

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141863.5 - phase: 0
(139 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
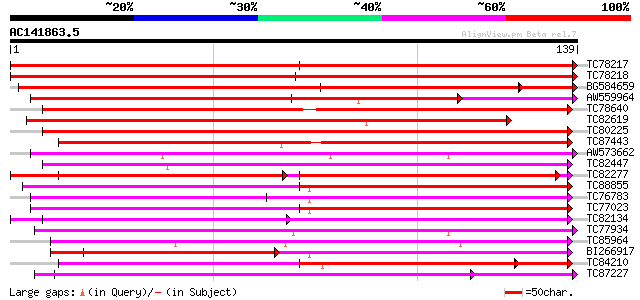
Score E
Sequences producing significant alignments: (bits) Value
TC78217 weakly similar to GP|20161870|dbj|BAB90783. hypothetical... 278 7e-76
TC78218 weakly similar to GP|12963415|gb|AAK11255.1 regulator of... 276 1e-75
BG584659 weakly similar to GP|12963415|gb| regulator of gene sil... 191 1e-49
AW559964 similar to GP|20161869|dbj hypothetical protein {Oryza ... 162 3e-41
TC78640 homologue to GP|13397927|emb|CAC34625. putative calmodul... 108 9e-25
TC82619 weakly similar to GP|22136314|gb|AAM91235.1 calmodulin-l... 106 3e-24
TC80225 similar to GP|14589311|emb|CAC43238. calcium binding pro... 105 6e-24
TC87443 similar to GP|13397927|emb|CAC34625. putative calmodulin... 105 8e-24
AW573662 weakly similar to PIR|D96689|D966 calmodulin-related pr... 97 2e-21
TC82447 similar to GP|14589311|emb|CAC43238. calcium binding pro... 92 7e-20
TC82277 weakly similar to PIR|A84532|A84532 probable calmodulin-... 92 7e-20
TC88855 PIR|A30900|MCBH calmodulin - barley, complete 87 2e-18
TC76783 calmodulin 1 [Medicago truncatula] 87 3e-18
TC77023 calmodulin 2 [Medicago truncatula] 87 3e-18
TC82134 similar to PIR|T07949|T07949 calcium binding protein - l... 87 3e-18
TC77934 similar to GP|9965747|gb|AAG10150.1| calmodulin-like MSS... 85 1e-17
TC85964 similar to PIR|D84841|D84841 calmodulin-like protein [im... 78 1e-15
BI266917 PIR|S01173|MC calmodulin [validated] - fruit fly (Droso... 77 3e-15
TC84210 homologue to PIR|T07122|T07122 calmodulin CAM5 - soybean... 73 4e-15
TC87227 similar to GP|17064926|gb|AAL32617.1 calcium-dependent p... 74 2e-14
>TC78217 weakly similar to GP|20161870|dbj|BAB90783. hypothetical protein
{Oryza sativa (japonica cultivar-group)}, partial (49%)
Length = 1027
Score = 278 bits (710), Expect = 7e-76
Identities = 139/139 (100%), Positives = 139/139 (100%)
Frame = +2
Query: 1 MKNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLE 60
MKNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLE
Sbjct: 212 MKNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLE 391
Query: 61 ELIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKH 120
ELIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKH
Sbjct: 392 ELIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKH 571
Query: 121 FDLDGDGLLSFDEFITMMQ 139
FDLDGDGLLSFDEFITMMQ
Sbjct: 572 FDLDGDGLLSFDEFITMMQ 628
Score = 45.4 bits (106), Expect = 7e-06
Identities = 22/68 (32%), Positives = 35/68 (51%)
Frame = +2
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E K +D + G ++P L++ L+ MGE + E + I+ D DGDG LS
Sbjct: 209 EMKNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSL 388
Query: 132 DEFITMMQ 139
+E I +M+
Sbjct: 389 EELIALME 412
>TC78218 weakly similar to GP|12963415|gb|AAK11255.1 regulator of gene
silencing {Nicotiana tabacum}, partial (58%)
Length = 793
Score = 276 bits (707), Expect = 1e-75
Identities = 138/139 (99%), Positives = 139/139 (99%)
Frame = +3
Query: 1 MKNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLE 60
MKNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLE
Sbjct: 261 MKNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLE 440
Query: 61 ELIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKH 120
ELIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKH
Sbjct: 441 ELIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKH 620
Query: 121 FDLDGDGLLSFDEFITMMQ 139
FDLDGDGLL+FDEFITMMQ
Sbjct: 621 FDLDGDGLLTFDEFITMMQ 677
Score = 44.7 bits (104), Expect = 1e-05
Identities = 22/69 (31%), Positives = 35/69 (49%)
Frame = +3
Query: 71 EEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLS 130
E K +D + G ++P L++ L+ MGE + E + I+ D DGDG LS
Sbjct: 255 EMMKNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLS 434
Query: 131 FDEFITMMQ 139
+E I +M+
Sbjct: 435 LEELIALME 461
>BG584659 weakly similar to GP|12963415|gb| regulator of gene silencing
{Nicotiana tabacum}, partial (34%)
Length = 677
Score = 191 bits (484), Expect(2) = 1e-49
Identities = 93/124 (75%), Positives = 105/124 (84%)
Frame = +3
Query: 3 NAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEEL 62
N FE VL YFDEDGDGK+SP ELR R+ + E LKE E+AIEA+DSDGDG LSLE+L
Sbjct: 171 NMEFERVLSYFDEDGDGKISPNELRSRMAKISGEFQLKEVEIAIEALDSDGDGLLSLEDL 350
Query: 63 IALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
IALME GGEE+KLKDLREAFEMYD E CG+ITPKSLKRMLKK+G+SKSI+ECK MIK FD
Sbjct: 351 IALMESGGEEEKLKDLREAFEMYDDEGCGYITPKSLKRMLKKLGDSKSIEECKVMIKRFD 530
Query: 123 LDGD 126
LDG+
Sbjct: 531 LDGE 542
Score = 40.0 bits (92), Expect = 3e-04
Identities = 18/63 (28%), Positives = 32/63 (50%)
Frame = +3
Query: 77 DLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFDEFIT 136
+ +D + G I+P L+ + K+ + E + I+ D DGDGLLS ++ I
Sbjct: 177 EFERVLSYFDEDGDGKISPNELRSRMAKISGEFQLKEVEIAIEALDSDGDGLLSLEDLIA 356
Query: 137 MMQ 139
+M+
Sbjct: 357 LME 365
Score = 21.2 bits (43), Expect(2) = 1e-49
Identities = 8/14 (57%), Positives = 12/14 (85%)
Frame = +1
Query: 126 DGLLSFDEFITMMQ 139
+G+LSF+EF MM+
Sbjct: 541 NGVLSFEEFRIMME 582
>AW559964 similar to GP|20161869|dbj hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (20%)
Length = 603
Score = 162 bits (411), Expect = 3e-41
Identities = 80/106 (75%), Positives = 90/106 (84%)
Frame = +3
Query: 6 FEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIAL 65
FE VL YFDEDGDGK+SP ELR R+ +G E LKE E+AIEA+DSDGDG LSL +LI L
Sbjct: 285 FERVLSYFDEDGDGKISPNELRSRMAKIGGEFQLKEVEIAIEALDSDGDGLLSLGDLITL 464
Query: 66 MEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSI 111
ME GGEE+KLKDLREAFEMYD+E CGFITPK LKRMLKK+G+SKSI
Sbjct: 465 MESGGEEEKLKDLREAFEMYDNEGCGFITPKKLKRMLKKLGDSKSI 602
Score = 43.9 bits (102), Expect = 2e-05
Identities = 24/76 (31%), Positives = 40/76 (52%), Gaps = 6/76 (7%)
Frame = +3
Query: 70 GEEQKLKDLREAFEM------YDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDL 123
G EQ + +++ E +D + G I+P L+ + K+G + E + I+ D
Sbjct: 243 GIEQYIVNMKREMEFERVLSYFDEDGDGKISPNELRSRMAKIGGEFQLKEVEIAIEALDS 422
Query: 124 DGDGLLSFDEFITMMQ 139
DGDGLLS + IT+M+
Sbjct: 423 DGDGLLSLGDLITLME 470
>TC78640 homologue to GP|13397927|emb|CAC34625. putative calmodulin-related
protein {Medicago sativa}, partial (84%)
Length = 1003
Score = 108 bits (269), Expect = 9e-25
Identities = 52/130 (40%), Positives = 83/130 (63%)
Frame = +2
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEE 68
+ FD++GDGK+S EL++ + +G + +E +E +D +GDGY+ L+E L
Sbjct: 437 IFNKFDKNGDGKISRTELKEMMTALGSKTTTEEVTRMMEELDRNGDGYIDLKEFGELHNG 616
Query: 69 GGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGL 128
GG+ K+LREAFEMYD +K G I+ K L +++++GE S+ +C+ MI + D D DG
Sbjct: 617 GGDT---KELREAFEMYDLDKNGLISAKELHAVMRRLGEKCSLGDCRKMIGNVDADADGN 787
Query: 129 LSFDEFITMM 138
++F+EF MM
Sbjct: 788 VNFEEFKKMM 817
>TC82619 weakly similar to GP|22136314|gb|AAM91235.1 calmodulin-like protein
{Arabidopsis thaliana}, partial (61%)
Length = 763
Score = 106 bits (264), Expect = 3e-24
Identities = 56/120 (46%), Positives = 74/120 (61%), Gaps = 1/120 (0%)
Frame = +3
Query: 5 GFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIA 64
G V +YFD DGDGK+S ELR +GE + +EAE I +D DGD L + I
Sbjct: 267 GLREVFKYFDGDGDGKISAYELRSYFGSIGEHMSHEEAERVINYLDGDGDNLLDFNDFIK 446
Query: 65 LMEEGGEEQKLKDLREAFEMYD-SEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDL 123
LM+ G KDLR+AFEM+ EK G ITPK L+RML+++G+ +S +EC MI FD+
Sbjct: 447 LMKGEGGRDDDKDLRKAFEMFVWEEKEGCITPKGLQRMLQRLGDDRSYEECVVMIDAFDI 626
>TC80225 similar to GP|14589311|emb|CAC43238. calcium binding protein
{Sesbania rostrata}, partial (87%)
Length = 704
Score = 105 bits (262), Expect = 6e-24
Identities = 52/130 (40%), Positives = 78/130 (60%)
Frame = +3
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEE 68
V FD +GDGK+S EL LR +G + E + +E +D+D DG+++L E A
Sbjct: 96 VFTRFDTNGDGKISVTELDNILRSLGSTVPKDELQRVMEDLDTDRDGFINLAEFAAFCRS 275
Query: 69 GGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGL 128
G + + +LREAF++YD +K G I+ L ++L +G S++EC MIK D DGDG
Sbjct: 276 GSADGDVSELREAFDLYDKDKNGLISATELCQVLNTLGMKCSVEECHTMIKSVDSDGDGN 455
Query: 129 LSFDEFITMM 138
++F+EF MM
Sbjct: 456 VNFEEFKKMM 485
>TC87443 similar to GP|13397927|emb|CAC34625. putative calmodulin-related
protein {Medicago sativa}, partial (69%)
Length = 918
Score = 105 bits (261), Expect = 8e-24
Identities = 52/128 (40%), Positives = 84/128 (65%), Gaps = 2/128 (1%)
Frame = +3
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIAL--MEEGG 70
FD++GDGK+S +EL++ L +G E +E + +E +D +GDG++ L+E E G
Sbjct: 51 FDKNGDGKISRSELKEMLLTLGSETTSEEVKRMMEELDQNGDGFIDLKEFADFHCTEPGK 230
Query: 71 EEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLS 130
+E +LR+AF++YD +K G I+ L +L K+GE S+++CK MI + D+DGDG ++
Sbjct: 231 DESS--ELRDAFDLYDLDKNGLISANELHAVLMKLGEKCSLNDCKKMISNVDVDGDGNVN 404
Query: 131 FDEFITMM 138
F+EF MM
Sbjct: 405 FEEFKKMM 428
>AW573662 weakly similar to PIR|D96689|D966 calmodulin-related protein
72976-72503 [imported] - Arabidopsis thaliana, partial
(29%)
Length = 615
Score = 97.1 bits (240), Expect = 2e-21
Identities = 54/143 (37%), Positives = 86/143 (59%), Gaps = 9/143 (6%)
Frame = +1
Query: 6 FEHVLRYFDEDGDGKVSPAELRQRLRIMGEE--ILLKEAEMAIEAMDSDGDGYLSLEELI 63
F + + D + DGK+S EL + L +G KEAE I +D +GDG++ ++E +
Sbjct: 115 FRQIFKLIDTNNDGKISTTELSEVLSCLGYNKCTASKEAESMIRVLDFNGDGFVDIDEFM 294
Query: 64 ALMEEGGEEQKLKD------LREAFEMYDSEKCGFITPKSLKRMLKKMG-ESKSIDECKA 116
+M+E G K K+ L +AF ++D +K G I+PK L+R+L +G E+ S+ +CK
Sbjct: 295 FVMKEDGNFGKGKEHDHDEYLMDAFLVFDIDKNGLISPKELRRVLVNLGCENCSLRDCKR 474
Query: 117 MIKHFDLDGDGLLSFDEFITMMQ 139
MIK D +GDG + F+EF +MM+
Sbjct: 475 MIKGVDKNGDGFVDFEEFRSMMK 543
>TC82447 similar to GP|14589311|emb|CAC43238. calcium binding protein
{Sesbania rostrata}, partial (79%)
Length = 660
Score = 92.0 bits (227), Expect = 7e-20
Identities = 49/131 (37%), Positives = 75/131 (56%), Gaps = 1/131 (0%)
Frame = +2
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEI-LLKEAEMAIEAMDSDGDGYLSLEELIALME 67
V FD +GDGK+S EL LR + ++ +E +E + +D DG+++L E A
Sbjct: 191 VFNNFDANGDGKISVNELETVLRTLRSDVPQQEELRRVMEELSTDRDGFINLSEFAAFCR 370
Query: 68 EGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDG 127
+ LR+AF++YD +K G I+ + L L ++G S++EC+ MIK D DGDG
Sbjct: 371 SDTTDGGDSALRDAFDLYDKDKNGLISTEELHLALDRLGMKCSVEECQDMIKSVDSDGDG 550
Query: 128 LLSFDEFITMM 138
++FDEF MM
Sbjct: 551 SVNFDEFKKMM 583
Score = 28.5 bits (62), Expect = 0.91
Identities = 11/29 (37%), Positives = 20/29 (68%)
Frame = +2
Query: 111 IDECKAMIKHFDLDGDGLLSFDEFITMMQ 139
+DE K + +FD +GDG +S +E T+++
Sbjct: 173 MDELKKVFNNFDANGDGKISVNELETVLR 259
>TC82277 weakly similar to PIR|A84532|A84532 probable calmodulin-like
protein [imported] - Arabidopsis thaliana, partial (91%)
Length = 780
Score = 92.0 bits (227), Expect = 7e-20
Identities = 44/126 (34%), Positives = 75/126 (58%)
Frame = +3
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGGEE 72
FD + DGK+S E + L+ +G E + E +D DGDG+++ EE + ++GG
Sbjct: 198 FDSNKDGKISQQEYKATLKSLGMEKSVNEVPNIFRVVDLDGDGFINFEEFMEAQKKGGGI 377
Query: 73 QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFD 132
+ L D++ AF +D G I+ + +K ML K+ E S+++C+ M++ D DGDG++ +
Sbjct: 378 RSL-DIQTAFRTFDKNGDGKISAEEIKEMLWKLEERCSLEDCRRMVRAVDTDGDGMVDMN 554
Query: 133 EFITMM 138
EF+ MM
Sbjct: 555 EFVAMM 572
Score = 52.8 bits (125), Expect = 5e-08
Identities = 23/64 (35%), Positives = 40/64 (61%)
Frame = +3
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
+ L +++ F+ +DS K G I+ + K LK +G KS++E + + DLDGDG ++F
Sbjct: 159 QPSLDEMKMVFDKFDSNKDGKISQQEYKATLKSLGMEKSVNEVPNIFRVVDLDGDGFINF 338
Query: 132 DEFI 135
+EF+
Sbjct: 339 EEFM 350
Score = 52.0 bits (123), Expect = 8e-08
Identities = 21/68 (30%), Positives = 42/68 (60%)
Frame = +3
Query: 1 MKNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLE 60
+++ + R FD++GDGK+S E+++ L + E L++ + A+D+DGDG + +
Sbjct: 375 IRSLDIQTAFRTFDKNGDGKISAEEIKEMLWKLEERCSLEDCRRMVRAVDTDGDGMVDMN 554
Query: 61 ELIALMEE 68
E +A+M +
Sbjct: 555 EFVAMMTQ 578
>TC88855 PIR|A30900|MCBH calmodulin - barley, complete
Length = 889
Score = 87.4 bits (215), Expect = 2e-18
Identities = 43/136 (31%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Frame = +1
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 106 AEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFL 285
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
LM ++ ++L+EAF ++D ++ GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 286 NLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 465
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+ +M
Sbjct: 466 VDGDGQINYEEFVKVM 513
Score = 51.2 bits (121), Expect = 1e-07
Identities = 21/67 (31%), Positives = 41/67 (60%)
Frame = +1
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
+ ++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 94 DDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 273
Query: 132 DEFITMM 138
EF+ +M
Sbjct: 274 PEFLNLM 294
>TC76783 calmodulin 1 [Medicago truncatula]
Length = 957
Score = 86.7 bits (213), Expect = 3e-18
Identities = 42/134 (31%), Positives = 79/134 (58%), Gaps = 1/134 (0%)
Frame = +3
Query: 6 FEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIAL 65
F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E + L
Sbjct: 156 FKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNL 335
Query: 66 MEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLD 124
M ++ ++L+EAF ++D ++ GFI+ L+ ++ +GE + +E MI+ D+D
Sbjct: 336 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 515
Query: 125 GDGLLSFDEFITMM 138
GDG ++++EF+ +M
Sbjct: 516 GDGQINYEEFVKVM 557
Score = 52.4 bits (124), Expect = 6e-08
Identities = 23/75 (30%), Positives = 44/75 (58%)
Frame = +3
Query: 64 ALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDL 123
A M + + ++ + +EAF ++D + G IT K L +++ +G++ + E + MI D
Sbjct: 114 ATMADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDA 293
Query: 124 DGDGLLSFDEFITMM 138
DG+G + F EF+ +M
Sbjct: 294 DGNGTIDFPEFLNLM 338
>TC77023 calmodulin 2 [Medicago truncatula]
Length = 999
Score = 86.7 bits (213), Expect = 3e-18
Identities = 42/134 (31%), Positives = 79/134 (58%), Gaps = 1/134 (0%)
Frame = +1
Query: 6 FEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIAL 65
F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E + L
Sbjct: 121 FKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNL 300
Query: 66 MEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLD 124
M ++ ++L+EAF ++D ++ GFI+ L+ ++ +GE + +E MI+ D+D
Sbjct: 301 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 480
Query: 125 GDGLLSFDEFITMM 138
GDG ++++EF+ +M
Sbjct: 481 GDGQINYEEFVKVM 522
Score = 52.4 bits (124), Expect = 6e-08
Identities = 21/67 (31%), Positives = 42/67 (62%)
Frame = +1
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
++++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 103 DEQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 282
Query: 132 DEFITMM 138
EF+ +M
Sbjct: 283 PEFLNLM 303
>TC82134 similar to PIR|T07949|T07949 calcium binding protein - loblolly
pine, partial (27%)
Length = 630
Score = 86.7 bits (213), Expect = 3e-18
Identities = 44/130 (33%), Positives = 74/130 (56%)
Frame = +1
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEE 68
V FD + DGK+S E + + + + I +A A MDSD DG++ +E + +
Sbjct: 79 VFEKFDTNKDGKISLEEYKAAAKSLDKGIGDPDAVKAFNVMDSDKDGFIDFKEFMEMFNG 258
Query: 69 GGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGL 128
+ K ++++ AF+++D G I+ + L ++ K++GES S+ CK M+K D DG GL
Sbjct: 259 ENNKIKEEEIKSAFQVFDINGDGKISAEELSQIFKRLGESCSLSACKKMVKGVDGDGHGL 438
Query: 129 LSFDEFITMM 138
+ +EF TMM
Sbjct: 439 IDLNEFTTMM 468
Score = 47.0 bits (110), Expect = 2e-06
Identities = 22/69 (31%), Positives = 35/69 (49%)
Frame = +1
Query: 1 MKNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLE 60
+K + + FD +GDGK+S EL Q + +GE L + ++ +D DG G + L
Sbjct: 271 IKEEEIKSAFQVFDINGDGKISAEELSQIFKRLGESCSLSACKKMVKGVDGDGHGLIDLN 450
Query: 61 ELIALMEEG 69
E +M G
Sbjct: 451 EFTTMMMNG 477
>TC77934 similar to GP|9965747|gb|AAG10150.1| calmodulin-like MSS3
{Arabidopsis thaliana}, partial (80%)
Length = 1325
Score = 84.7 bits (208), Expect = 1e-17
Identities = 47/138 (34%), Positives = 80/138 (57%), Gaps = 5/138 (3%)
Frame = +1
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
+ V + FD + DG+++ EL L +G I KE IE +D + DG + +EE L
Sbjct: 292 KRVFQMFDRNDDGRITKKELNDSLENLGIFIPDKELSQMIEKIDVNRDGCVDIEEFRELY 471
Query: 67 EE---GGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMG--ESKSIDECKAMIKHF 121
E +E++ +D+REAF ++D GFI+ L+ +L +G + +++++CK MI
Sbjct: 472 ESIMSERDEEEEEDMREAFNVFDQNGDGFISVDELRSVLVSLGLKQGRTVEDCKKMIGTV 651
Query: 122 DLDGDGLLSFDEFITMMQ 139
D+DG+GL+ + EF MM+
Sbjct: 652 DVDGNGLVDYKEFKQMMK 705
>TC85964 similar to PIR|D84841|D84841 calmodulin-like protein [imported] -
Arabidopsis thaliana, partial (69%)
Length = 825
Score = 77.8 bits (190), Expect = 1e-15
Identities = 46/134 (34%), Positives = 76/134 (56%), Gaps = 6/134 (4%)
Frame = +1
Query: 11 RYFDEDGDGKVSPAELRQRLRIMGEEILL-KEAEMAIEAMDSDGDGYLSLEELIALM--- 66
R D D DG VS EL L +G +E + + +DSDG G +S+E ++ +
Sbjct: 298 RLIDRDNDGVVSREELEAVLTRLGARPPTPEEIALMLSEVDSDGKGCISVETIMNRVGSG 477
Query: 67 -EEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESK-SIDECKAMIKHFDLD 124
G + ++LREAFE++D+++ G I+ + L R+ + +G+ + +++ECK MI D +
Sbjct: 478 SSSGSDPNPEEELREAFEVFDTDRDGRISAEELLRVFRAIGDERCTLEECKRMIAGVDKN 657
Query: 125 GDGLLSFDEFITMM 138
GDG + F EF MM
Sbjct: 658 GDGFVCFQEFSLMM 699
>BI266917 PIR|S01173|MC calmodulin [validated] - fruit fly (Drosophila
melanogaster), complete
Length = 576
Score = 76.6 bits (187), Expect = 3e-15
Identities = 37/121 (30%), Positives = 71/121 (58%), Gaps = 1/121 (0%)
Frame = +2
Query: 19 GKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGGEE-QKLKD 77
G ++ EL +R +G+ E + I +D+DG+G + E + +M ++ ++
Sbjct: 86 GTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLTMMARKMKDTDSEEE 265
Query: 78 LREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFDEFITM 137
+REAF ++D + GFI+ L+ ++ +GE + +E MI+ D+DGDG ++++EF+TM
Sbjct: 266 IREAFRVFDKDGNGFISAAELRHVMTNLGEKLTDEEVDEMIREADIDGDGQVNYEEFVTM 445
Query: 138 M 138
M
Sbjct: 446 M 448
Score = 50.1 bits (118), Expect = 3e-07
Identities = 23/56 (41%), Positives = 36/56 (64%)
Frame = +2
Query: 11 RYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
R FD+DG+G +S AELR + +GE++ +E + I D DGDG ++ EE + +M
Sbjct: 281 RVFDKDGNGFISAAELRHVMTNLGEKLTDEEVDEMIREADIDGDGQVNYEEFVTMM 448
>TC84210 homologue to PIR|T07122|T07122 calmodulin CAM5 - soybean, partial
(96%)
Length = 627
Score = 72.8 bits (177), Expect(2) = 4e-15
Identities = 38/114 (33%), Positives = 68/114 (59%), Gaps = 1/114 (0%)
Frame = +1
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGGEE 72
FD+DGDG ++ EL +R + + +E + I +D+DG+G + E + LM + +E
Sbjct: 40 FDKDGDGCITVEELATVIRSLDQNPTEEELQEMINEVDADGNGTIEFVEFLNLMAKKMKE 219
Query: 73 QKL-KDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDG 125
+DL+EAF+++D ++ G+I+ L+ ++ +GE + +E MIK DLDG
Sbjct: 220 TDADEDLKEAFKVFDKDQNGYISASELRHVMINLGEKLTDEEVDQMIKEADLDG 381
Score = 48.1 bits (113), Expect = 1e-06
Identities = 20/67 (29%), Positives = 42/67 (61%)
Frame = +1
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E ++ +++EAF ++D + G IT + L +++ + ++ + +E + MI D DG+G + F
Sbjct: 1 EDQIVEIKEAFCLFDKDGDGCITVEELATVIRSLDQNPTEEELQEMINEVDADGNGTIEF 180
Query: 132 DEFITMM 138
EF+ +M
Sbjct: 181 VEFLNLM 201
Score = 33.5 bits (75), Expect = 0.028
Identities = 15/47 (31%), Positives = 28/47 (58%)
Frame = +1
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDG 53
+ + FD+D +G +S +ELR + +GE++ +E + I+ D DG
Sbjct: 241 KEAFKVFDKDQNGYISASELRHVMINLGEKLTDEEVDQMIKEADLDG 381
Score = 23.5 bits (49), Expect(2) = 4e-15
Identities = 8/14 (57%), Positives = 12/14 (85%)
Frame = +2
Query: 125 GDGLLSFDEFITMM 138
GDG ++F+EF+ MM
Sbjct: 380 GDGQVNFEEFVKMM 421
>TC87227 similar to GP|17064926|gb|AAL32617.1 calcium-dependent protein kinase
{Arabidopsis thaliana}, partial (95%)
Length = 2319
Score = 73.9 bits (180), Expect = 2e-14
Identities = 41/133 (30%), Positives = 67/133 (49%)
Frame = +1
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
+ + + D D DG VS EL+ R ++ E +M IEA+D++G G L E +A+
Sbjct: 1291 KEMFKKMDTDNDGIVSIEELKVGFRNHQSQLAESEVQMFIEAVDNNGKGTLDYGEFVAIS 1470
Query: 67 EEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGD 126
+ L +AF +D + G+I P+ L+ L + G D + + D D D
Sbjct: 1471 LHLKRMANDEHLHKAFSYFDKDGNGYIEPEELRNALMEDGTDDCTDVANDIFQEVDTDKD 1650
Query: 127 GLLSFDEFITMMQ 139
G +S+DEF+ MM+
Sbjct: 1651 GKISYDEFVAMMK 1689
Score = 49.7 bits (117), Expect = 4e-07
Identities = 30/103 (29%), Positives = 49/103 (47%)
Frame = +1
Query: 12 YFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGGE 71
YFD+DG+G + P ELR L G + A + +D+D DG +S +E +A+M+ G
Sbjct: 1522 YFDKDGNGYIEPEELRNALMEDGTDDCTDVANDIFQEVDTDKDGKISYDEFVAMMKTG-- 1695
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDEC 114
D R+A Y + ++ K +K +G + C
Sbjct: 1696 ----TDWRKASRHYSRGRFNSLSLKLMKDGSLNLGND*CLSVC 1812
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.138 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,933,821
Number of Sequences: 36976
Number of extensions: 26849
Number of successful extensions: 360
Number of sequences better than 10.0: 99
Number of HSP's better than 10.0 without gapping: 234
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 298
length of query: 139
length of database: 9,014,727
effective HSP length: 86
effective length of query: 53
effective length of database: 5,834,791
effective search space: 309243923
effective search space used: 309243923
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 53 (25.0 bits)
Medicago: description of AC141863.5


