

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141436.5 + phase: 0 /pseudo
(364 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
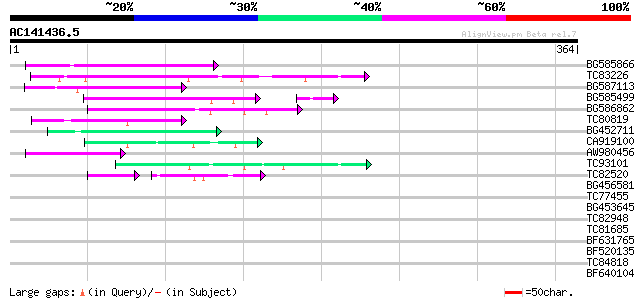
Score E
Sequences producing significant alignments: (bits) Value
BG585866 102 2e-22
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 87 1e-17
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 79 2e-15
BG585499 64 2e-11
BG586862 53 2e-07
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 48 5e-06
BG452711 45 3e-05
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 45 4e-05
AW980456 44 1e-04
TC93101 41 6e-04
TC82520 30 7e-04
BG456581 39 0.003
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 38 0.005
BG453645 37 0.016
TC82948 35 0.046
TC81685 34 0.078
BF631765 31 0.66
BF520135 30 1.1
TC84818 weakly similar to GP|18252325|gb|AAL66194.1 cytochrome P... 30 1.1
BF640104 30 1.5
>BG585866
Length = 828
Score = 102 bits (254), Expect = 2e-22
Identities = 45/124 (36%), Positives = 64/124 (51%)
Frame = +3
Query: 11 DIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETWQWIWKLHLPENLKHFIWLAMHG 70
D IW H+ G+Y+ KSGY W+ S + +W WIW+L +PE K F+WLA H
Sbjct: 348 DAYIWPHNSNGVYTAKSGYSWILSQTETV--NYNNSSWSWIWRLKIPEKYKFFLWLACHN 521
Query: 71 NLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLFNKDTNFNSQDFN 130
+PT + +++ + + C RCG ES H RDC S IW + F+ F+S
Sbjct: 522 AVPTLSLLNHRNMVNSAICSRCGEHEESFFHCVRDCRFSKIIWHKIGFSSPDFFSSSSVQ 701
Query: 131 WWFK 134
W K
Sbjct: 702 DWLK 713
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 86.7 bits (213), Expect = 1e-17
Identities = 71/253 (28%), Positives = 104/253 (41%), Gaps = 35/253 (13%)
Frame = +3
Query: 14 IWTHSPTGMYSVKSGYQ----WLTSNCLSFAGQTQQET--WQWIWKLHLPENLKHFIWLA 67
+W H+PTG+YSVKSGY W T ++ + ET W+ IW LH K +W
Sbjct: 3 MWMHNPTGIYSVKSGYNTLRTWQTQQ-INNTSTSSDETLIWKKIWSLHTIPRHKVLLWRI 179
Query: 68 MHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW--DDLLFNKDTNFN 125
++ +LP + + + C RC + E++ H F CPLS +W +L N D N
Sbjct: 180 LNDSLPVRSSLRKRGIQCYPLCPRCHSKTETITHLFMSCPLSKRVWFGSNLCINFDNLPN 359
Query: 126 SQDFNWWFKHYALALNGNIFVI----TCWVLWKARNHKVFNGNDWVCLSHIYSLHSDVVA 181
N ++ A+ I + LW ARN V L L D++
Sbjct: 360 PNFIN*LYE--AIL*KDECITI*IAAIIYNLWHARNLSV--------LEDQTILEMDIIQ 509
Query: 182 HISNDVI-----------------------QKPIK*VKWIPPLDNIIKVNVDGISLSNHG 218
SN + +P K KW P ++KVN D +L NHG
Sbjct: 510 RASNCISDYKQANTQAPPSMARTGYDPRSQHRPAKNTKWKRPNLGLVKVNTDA-NLQNHG 686
Query: 219 RSVFGGLIRNNNG 231
+ G +IR+ G
Sbjct: 687 KWGLGIIIRDEVG 725
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 79.3 bits (194), Expect = 2e-15
Identities = 37/111 (33%), Positives = 61/111 (54%), Gaps = 7/111 (6%)
Frame = -3
Query: 10 QDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQ-------TQQETWQWIWKLHLPENLKH 62
+D W +S +G YSVKSGY ++ +N ++ A Q + + +Q +WK + ++H
Sbjct: 429 EDSYSWEYSKSGHYSVKSGY-YVQTNIIAAANQRGTVDQPSLDDLYQRVWKYNTSPKVRH 253
Query: 63 FIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW 113
F+W + +LPT A +H++ D SC RCG E++ H CP + IW
Sbjct: 252 FLWRCISNSLPTAANMRSRHISKDGSCSRCGMESETVNHILFQCPYARLIW 100
>BG585499
Length = 792
Score = 63.5 bits (153), Expect(2) = 2e-11
Identities = 39/124 (31%), Positives = 60/124 (47%), Gaps = 10/124 (8%)
Frame = +3
Query: 48 WQWIWKLHLPENLKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCP 107
W+ +W P + F+WL HG + TN R+ ++C CG + E++LH DC
Sbjct: 225 WKMLWGWRGPHRTQTFMWLVAHGCILTNYRRSRWGTRVLATCPCCGNADETVLHVLCDCR 404
Query: 108 LSLAIWDDLL-FNKDTNFNSQD--FNWWFKHYALALNG-------NIFVITCWVLWKARN 157
+ +W L+ + TNF S D +W FK+ + NG F+ TCW +W RN
Sbjct: 405 PASQVWIRLVPSDWITNFFSFDDCRDWVFKNLSKRSNGVSKFKWQPTFMTTCWHMWTWRN 584
Query: 158 HKVF 161
+F
Sbjct: 585 KAIF 596
Score = 22.3 bits (46), Expect(2) = 2e-11
Identities = 11/27 (40%), Positives = 16/27 (58%)
Frame = +2
Query: 185 NDVIQKPIK*VKWIPPLDNIIKVNVDG 211
N +I+K + + W PLD K+N DG
Sbjct: 692 NLIIRKNVY-IAWKRPLDGWAKLNCDG 769
>BG586862
Length = 804
Score = 52.8 bits (125), Expect = 2e-07
Identities = 40/148 (27%), Positives = 64/148 (43%), Gaps = 10/148 (6%)
Frame = -1
Query: 51 IWKLHLPENLKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSL 110
+W + K F+W +H LP + + C RC + IE++ H F +C ++
Sbjct: 654 VWGIKTIPRHKSFLWRLLHNALPVKDELHKRGIRCSLLCPRCESKIETVQHLFLNCEVTQ 475
Query: 111 AIWDDLLFNKDTNFNSQ---DFNWWFKHYALALN-GNIFVITC--WVLWKARNHKVFNG- 163
W NF+S F+ W ++ L + I +T + +W ARN KVF
Sbjct: 474 KEWFGSQLG--INFHSSGVLHFHDWITNFILKNDEETIIALTALLYSIWHARNQKVFENI 301
Query: 164 ---NDWVCLSHIYSLHSDVVAHISNDVI 188
D V SLHS +A +S+ V+
Sbjct: 300 DVPGDVVIQRASSSLHSFKMAQVSDSVL 217
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 48.1 bits (113), Expect = 5e-06
Identities = 27/101 (26%), Positives = 44/101 (42%), Gaps = 2/101 (1%)
Frame = +2
Query: 15 WTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETWQWIWKLHLPENLKHFIWLAMHGNLPT 74
W P YSVK Y+++TS + + +W H+P + F+W + LPT
Sbjct: 530 WLLDPVNGYSVKVFYRYITST----GHISDRSLVDDVWHKHIPSKVSLFVWRLLRNRLPT 697
Query: 75 --NAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW 113
N G L ++++C ES H F C + ++W
Sbjct: 698 KDNLVHRGVLLATNAACVCGCVDSESTTHLFLHCNVFCSLW 820
Score = 28.5 bits (62), Expect = 4.3
Identities = 23/92 (25%), Positives = 38/92 (41%), Gaps = 17/92 (18%)
Frame = +1
Query: 87 SSCKRCGA---------------SIESLLHTFRDCPLSLAIWDDLLF--NKDTNFNSQDF 129
SSC +CG +++ L T C LA + +F +T ++
Sbjct: 724 SSCNKCGMCMWVR*LRIHYTPLFTLQRFLFTLVSCE-KLAWYSINVFW*ASNTFYSVCQN 900
Query: 130 NWWFKHYALALNGNIFVITCWVLWKARNHKVF 161
W K ++ L N WV+WK RN+++F
Sbjct: 901 GWHAKSFSFILQVNF-----WVIWKERNNRLF 981
>BG452711
Length = 672
Score = 45.4 bits (106), Expect = 3e-05
Identities = 29/112 (25%), Positives = 45/112 (39%)
Frame = +3
Query: 25 VKSGYQWLTSNCLSFAGQTQQETWQWIWKLHLPENLKHFIWLAMHGNLPTNAFRAGKHLT 84
V GY WL SF G+ E+ WI +L P N+K F+ + + +L
Sbjct: 225 VSKGYWWLMGVHTSFLGK---ES*NWISRLCAPSNIKLFL*QL*RDYVHFRSILLFCNLI 395
Query: 85 SDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLFNKDTNFNSQDFNWWFKHY 136
S + C C + +LH C + +W L T+ N + W K +
Sbjct: 396 SSNLCPICNQRSQDMLHALFSCTRAKDVWSFCLHELVTSSNQDNLITWMKEF 551
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 45.1 bits (105), Expect = 4e-05
Identities = 37/131 (28%), Positives = 51/131 (38%), Gaps = 17/131 (12%)
Frame = -2
Query: 49 QWIWKLHLPENLKHFIWLAMHGNLPT--NAFRAGKHLTSDSSC-KRCGASIESLLHTFRD 105
+ IW +P + F W + LPT N G T C CGA +ES H F
Sbjct: 590 EMIWHRQVPLKVSVFAWRLLRDRLPTKSNLIYRGVIPTEAGLCVSGCGA-LESAQHLFLS 414
Query: 106 CPLSLAIWDDL-----LFNKDTNFNSQDFNWWFKHYALALNGN---------IFVITCWV 151
C ++W + DTN S F + + GN I+++ WV
Sbjct: 413 CSYFASLWSLVRDWIGFVGVDTNVLSDHF----VQFVHSTGGNKASQSFLQLIWLLCAWV 246
Query: 152 LWKARNHKVFN 162
LW RN+ FN
Sbjct: 245 LWTERNNMCFN 213
>AW980456
Length = 779
Score = 43.9 bits (102), Expect = 1e-04
Identities = 20/64 (31%), Positives = 29/64 (45%)
Frame = -2
Query: 11 DIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETWQWIWKLHLPENLKHFIWLAMHG 70
D IW G Y VKS Y++ + + W IWKL +P +++ +W G
Sbjct: 208 DRLIWKDEKHGKYYVKSAYRFCVEELFDSSYLHRPGNWSGIWKLKVPPKVQNLVWRMCRG 29
Query: 71 NLPT 74
LPT
Sbjct: 28 CLPT 17
>TC93101
Length = 675
Score = 41.2 bits (95), Expect = 6e-04
Identities = 45/192 (23%), Positives = 71/192 (36%), Gaps = 28/192 (14%)
Frame = -1
Query: 69 HGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWD----DLLFNKDTNF 124
H LP + + C RC ++E+ H F C + +W + F ++
Sbjct: 672 HTTLPVRXELNKRGVNCPPLCPRCYFNLETTNHIFMSCERTQRVWFGSQLSIRFPDNSTI 493
Query: 125 NSQDFNWWFKHYALALNGNIFVITC--WVLWKARNHKVFNGNDWVCLSHIYS-------- 174
N D W F + I I+ + +W ARN +F N +V I
Sbjct: 492 NFSD--WLFDAISNQTEEIIIKISAITYSIWHARNKAIFE-NQFVSEDTIIQ*AQNSILA 322
Query: 175 --------------LHSDVVAHISNDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRS 220
L S +N ++ +W PL+NI+K N D +L GR
Sbjct: 321 YEQATIKPQNPNIVLSSLSATSNTNTTRRRSNVRSRWQKPLNNILKANCDA-NLQVQGRW 145
Query: 221 VFGGLIRNNNGD 232
G +IRN +G+
Sbjct: 144 GLGCIIRNADGE 109
>TC82520
Length = 833
Score = 30.0 bits (66), Expect(2) = 7e-04
Identities = 24/86 (27%), Positives = 37/86 (42%), Gaps = 13/86 (15%)
Frame = +2
Query: 92 CGASIESLLHTFRDCPLSLAIWDDLL--------FNKDTN-----FNSQDFNWWFKHYAL 138
CG S E+ H F C + ++W +L D F S + F H L
Sbjct: 317 CGDS-ETATHLFLHCDIFGSLWSHVLRWLHLLLVLPADIRQFFIQFTSMAGSPRFTHSFL 493
Query: 139 ALNGNIFVITCWVLWKARNHKVFNGN 164
+ ++ + WVLWK RN++VF +
Sbjct: 494 QI---MWFASVWVLWKKRNNRVFQNS 562
Score = 30.0 bits (66), Expect(2) = 7e-04
Identities = 9/33 (27%), Positives = 17/33 (51%)
Frame = +3
Query: 51 IWKLHLPENLKHFIWLAMHGNLPTNAFRAGKHL 83
+W+ ++P + F+W H LPT +H+
Sbjct: 186 VWQKNIPSKVSMFVWRLFHNRLPTKVNLMQRHV 284
>BG456581
Length = 683
Score = 38.9 bits (89), Expect = 0.003
Identities = 50/227 (22%), Positives = 77/227 (33%), Gaps = 29/227 (12%)
Frame = +2
Query: 21 GMYSVKSGYQWLTSNCLSFAGQTQQETW-QWIWKLHLPENLKHFIWLAMHGNLPTNAFRA 79
G ++K Y +LTS+ T W IW +P + W H LPT+ +
Sbjct: 8 GKLTIKLVYSFLTSH-------TSCAPWASTIWNSCIPPSHSFICWRLAHDRLPTDDNLS 166
Query: 80 GKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLFNKDTNFNSQDFNWWFKHYALA 139
+ S C C +E+ H F C + +W L + F
Sbjct: 167 SRGCALVSMCSFCLEQVETSDHLFLRCKFVVTLWSWLCSQLRVGLDFSSFKALLSSLPRH 346
Query: 140 LNGNI-------FVITCWVLWKARNHKVFNG------------NDWVCLSHIYSLHSDVV 180
+ + V +W ARN+ F+ + + LS S +
Sbjct: 347 CSSQVRDLYVAAVVHMVHSIWWARNNVRFSSAKVSAHAVQVRVHALIGLSGAVSTGKCIA 526
Query: 181 AHIS-NDVIQKP--------IK*VKWIPPLDNIIKVNVDGISLSNHG 218
A + DV + P + V W PP +K N DG L+N G
Sbjct: 527 ADAAILDVFRIPPHRRSMXEMVSVCWKPPSAPWVKGNTDGSXLNNSG 667
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 38.1 bits (87), Expect = 0.005
Identities = 27/102 (26%), Positives = 41/102 (39%), Gaps = 6/102 (5%)
Frame = -2
Query: 11 DIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQET---WQWIWKLHLPENLKHFIWLA 67
DI +W G++SV S Y +L N + E ++ +WK P + F W
Sbjct: 448 DIWVWKPDKEGVFSVNSCY-FLLQNLRLLEDRLSYEEEVIFRELWKSKAPAKVLAFSWTL 272
Query: 68 MHGNLPTNAFRAGKHLTSDSSCKR---CGASIESLLHTFRDC 106
+PT + L KR CG E+++H F C
Sbjct: 271 FLDRIPTMVNLGKRRLLRVEDSKRCVFCGCQDETVVHLFLHC 146
>BG453645
Length = 622
Score = 36.6 bits (83), Expect = 0.016
Identities = 23/88 (26%), Positives = 35/88 (39%), Gaps = 11/88 (12%)
Frame = +3
Query: 87 SSCKRCGASIESLLHTFRDCPLSLAIWDDLLFNKDTNFNSQDFNWWFKHYALALNGN--- 143
S+C C +E+ H F C ++ +W ++ D NF F H+ G
Sbjct: 27 STCVLCNLKVETTTHIFLHCDVASLVWFRVMKWFDVNFLIPPN--LFVHWECWSEGGSAN 200
Query: 144 --------IFVITCWVLWKARNHKVFNG 163
++ T WVLW RN +F G
Sbjct: 201 RVTKGHWLVWHTTIWVLWAKRNDLIFKG 284
>TC82948
Length = 705
Score = 35.0 bits (79), Expect = 0.046
Identities = 18/48 (37%), Positives = 24/48 (49%), Gaps = 2/48 (4%)
Frame = +3
Query: 86 DSSCKRCGASIESLLHTFRDCPLSLAIWD--DLLFNKDTNFNSQDFNW 131
D C C S ES H F +C +S+ IWD L N + + N+ D W
Sbjct: 435 DFMCSLCFKSFESTHHLFFECSVSVQIWDWFSNLINLNFHINNTDDLW 578
>TC81685
Length = 1706
Score = 34.3 bits (77), Expect = 0.078
Identities = 21/53 (39%), Positives = 28/53 (52%)
Frame = -2
Query: 185 NDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIRNNNGDWLLGF 237
NDV K V W P + K+N DG+ + R+ GGLI + G+WL GF
Sbjct: 208 NDVNAKQQITVSWTLPQSDWDKINSDGMCQGSL-RAGCGGLI*GDGGEWLCGF 53
>BF631765
Length = 498
Score = 31.2 bits (69), Expect = 0.66
Identities = 30/94 (31%), Positives = 42/94 (43%), Gaps = 6/94 (6%)
Frame = +1
Query: 144 IFVITCWVLWKARNHKVFNGNDWVCLSHIYSLHSDVVAHISNDVIQKPIK*VKWI----- 198
+F + W++WK RN KVFN D+ L S +V S + P+K +
Sbjct: 115 LFGTSLWLIWKHRNQKVFNNVDF-----DPPLVSALVEEFS--IANLPLKRGQVCVVPR* 273
Query: 199 -PPLDNIIKVNVDGISLSNHGRSVFGGLIRNNNG 231
PP IKVNV G G + +G + RN G
Sbjct: 274 QPPPFGSIKVNV-GAGCFGDGSTGWGFIARNAAG 372
>BF520135
Length = 202
Score = 30.4 bits (67), Expect = 1.1
Identities = 20/59 (33%), Positives = 26/59 (43%), Gaps = 3/59 (5%)
Frame = +3
Query: 51 IWKLHLPENLKHFIWLAMHGNLPT--NAFRAG-KHLTSDSSCKRCGASIESLLHTFRDC 106
+W+ + F W HG LPT N F+ G H + RC IES +H F C
Sbjct: 15 LWRKEDLSKVSIFAWRLFHGRLPTKANVFKRGIVHHDAHMCVTRC-RLIESDVHLFLHC 188
>TC84818 weakly similar to GP|18252325|gb|AAL66194.1 cytochrome P450 {Pyrus
communis}, partial (27%)
Length = 541
Score = 30.4 bits (67), Expect = 1.1
Identities = 23/78 (29%), Positives = 35/78 (44%)
Frame = +1
Query: 129 FNWWFKHYALALNGNIFVITCWVLWKARNHKVFNGNDWVCLSHIYSLHSDVVAHISNDVI 188
F WW+ + +AL GN +I V+W + D +CL I L ++A + + I
Sbjct: 268 FVWWWLNLCIALIGNCLIICYQVIWI*Q-----RSLDLLCLGQII*LPFPLIAFLVDIDI 432
Query: 189 QKPIK*VKWIPPLDNIIK 206
I+ VK P I K
Sbjct: 433 CV*IRLVKKYSPFHTIDK 486
>BF640104
Length = 344
Score = 30.0 bits (66), Expect = 1.5
Identities = 15/37 (40%), Positives = 22/37 (58%)
Frame = -1
Query: 195 VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIRNNNG 231
VKWI P +IK+N D +L++ G + RN+NG
Sbjct: 344 VKWIKPHQGVIKINCDA-NLTSEDVWGIGVITRNDNG 237
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.337 0.147 0.506
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,464,684
Number of Sequences: 36976
Number of extensions: 282131
Number of successful extensions: 2494
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 2449
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2480
length of query: 364
length of database: 9,014,727
effective HSP length: 97
effective length of query: 267
effective length of database: 5,428,055
effective search space: 1449290685
effective search space used: 1449290685
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC141436.5


