

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141865.9 - phase: 0 /pseudo
(547 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
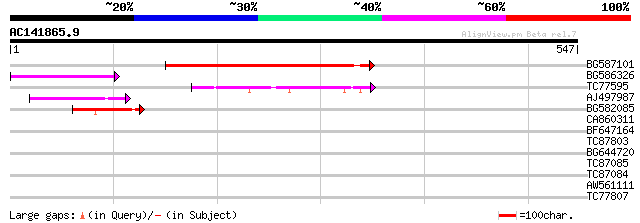
Score E
Sequences producing significant alignments: (bits) Value
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 244 6e-65
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 70 2e-12
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 59 4e-09
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 51 1e-06
BG582085 49 5e-06
CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product ... 40 0.003
BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis t... 37 0.025
TC87803 homologue to GP|1871187|gb|AAB63547.1| unknown protein {... 32 0.82
BG644720 30 2.4
TC87085 similar to PIR|T01511|T01511 hypothetical protein T10M13... 29 4.1
TC87084 similar to GP|2529683|gb|AAC62866.1| unknown protein {Ar... 29 4.1
AW561111 homologue to GP|13876340|gb| protocadherin gamma A7 {Mu... 29 5.3
TC77807 similar to PIR|T52601|T52601 squamosa promoter binding p... 28 6.9
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 244 bits (623), Expect = 6e-65
Identities = 120/202 (59%), Positives = 146/202 (71%)
Frame = +2
Query: 151 DLKHYYSDEPYLFKRG*DGIFRRCIPENEVSSILTHCHSSSYGGHASTQKASFKILHSGF 210
D++ Y DEP+L+K+ D I+ RC+ E E+ IL HCH S+Y GH + K KI +GF
Sbjct: 8 DVRRYLWDEPFLYKQCADNIYIRCVAEEEIPGILFHCHGSNYAGHFAVSKTVSKIQQAGF 187
Query: 211 WWPSLFKDVHLFISKCDKCQRTGNITKRNEMPLNNILEVEIFDVWVIDFIGPFPSSFVNQ 270
WWP++FKD H FISKCD CQR GNI+ RNEMP N ILEVE+FDVW IDF+GPFPSS+ N+
Sbjct: 188 WWPTMFKDAHSFISKCDPCQRQGNIS*RNEMPQNFILEVEVFDVWGIDFMGPFPSSYNNK 367
Query: 271 YILVAVDYVSKWVEVIASLTNDAQVVIKLFKKNHFSRFGVPRVVISDGGSQNILKNFCKN 330
YILVAVDYVSKWVE IAS TNDA VV+K+FK F RFGVPRVVISDGGS I K F K
Sbjct: 368 YILVAVDYVSKWVEAIASPTNDATVVVKMFKSVIFPRFGVPRVVISDGGSHFINKVFEKL 547
Query: 331 LE*GTKLQHPITHKLVDKWRSQ 352
L+ ++ + HK+ + Q
Sbjct: 548 LK-----KNGVRHKVATAYHPQ 598
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 70.5 bits (171), Expect = 2e-12
Identities = 39/106 (36%), Positives = 61/106 (56%)
Frame = +2
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I YAS+ L NY T + E+ AVV+A+ +R YL G+K+ ++TDH ++KY+ + +
Sbjct: 113 IAYASRQLRKHEGNYPTHDLEMAAVVFALKIWRSYLYGAKVQIHTDHKSLKYIFTQPELN 292
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDD 106
R + + + ++DL+I G N+VAD LSR R E DD
Sbjct: 293 LRQRRWMEFVADYDLDITYYPGKANLVADALSRRRVDVSAEREADD 430
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial
(14%)
Length = 1708
Score = 59.3 bits (142), Expect = 4e-09
Identities = 52/193 (26%), Positives = 88/193 (44%), Gaps = 15/193 (7%)
Frame = +2
Query: 176 PENEV-SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQ---- 230
P NE+ + ++ H S+ GH + + +I+ F+WP + V F+ CD C
Sbjct: 155 PLNELRTKLVQESHDSTAAGHPG-RNGTLEIVSRKFFWPGQSQTVRRFVRNCDVCGGIHI 331
Query: 231 -RTGNITKRNEMPLNNILEVEIFDVWVIDFIGPFPSSFV--NQYILVAVDYVSKWVEVIA 287
R +P+ N L ++ +DFI P + +QY+ V VD +SK V +
Sbjct: 332 WRQAKRGFLKPLPVPNRLHSDLS----MDFITSLPPTRGRGSQYLWVIVDRLSKSVTLEE 499
Query: 288 SLTNDAQVVIKLFKKNHFSRFGVPRVVISDGGSQ---NILKNFCKNLE*GTKL----QHP 340
T +A+ + F H+ G+P+ ++SD GS + FC+ L T+L HP
Sbjct: 500 MDTMEAEACAQRFLSCHYRFHGMPQSIVSDRGSNWVGRFWREFCR-LTGVTQLLSTSYHP 676
Query: 341 ITHKLVDKWRSQI 353
T ++W +I
Sbjct: 677 QTDGGTERWNQEI 715
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 51.2 bits (121), Expect = 1e-06
Identities = 27/98 (27%), Positives = 52/98 (52%), Gaps = 1/98 (1%)
Frame = -2
Query: 20 KELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAKPRLIQLIFLLQEFDLEIKD 79
K A+ +A + R Y++ + + IKY+ K R+ + LL E+D+E +
Sbjct: 635 KTCCALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRS 456
Query: 80 KKGVE-NVVADHLSRLRETNEDELPLDDSFPDDQLFLL 116
+K ++ +++ADHL+ + ED P+ FPD+++ L
Sbjct: 455 QKAIKGSILADHLA--HQPLEDYRPIKFDFPDEEIMYL 348
>BG582085
Length = 824
Score = 48.9 bits (115), Expect = 5e-06
Identities = 25/72 (34%), Positives = 45/72 (61%), Gaps = 2/72 (2%)
Frame = -3
Query: 61 PRLIQLIFLLQEFDLEIKDKK--GVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQ 118
P ++ I L QEFDLE + ++ G+ ++A R ++ ++E+ + + FP++QLF ++Q
Sbjct: 771 P*RLRCILLCQEFDLESRQERETGMW*LIAFPCLRNKQVTQNEISIKERFPNEQLFAISQ 592
Query: 119 TNAPWYADFVNF 130
PW+AD NF
Sbjct: 591 --RPWFADMTNF 562
Score = 31.6 bits (70), Expect = 0.82
Identities = 19/53 (35%), Positives = 27/53 (50%)
Frame = -2
Query: 44 YTDHSTIKYLLNKKDAKPRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRE 96
+ DH+T++Y K DAKP + + I +KG NVVA LS L +
Sbjct: 823 FIDHATLEYC*PKGDAKPVTA*VHSFMPGIRFRI*TRKGDRNVVAYRLSMLEK 665
Score = 29.6 bits (65), Expect = 3.1
Identities = 13/24 (54%), Positives = 15/24 (62%)
Frame = -1
Query: 174 CIPENEVSSILTHCHSSSYGGHAS 197
C+ E + IL H HSSSY GH S
Sbjct: 530 CVGEKVIKDIL*HHHSSSYMGHHS 459
>CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product {Drosophila
melanogaster}, partial (20%)
Length = 192
Score = 39.7 bits (91), Expect = 0.003
Identities = 20/40 (50%), Positives = 24/40 (60%)
Frame = +1
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSK 40
I YAS+ L A NY E+E LA ++AI FR YL G K
Sbjct: 64 IAYASRLLTAAERNYTVVERECLAAIWAIRNFRHYLHGPK 183
>BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana},
partial (6%)
Length = 469
Score = 36.6 bits (83), Expect = 0.025
Identities = 23/48 (47%), Positives = 30/48 (61%)
Frame = +2
Query: 413 LIGQLKTST*TPI*PVKKENFN*AS*KNLG*MPMRMPAFTRKELRNGM 460
L+G LK T P +K +* S*++ * PMRMP FT+KE +NGM
Sbjct: 41 LLGLLKPLTWITKPPEEKGY*S*MS*RS*D*TPMRMPRFTKKEQKNGM 184
>TC87803 homologue to GP|1871187|gb|AAB63547.1| unknown protein {Arabidopsis
thaliana}, partial (0%)
Length = 840
Score = 31.6 bits (70), Expect = 0.82
Identities = 16/35 (45%), Positives = 18/35 (50%)
Frame = +2
Query: 202 SFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNIT 236
S K S W SLF V ISKC C +T NI+
Sbjct: 110 SLKKFDSSIWCRSLFISVESRISKCSPCYKTMNIS 214
>BG644720
Length = 678
Score = 30.0 bits (66), Expect = 2.4
Identities = 16/30 (53%), Positives = 20/30 (66%), Gaps = 1/30 (3%)
Frame = -1
Query: 259 FIGPFPSSFVNQ-YILVAVDYVSKWVEVIA 287
F+ FP+ V+ YIL AV Y KWVEV+A
Sbjct: 93 FLDYFPNLLVDTWYILAAVYYF*KWVEVVA 4
>TC87085 similar to PIR|T01511|T01511 hypothetical protein T10M13.11 -
Arabidopsis thaliana, partial (12%)
Length = 639
Score = 29.3 bits (64), Expect = 4.1
Identities = 23/72 (31%), Positives = 37/72 (50%), Gaps = 5/72 (6%)
Frame = +3
Query: 34 QYLVGSKITVYT-DHSTIKYLLNKKDA----KPRLIQLIFLLQEFDLEIKDKKGVENVVA 88
+Y+ ++ + T + S I LN DA PRL Q + L L ++ K V +++
Sbjct: 339 KYIKDARSLIATQEQSEILSALNLLDAALAISPRLDQALELRARSLLYLRRFKDVADMLQ 518
Query: 89 DHLSRLRETNED 100
D++ LR TNED
Sbjct: 519 DYIPSLRMTNED 554
>TC87084 similar to GP|2529683|gb|AAC62866.1| unknown protein {Arabidopsis
thaliana}, partial (81%)
Length = 2109
Score = 29.3 bits (64), Expect = 4.1
Identities = 23/72 (31%), Positives = 37/72 (50%), Gaps = 5/72 (6%)
Frame = +3
Query: 34 QYLVGSKITVYT-DHSTIKYLLNKKDA----KPRLIQLIFLLQEFDLEIKDKKGVENVVA 88
+Y+ ++ + T + S I LN DA PRL Q + L L ++ K V +++
Sbjct: 222 KYIKDARSLIATQEQSEILSALNLLDAALAISPRLDQALELRARSLLYLRRFKDVADMLQ 401
Query: 89 DHLSRLRETNED 100
D++ LR TNED
Sbjct: 402 DYIPSLRMTNED 437
>AW561111 homologue to GP|13876340|gb| protocadherin gamma A7 {Mus musculus},
partial (1%)
Length = 435
Score = 28.9 bits (63), Expect = 5.3
Identities = 14/42 (33%), Positives = 23/42 (54%)
Frame = -1
Query: 13 VNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLL 54
V Y+T E+ L +Y + K +++ + K T H TI+ LL
Sbjct: 129 VYYSTKEQNLKQKIYQVMKLKKHYLWKKSESTTSHRTIQQLL 4
>TC77807 similar to PIR|T52601|T52601 squamosa promoter binding protein 1
[imported] - Arabidopsis thaliana, partial (55%)
Length = 2923
Score = 28.5 bits (62), Expect = 6.9
Identities = 15/44 (34%), Positives = 24/44 (54%)
Frame = +3
Query: 69 LLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQ 112
+L+EFD + K+ +A H R R+TN++ +P DDQ
Sbjct: 165 ILEEFD---EGKRSCRRRLAGHNKRRRKTNQEAVPNGSPTNDDQ 287
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.336 0.148 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,361,070
Number of Sequences: 36976
Number of extensions: 232306
Number of successful extensions: 1705
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1693
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1702
length of query: 547
length of database: 9,014,727
effective HSP length: 101
effective length of query: 446
effective length of database: 5,280,151
effective search space: 2354947346
effective search space used: 2354947346
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC141865.9


