753. putative nucleosome... 37 0.007
TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Atel... 36 0.013
TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 36 0.013
TC85487 homologue to SP|P49688|RS2_ARATH 40S ribosomal protein S... 35 0.016
TC93086 similar to PIR|F96695|F96695 hypothetical protein F5A8.8... 35 0.021
TC81307 weakly similar to GP|4335772|gb|AAD17449.1| unknown prot... 35 0.021
TC78402 weakly similar to PIR|B84853|B84853 hypothetical protein... 35 0.028
TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~un... 35 0.028
>TC92260 homologue to GP|15912211|gb|AAL08239.1 At2g31160/T16B12.3
{Arabidopsis thaliana}, partial (60%)
Length = 842
Score = 306 bits (784), Expect = 4e-84
Identities = 152/202 (75%), Positives = 168/202 (82%)
Frame = +1
Query: 5 QEFESSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
+EF SS++ K++ IN TN ++S S+ T+S+SS+ P T+T T S
Sbjct: 1 EEFNSSSSTKNI------INFTNTTTSEDNKNISNFTSSSSSAV------PPATNTNTLS 144
Query: 65 RYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHTLICPFYGHP 124
RYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVH+ ICPF+GHP
Sbjct: 145 RYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHSQICPFFGHP 324
Query: 125 NPPASCPCPLRQAWGSLDALIGRLRAAFEENGGKPEANPFGARAVRLYLREVRDSQAKAR 184
NPPA CPCPLRQAWGSLDALIGRLRAAFEENGGKPE NPFGARAVRL+LREVRDSQ+KAR
Sbjct: 325 NPPAPCPCPLRQAWGSLDALIGRLRAAFEENGGKPEDNPFGARAVRLFLREVRDSQSKAR 504
Query: 185 GISYEKKKRKRPPQPPPPPPSN 206
GISYEKKKRKRP PPPPPSN
Sbjct: 505 GISYEKKKRKRPQIQPPPPPSN 570
>TC87444 similar to GP|21555695|gb|AAM63916.1 unknown {Arabidopsis
thaliana}, partial (68%)
Length = 1607
Score = 256 bits (653), Expect = 6e-69
Identities = 118/160 (73%), Positives = 136/160 (84%)
Frame = +2
Query: 37 PSSSTTSASSSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGA 96
P S S + TA P S+ +T TPSRYE+QKRRDWNTF QYLRNH+PPL+L+ CSGA
Sbjct: 533 PPPSIPSETPPETAQPQP-SSPATPTPSRYESQKRRDWNTFQQYLRNHKPPLTLALCSGA 709
Query: 97 HVLEFLRYLDQFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEENG 156
HV+EFL+YLDQFGKTKVH + CP++G PNPP+ C CPL+QAWGSLDALIGRLRAAFEENG
Sbjct: 710 HVIEFLKYLDQFGKTKVHVIGCPYFGQPNPPSPCACPLKQAWGSLDALIGRLRAAFEENG 889
Query: 157 GKPEANPFGARAVRLYLREVRDSQAKARGISYEKKKRKRP 196
GKPE+NPFG RAVR+YLREV+D QAKARG+ YEKKKRKRP
Sbjct: 890 GKPESNPFGTRAVRIYLREVKDGQAKARGVPYEKKKRKRP 1009
>TC78449 similar to GP|21555695|gb|AAM63916.1 unknown {Arabidopsis
thaliana}, partial (70%)
Length = 1031
Score = 252 bits (643), Expect = 9e-68
Identities = 121/175 (69%), Positives = 139/175 (79%), Gaps = 11/175 (6%)
Frame = +2
Query: 33 TMTIPSSSTTSASSSS--TATTSPPSTTST---------TTPSRYENQKRRDWNTFGQYL 81
T+ PSS A S ++TTSPP PSRYE+QKRRDWNTF QYL
Sbjct: 197 TIMDPSSEGAGAPSEQLPSSTTSPPIVVQPEGSSPAPPPAPPSRYESQKRRDWNTFLQYL 376
Query: 82 RNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSL 141
+NH+PPL+L RCSGAHV+EFL+YLDQFGKTKVH CP++GHPNPPA C CPL+QAWGSL
Sbjct: 377 QNHKPPLTLVRCSGAHVIEFLKYLDQFGKTKVHVSGCPYFGHPNPPAPCACPLKQAWGSL 556
Query: 142 DALIGRLRAAFEENGGKPEANPFGARAVRLYLREVRDSQAKARGISYEKKKRKRP 196
DALIGRLRAA+EENGG+PE+NPFGA+AVR+YLREVR+ QAKARGI YEKKKRKRP
Sbjct: 557 DALIGRLRAAYEENGGRPESNPFGAKAVRIYLREVREGQAKARGIPYEKKKRKRP 721
>TC90792 similar to GP|4559331|gb|AAD22993.1| expressed protein {Arabidopsis
thaliana}, partial (74%)
Length = 710
Score = 222 bits (566), Expect = 7e-59
Identities = 107/168 (63%), Positives = 131/168 (77%), Gaps = 1/168 (0%)
Frame = +2
Query: 29 SSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTP-SRYENQKRRDWNTFGQYLRNHRPP 87
S +M+ ++ + + ++SSSS S T+ P SRYE+QK+RDWNTFGQYLRN PP
Sbjct: 50 SPTMSSSMQNDTVQASSSSSRPGGSDQQPTAAVAPLSRYESQKKRDWNTFGQYLRNQSPP 229
Query: 88 LSLSRCSGAHVLEFLRYLDQFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGR 147
+SLS+C+ HVLEFLRYL QFGKTKVH C F+G P+PPA C CPLRQAWGSLDALIGR
Sbjct: 230 VSLSQCNFNHVLEFLRYLHQFGKTKVHLHGCIFFGQPDPPAPCTCPLRQAWGSLDALIGR 409
Query: 148 LRAAFEENGGKPEANPFGARAVRLYLREVRDSQAKARGISYEKKKRKR 195
LRAA+EE+GG PE NPFG A+R+YLREV++ Q+KARGI Y KKK+KR
Sbjct: 410 LRAAYEEHGGSPENNPFGTGAIRVYLREVKECQSKARGIPYTKKKKKR 553
>TC78034 similar to GP|21592379|gb|AAM64330.1 unknown {Arabidopsis
thaliana}, partial (73%)
Length = 947
Score = 218 bits (556), Expect = 1e-57
Identities = 100/132 (75%), Positives = 113/132 (84%)
Frame = +1
Query: 64 SRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHTLICPFYGH 123
SRYE+QKRRDWNTFGQYLRN RPP++LS C+ HVLEFLRYLDQFGKTKVH C F+G
Sbjct: 196 SRYESQKRRDWNTFGQYLRNQRPPVALSNCNSNHVLEFLRYLDQFGKTKVHLQGCLFFGQ 375
Query: 124 PNPPASCPCPLRQAWGSLDALIGRLRAAFEENGGKPEANPFGARAVRLYLREVRDSQAKA 183
PP C CPL+QAWGSLDALIGRLRAA+EENGG PE NPF + ++R+YLREVRDSQAKA
Sbjct: 376 TEPPGPCTCPLKQAWGSLDALIGRLRAAYEENGGLPETNPFASGSIRIYLREVRDSQAKA 555
Query: 184 RGISYEKKKRKR 195
RGI Y+KKK+KR
Sbjct: 556 RGIPYKKKKKKR 591
>TC84562 homologue to GP|15912211|gb|AAL08239.1 At2g31160/T16B12.3
{Arabidopsis thaliana}, partial (36%)
Length = 542
Score = 188 bits (477), Expect = 2e-48
Identities = 93/141 (65%), Positives = 106/141 (74%)
Frame = +1
Query: 1 MNSLQEFESSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTST 60
M+S+Q+F S N+ N S TI ++S+ + S++ +S PS ST
Sbjct: 88 MDSIQDFMDSCNSD------------NSCSLTNSTITTNSSNNNISNAIVGSSSPSG-ST 228
Query: 61 TTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHTLICPF 120
TT SRYENQKRRDWNTFGQYL+NHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHT ICPF
Sbjct: 229 TTSSRYENQKRRDWNTFGQYLKNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHTPICPF 408
Query: 121 YGHPNPPASCPCPLRQAWGSL 141
YGHPNPPA CPCPLRQAWG+L
Sbjct: 409 YGHPNPPAPCPCPLRQAWGNL 471
>TC89208 homologue to GP|15912211|gb|AAL08239.1 At2g31160/T16B12.3
{Arabidopsis thaliana}, partial (17%)
Length = 678
Score = 89.7 bits (221), Expect = 7e-19
Identities = 47/97 (48%), Positives = 66/97 (67%), Gaps = 3/97 (3%)
Frame = +1
Query: 1 MNSLQEFESSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTST 60
M+S+QEF + NN+++ N M N N + S T T +++TT+AS SS+++ + S
Sbjct: 376 MDSIQEFIGTCNNENL-TCNFMNNNNNNTISTTTTSLTTTTTTASGSSSSSAASTIINSP 552
Query: 61 TTPSRYENQKRRDWNTFGQYLRNHRP---PLSLSRCS 94
+ SRYENQKRRDWNTFGQYL+NHRP PL + RC+
Sbjct: 553 NSSSRYENQKRRDWNTFGQYLKNHRPPFFPLQVQRCT 663
Score = 30.4 bits (67), Expect = 0.53
Identities = 13/13 (100%), Positives = 13/13 (100%)
Frame = +2
Query: 89 SLSRCSGAHVLEF 101
SLSRCSGAHVLEF
Sbjct: 638 SLSRCSGAHVLEF 676
>TC89209 homologue to GP|15912211|gb|AAL08239.1 At2g31160/T16B12.3
{Arabidopsis thaliana}, partial (21%)
Length = 713
Score = 87.4 bits (215), Expect = 4e-18
Identities = 40/61 (65%), Positives = 46/61 (74%)
Frame = +2
Query: 55 PSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVH 114
PS+T T + + DWNTFGQYL+NHRPPLSLSRCSGAHVLEFLRYLDQF + +
Sbjct: 530 PSSTVPTVAAAMRTKNAGDWNTFGQYLKNHRPPLSLSRCSGAHVLEFLRYLDQFWQRQKF 709
Query: 115 T 115
T
Sbjct: 710 T 712
Score = 54.3 bits (129), Expect = 3e-08
Identities = 41/118 (34%), Positives = 64/118 (53%), Gaps = 4/118 (3%)
Frame = +1
Query: 1 MNSLQEFESSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTST 60
M+S+QEF + NN+++ N M N N + S T T +++TT+AS SS+++ + S
Sbjct: 370 MDSIQEFIGTCNNENL-TCNFMNNNNNNTISTTTTSLTTTTTTASGSSSSSAASTIINSP 546
Query: 61 TTPSRYENQKRR----DWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVH 114
+ SRYENQKRR W+ Q + PL + RC+ + + + KTKVH
Sbjct: 547 NSSSRYENQKRR*LEHLWSV-PQESSSTSFPLQVQRCTCP*ISKVFGSI--LAKTKVH 711
>AL379566 homologue to GP|15912211|gb| At2g31160/T16B12.3 {Arabidopsis
thaliana}, partial (15%)
Length = 488
Score = 73.6 bits (179), Expect = 5e-14
Identities = 35/39 (89%), Positives = 36/39 (91%)
Frame = +2
Query: 163 PFGARAVRLYLREVRDSQAKARGISYEKKKRKRPPQPPP 201
PFGARAVRLYLREVRD Q+KARGISYEKKKRKRPPQ P
Sbjct: 2 PFGARAVRLYLREVRDLQSKARGISYEKKKRKRPPQQQP 118
>BQ750934 similar to PIR|T19673|T19 hypothetical protein C33B4.3 -
Caenorhabditis elegans, partial (1%)
Length = 627
Score = 37.4 bits (85), Expect = 0.004
Identities = 20/40 (50%), Positives = 32/40 (80%)
Frame = +2
Query: 24 NITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTP 63
N T+P+SS + +I +S+T+S+SSSS++TTS S++S T P
Sbjct: 500 NTTSPASSPS-SITTSTTSSSSSSSSSTTSSSSSSSPTPP 616
Score = 32.3 bits (72), Expect = 0.14
Identities = 17/55 (30%), Positives = 35/55 (62%)
Frame = +2
Query: 2 NSLQEFESSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPS 56
+S+ E S++ ++ N ++PSS T T SSS++S+S++S++++S P+
Sbjct: 446 SSIPESSSTSLPPTLVPQNTTSPASSPSSITTSTTSSSSSSSSSTTSSSSSSSPT 610
Score = 28.5 bits (62), Expect = 2.0
Identities = 16/41 (39%), Positives = 26/41 (63%), Gaps = 4/41 (9%)
Frame = +2
Query: 28 PSSSMTMT----IPSSSTTSASSSSTATTSPPSTTSTTTPS 64
P SS T +P ++T+ ASS S+ TTS S++S+++ S
Sbjct: 455 PESSSTSLPPTLVPQNTTSPASSPSSITTSTTSSSSSSSSS 577
>TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Arabidopsis
thaliana}, partial (3%)
Length = 805
Score = 37.4 bits (85), Expect = 0.004
Identities = 21/46 (45%), Positives = 30/46 (64%), Gaps = 6/46 (13%)
Frame = +3
Query: 24 NITNPSSSMTMTIPSSSTTS------ASSSSTATTSPPSTTSTTTP 63
N T+PSSS + + P S+TTS ASS + TTSPP++T ++P
Sbjct: 81 NPTSPSSSPSASSPVSTTTSPPASTPASSPVSTTTSPPASTPASSP 218
Score = 34.7 bits (78), Expect = 0.028
Identities = 16/46 (34%), Positives = 29/46 (62%)
Frame = +3
Query: 19 TNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
++P+ T+P +S + P S+TTS +S+ A++ P+TTS P+
Sbjct: 114 SSPVSTTTSPPASTPASSPVSTTTSPPASTPASSPVPTTTSPPAPT 251
Score = 26.6 bits (57), Expect = 7.6
Identities = 18/51 (35%), Positives = 30/51 (58%), Gaps = 8/51 (15%)
Frame = +3
Query: 23 INITNPSSSMTMTIPS------SSTTSASSSSTATTSPPSTTST--TTPSR 65
++ +P++S ++P+ SSTT+ S S ++T PP T S+ TTP R
Sbjct: 267 VSTNSPTASPAGSLPAAATPSPSSTTAGSPSPSST--PPGTRSSNETTPGR 413
Score = 26.2 bits (56), Expect = 9.9
Identities = 15/43 (34%), Positives = 24/43 (54%)
Frame = +3
Query: 19 TNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTT 61
++P+ T+P +S + P +TTS + T +SP ST S T
Sbjct: 162 SSPVSTTTSPPASTPASSPVPTTTS-PPAPTPASSPVSTNSPT 287
>TC86742 similar to PIR|T01541|T01541 hypothetical protein A_IG005I10.16 -
Arabidopsis thaliana, partial (19%)
Length = 2073
Score = 37.0 bits (84), Expect = 0.006
Identities = 18/39 (46%), Positives = 27/39 (69%)
Frame = +3
Query: 26 TNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
T P S T+++ SSS+ S S S T+S PSTT+++TP+
Sbjct: 663 TQP*KSSTLSLSSSSSPSPPSPSPPTSSHPSTTTSSTPA 779
>TC91658 similar to GP|14329812|emb|CAC40753. putative nucleosome assembly
protein 1 {Atropa belladonna}, partial (46%)
Length = 583
Score = 36.6 bits (83), Expect = 0.007
Identities = 18/37 (48%), Positives = 31/37 (83%)
Frame = -2
Query: 28 PSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
PSSS + + SSS++S+SSSS++++SP S++S+T+ S
Sbjct: 183 PSSSSSSSSSSSSSSSSSSSSSSSSSPSSSSSSTSIS 73
Score = 32.7 bits (73), Expect = 0.11
Identities = 15/37 (40%), Positives = 28/37 (75%)
Frame = -2
Query: 28 PSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
P + PSSS++S+SSSS++++S S++S+++PS
Sbjct: 210 PLLLLLFPFPSSSSSSSSSSSSSSSSSSSSSSSSSPS 100
Score = 30.4 bits (67), Expect = 0.53
Identities = 16/39 (41%), Positives = 29/39 (74%)
Frame = -2
Query: 26 TNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
++ SSS + + SSS++S+SSSS++ +S S+TS ++ S
Sbjct: 180 SSSSSSSSSSSSSSSSSSSSSSSSSPSSSSSSTSISSKS 64
Score = 29.6 bits (65), Expect = 0.90
Identities = 15/37 (40%), Positives = 28/37 (75%)
Frame = -2
Query: 29 SSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPSR 65
SSS + + SSS++S+SSSS++++ S++ST+ S+
Sbjct: 177 SSSSSSSSSSSSSSSSSSSSSSSSPSSSSSSTSISSK 67
Score = 29.6 bits (65), Expect = 0.90
Identities = 16/43 (37%), Positives = 32/43 (74%), Gaps = 1/43 (2%)
Frame = -2
Query: 21 PMINITNPSSSMTMTIPSSSTTS-ASSSSTATTSPPSTTSTTT 62
P++ + P S + + SSS++S +SSSS++++S PS++S++T
Sbjct: 210 PLLLLLFPFPSSSSSSSSSSSSSSSSSSSSSSSSSPSSSSSST 82
Score = 27.7 bits (60), Expect = 3.4
Identities = 14/42 (33%), Positives = 27/42 (63%)
Frame = -2
Query: 26 TNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPSRYE 67
++ SSS + + SSS++S SSSS++T+ + S +P ++
Sbjct: 159 SSSSSSSSSSSSSSSSSSPSSSSSSTSISSKSLSPASPVNHD 34
Score = 26.9 bits (58), Expect = 5.8
Identities = 13/44 (29%), Positives = 30/44 (67%)
Frame = -2
Query: 21 PMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
P + ++ SSS + + SSS++S+SS S++++S ++ + +P+
Sbjct: 183 PSSSSSSSSSSSSSSSSSSSSSSSSSPSSSSSSTSISSKSLSPA 52
>TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Ateline
herpesvirus 3}, partial (8%)
Length = 986
Score = 35.8 bits (81), Expect = 0.013
Identities = 17/46 (36%), Positives = 35/46 (75%)
Frame = -2
Query: 18 NTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTP 63
+++ + + + PSSS + SSS++S+SSSS+++ S PS++S+++P
Sbjct: 727 SSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSSPSSSSSSSP 590
Score = 32.0 bits (71), Expect = 0.18
Identities = 16/43 (37%), Positives = 31/43 (71%)
Frame = -2
Query: 26 TNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPSRYEN 68
++ SSS + ++ SSST S+SSSS ++S S++S+++ S + +
Sbjct: 745 SSSSSSSSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSS 617
Score = 30.4 bits (67), Expect = 0.53
Identities = 17/55 (30%), Positives = 34/55 (60%)
Frame = -2
Query: 9 SSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTP 63
SS+++ +T + + PSSS + + SSS++S SS S++++S P +++ P
Sbjct: 730 SSSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSSPSSSSSSSPFSSAPPAP 566
Score = 29.6 bits (65), Expect = 0.90
Identities = 16/51 (31%), Positives = 32/51 (62%)
Frame = -2
Query: 9 SSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTS 59
SS++ +++P + ++ SSS + + SS +S+SSSS +++PP+ S
Sbjct: 712 SSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSSPSSSSSSSPFSSAPPAPPS 560
Score = 29.6 bits (65), Expect = 0.90
Identities = 13/35 (37%), Positives = 28/35 (79%)
Frame = -2
Query: 30 SSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
+S + + SSS +S+S+ S++++SPPS++S+++ S
Sbjct: 748 ASSSSSSSSSSLSSSSTPSSSSSSPPSSSSSSSSS 644
Score = 28.5 bits (62), Expect = 2.0
Identities = 15/39 (38%), Positives = 28/39 (71%)
Frame = -2
Query: 26 TNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
++ S S + T SSS++ SSSS++++S S++S ++PS
Sbjct: 727 SSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSSPS 611
Score = 28.5 bits (62), Expect = 2.0
Identities = 14/39 (35%), Positives = 24/39 (60%)
Frame = -2
Query: 38 SSSTTSASSSSTATTSPPSTTSTTTPSRYENQKRRDWNT 76
SSS++S SSSST ++S S S+++ S + W++
Sbjct: 733 SSSSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSS 617
Score = 27.7 bits (60), Expect = 3.4
Identities = 14/34 (41%), Positives = 22/34 (64%)
Frame = -2
Query: 28 PSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTT 61
PSS + PS + S SSSS++++S PS+ S +
Sbjct: 508 PSSDDNSSSPSPNAPSPSSSSSSSSSLPSSPSAS 407
Score = 26.6 bits (57), Expect = 7.6
Identities = 18/57 (31%), Positives = 32/57 (55%), Gaps = 2/57 (3%)
Frame = -2
Query: 2 NSLQEFESSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSS--STATTSPPS 56
+S SS+ ++ P + ++ SSS + + SS ++S+SSS S+A +PPS
Sbjct: 730 SSSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSSPSSSSSSSPFSSAPPAPPS 560
>TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (69%)
Length = 1615
Score = 35.8 bits (81), Expect = 0.013
Identities = 20/55 (36%), Positives = 34/55 (61%), Gaps = 1/55 (1%)
Frame = -3
Query: 8 ESSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPS-TTSTT 61
ESST T+ + S+S + T P++ +T++S+S+ +TT+ PS +TSTT
Sbjct: 1277 ESSTTTSSSSTTSTATIPSTKSTSTSSTTPTTKSTASSTSTKSTTTTPSTSTSTT 1113
Score = 33.5 bits (75), Expect = 0.062
Identities = 16/26 (61%), Positives = 22/26 (84%)
Frame = -3
Query: 39 SSTTSASSSSTATTSPPSTTSTTTPS 64
SSTT++SSS+T+T + PST ST+T S
Sbjct: 1274 SSTTTSSSSTTSTATIPSTKSTSTSS 1197
Score = 32.3 bits (72), Expect = 0.14
Identities = 18/39 (46%), Positives = 23/39 (58%)
Frame = -3
Query: 26 TNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
T SSS T T ST S S+SST T+ + +ST+T S
Sbjct: 1265 TTSSSSTTSTATIPSTKSTSTSSTTPTTKSTASSTSTKS 1149
>TC85487 homologue to SP|P49688|RS2_ARATH 40S ribosomal protein S2.
[Mouse-ear cress] {Arabidopsis thaliana}, partial (88%)
Length = 1114
Score = 35.4 bits (80), Expect = 0.016
Identities = 19/45 (42%), Positives = 26/45 (57%)
Frame = -2
Query: 20 NPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
NP + SSS T T +S TTS +++S TT+ +T TTT S
Sbjct: 222 NPFLFFFASSSSSTTTTTTSVTTSTTTTSVTTTTTKTTAKTTTIS 88
>TC93086 similar to PIR|F96695|F96695 hypothetical protein F5A8.8 [imported]
- Arabidopsis thaliana, partial (30%)
Length = 752
Score = 35.0 bits (79), Expect = 0.021
Identities = 27/77 (35%), Positives = 37/77 (47%), Gaps = 2/77 (2%)
Frame = -1
Query: 24 NITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRN 83
N+T PSS +T PS S T SSS ++ + ST + TT SR R T + +
Sbjct: 557 NVTGPSSGSLITSPSRSWTRLSSSRSSCS---STAAATTSSRLRTTSLRSSITIFICILS 387
Query: 84 HRPPLSL--SRCSGAHV 98
+R SL C GA +
Sbjct: 386 NRKTRSLVIISCRGATI 336
>TC81307 weakly similar to GP|4335772|gb|AAD17449.1| unknown protein
{Arabidopsis thaliana}, partial (31%)
Length = 1171
Score = 35.0 bits (79), Expect = 0.021
Identities = 19/52 (36%), Positives = 31/52 (59%), Gaps = 1/52 (1%)
Frame = +2
Query: 9 SSTNNKDMMNTNPMINITNPSSSMTMTI-PSSSTTSASSSSTATTSPPSTTS 59
SS N + P+ ++ +PSSS +++ PSSS +S+SS +PP +TS
Sbjct: 104 SSLNPNESSTLTPVTSLLSPSSSSFLSLSPSSSLPPSSTSSAPHPTPPPSTS 259
Score = 26.9 bits (58), Expect = 5.8
Identities = 18/39 (46%), Positives = 24/39 (61%), Gaps = 5/39 (12%)
Frame = +2
Query: 27 NPSSSMTMTIPSSSTTSASSSSTATTS-----PPSTTST 60
NP+ S T+T P +S S SSSS + S PPS+TS+
Sbjct: 113 NPNESSTLT-PVTSLLSPSSSSFLSLSPSSSLPPSSTSS 226
>TC78402 weakly similar to PIR|B84853|B84853 hypothetical protein At2g42370
[imported] - Arabidopsis thaliana, partial (14%)
Length = 1982
Score = 34.7 bits (78), Expect = 0.028
Identities = 22/53 (41%), Positives = 31/53 (57%), Gaps = 6/53 (11%)
Frame = -3
Query: 18 NTNPMINITNPSSSMTMTIPSSSTTSA------SSSSTATTSPPSTTSTTTPS 64
+T P+ PS+S T PSSS++S+ SSS T +SP +TSTT+ S
Sbjct: 570 STPPIFTSPLPSTSSFTTTPSSSSSSSFASPSSSSSFTTPSSPSPSTSTTSVS 412
Score = 26.2 bits (56), Expect = 9.9
Identities = 14/28 (50%), Positives = 21/28 (75%)
Frame = -3
Query: 38 SSSTTSASSSSTATTSPPSTTSTTTPSR 65
SSS+TS+SSSS+ + S S +S+ T S+
Sbjct: 879 SSSSTSSSSSSSCSCSSSSPSSSFTCSQ 796
Score = 26.2 bits (56), Expect = 9.9
Identities = 11/30 (36%), Positives = 20/30 (66%)
Frame = -3
Query: 24 NITNPSSSMTMTIPSSSTTSASSSSTATTS 53
+ +PSSS + T PSS + S S++S ++ +
Sbjct: 492 SFASPSSSSSFTTPSSPSPSTSTTSVSSNN 403
>TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~unknown
protein {Arabidopsis thaliana}, partial (46%)
Length = 2077
Score = 34.7 bits (78), Expect = 0.028
Identities = 17/45 (37%), Positives = 30/45 (65%)
Frame = -2
Query: 21 PMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPSR 65
P+ + SSS + ++PSSS++S+SS S++ + PP +TPS+
Sbjct: 561 PLTFSFSSSSSSSSSLPSSSSSSSSSKSSSPSLPPFAFFFSTPSK 427
Score = 27.7 bits (60), Expect = 3.4
Identities = 15/42 (35%), Positives = 28/42 (65%), Gaps = 5/42 (11%)
Frame = -2
Query: 28 PSSSMTMTIP-----SSSTTSASSSSTATTSPPSTTSTTTPS 64
P S +T P SSS++S+SSSS++ ++ P+ +S+++ S
Sbjct: 798 PKKSSYLTFPISSASSSSSSSSSSSSSSISNSPAFSSSSSSS 673
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.128 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,270,461
Number of Sequences: 36976
Number of extensions: 170765
Number of successful extensions: 7847
Number of sequences better than 10.0: 533
Number of HSP's better than 10.0 without gapping: 3589
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6461
length of query: 213
length of database: 9,014,727
effective HSP length: 92
effective length of query: 121
effective length of database: 5,612,935
effective search space: 679165135
effective search space used: 679165135
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC142096.4


