

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141414.10 - phase: 0 /pseudo
(1147 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
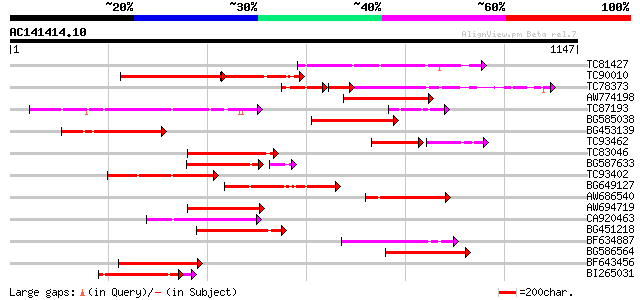
Sequences producing significant alignments: (bits) Value
TC81427 weakly similar to GP|11994216|dbj|BAB01338. disease resi... 245 6e-65
TC90010 similar to GP|5817349|gb|AAD52718.1| putative NBS-LRR ty... 233 4e-61
TC78373 weakly similar to GP|14348622|gb|AAK61318.1 NBS-LRR resi... 154 2e-56
AW774198 213 4e-55
TC87193 similar to GP|21616922|gb|AAM66423.1 NBS-LRR protein {Or... 197 1e-50
BG585038 weakly similar to GP|11994216|db disease resistance com... 190 2e-48
BG453139 similar to GP|5817349|gb| putative NBS-LRR type disease... 190 2e-48
TC93462 121 2e-48
TC83046 weakly similar to GP|10280097|emb|CAC09980. nucleotide s... 189 4e-48
BG587633 similar to GP|20385442|gb| resistance gene analog {Viti... 167 4e-46
TC93402 weakly similar to GP|16588610|gb|AAL26857.1 NBS-type put... 183 4e-46
BG649127 weakly similar to GP|11994217|db disease resistance com... 166 6e-41
AW686540 weakly similar to GP|15289776|db Lycopersicon esculentu... 165 8e-41
AW694719 similar to GP|11994217|db disease resistance comples pr... 164 2e-40
CA920463 weakly similar to GP|2852684|gb|A resistance protein ca... 161 2e-39
BG451218 similar to GP|11994217|db disease resistance comples pr... 155 8e-38
BF634887 weakly similar to GP|10280097|em nucleotide sequence of... 152 7e-37
BG586564 weakly similar to GP|14348622|gb NBS-LRR resistance-lik... 151 2e-36
BF643456 weakly similar to GP|14348622|gb NBS-LRR resistance-lik... 148 1e-35
BI265031 weakly similar to GP|13872974|db putative NBS-LRR type ... 127 1e-32
>TC81427 weakly similar to GP|11994216|dbj|BAB01338. disease resistance
comples protein {Arabidopsis thaliana}, partial (6%)
Length = 1253
Score = 245 bits (626), Expect = 6e-65
Identities = 151/404 (37%), Positives = 213/404 (52%), Gaps = 23/404 (5%)
Frame = +3
Query: 583 LLSEIRPLRVLSLSYYLNITDLPQYLGNLIHLRYLDLSNTKIQRLPYETCKLYNLQTLLL 642
LL + +PLRV SLS Y IT LP +G+L+HLRYLDLS T I LP C LYNL LLL
Sbjct: 75 LLKKPKPLRVFSLSEY-PITLLPSSIGHLLHLRYLDLSRTPITSLPDSICNLYNLXALLL 251
Query: 643 SRCWLLIELPEDMGNLINLRHLDICGTNLKYMPSQIAKLQNLQTLSAFIVSKSQDGLKVG 702
C L LP LINLR LDI G+ +K MP+ + KL++LQ+L F+VS + G VG
Sbjct: 252 VGCADLTLLPTKTSKLINLRQLDISGSGIKKMPTNLGKLKSLQSLPRFVVS-NDGGSNVG 428
Query: 703 ELKNFTNLQGKLSISKLQNVTDPFEAFRANLKSKEKVDELSLEWDYGATLDTQIERLVLE 762
EL L+G LSI L+NV A A LK K+ + E+ +W T + E ++ +
Sbjct: 429 ELGEMLELRGSLSIVNLENVLLKEXASNAGLKRKKYLHEVEFKWTT-PTHSQESENIIFD 605
Query: 763 QLQPPSSLKKLTIKSYGGTSFPNWFGDSSFAHMVYLCISDCDHCWSLPPLGQLLGLRELY 822
L P +LK+L I ++GG FPNW G +S + M+ L + +C +C SLP LGQL LRE+Y
Sbjct: 606 MLXPHRNLKRLKINNFGGEKFPNWLGSNSGSTMMSLYLDECGNCLSLPSLGQLSNLREIY 785
Query: 823 ISGMKSVKIVGAEFYGSSSSSSLFQPFPSLQVLRFRDMPEWEDWN--------------- 867
I+ + ++ VG EFYG+ F+ F SL++++F+DM WE+W+
Sbjct: 786 ITSVTRLQKVGPEFYGNG-----FEAFSSLRIIKFKDMLNWEEWSVNNQSGSEGFTLLQE 950
Query: 868 --------LIGDTTTDFPNLLHLSLKDCPKLKGTLPINQISSTFELSGCPLLFPNSMLYF 919
LIG + P+L L + C L T+P ++SGC
Sbjct: 951 LYIENCPKLIGKLPGNLPSLDKLVITSCQTLSDTMPCVPRLRELKISGCEAFVS------ 1112
Query: 920 TENIPTNFHSSLVLNCTNLILDLTLSRIPSSASFPRDGLPTTLR 963
S ++ C + + + +S PS S P D + TL+
Sbjct: 1113--------LSEQMMKCNDCLQTMAISNCPSLVSIPMDCVSRTLK 1220
>TC90010 similar to GP|5817349|gb|AAD52718.1| putative NBS-LRR type disease
resistance protein {Pisum sativum}, partial (88%)
Length = 1107
Score = 233 bits (593), Expect = 4e-61
Identities = 115/217 (52%), Positives = 152/217 (69%), Gaps = 2/217 (0%)
Frame = +3
Query: 224 EVEDNFDLKAWAYISKDFDVCRVTKVILESITFKPVDTNNLNILQVELQQSLRNRRFLLV 283
EV+ +FDLKAW +S+DFD+ RVTK +LES+T D+ NL++L+VEL++ R +RFL V
Sbjct: 12 EVQQHFDLKAWVCVSEDFDIMRVTKSLLESVTSTTWDSKNLDVLRVELKKISREKRFLFV 191
Query: 284 LDDIWDGSYVDWNNLMDIFSAGEKGSRIIVTTRDESVARSMQTSFPIYHLLPLASEDCWS 343
DD+W+ +Y DWN L F G+ GS +I+TTR + VA T FPI+ L L++EDCWS
Sbjct: 192 FDDLWNDNYNDWNELASPFINGKPGSMVIITTRQQKVAEVAHT-FPIHKLKLLSNEDCWS 368
Query: 344 LLAKHAFGPYNCRNRSN--LEFIGKEIVKKCDGLPIAAVALGGLLRSELSENRWNKVLKS 401
LL+KHA G + SN LE G++I +KC GLPIAA +GGLLRS++ W +L S
Sbjct: 369 LLSKHALGSDEFHHSSNTALEETGRKIARKCGGLPIAAKTIGGLLRSKVDITEWTSILNS 548
Query: 402 NIWDLPNVKVLPALLLSYHHLPSPLKQCFTYCSIFPK 438
NIW+L N +LPAL LSY +LPS LK+CF YCSIF K
Sbjct: 549 NIWNLRNDNILPALHLSYQYLPSHLKRCFAYCSIFSK 659
Score = 115 bits (287), Expect = 1e-25
Identities = 72/171 (42%), Positives = 104/171 (60%), Gaps = 1/171 (0%)
Frame = +1
Query: 427 KQCFTYCSIFPKNFILEKQMVVQLWIAEGFVHQSKSGKTMEEVADEYFDELVSRSLIHRW 486
K FPK++ L+++ +V LW+AEGF+ S+ GK MEE+ D+ F EL+SRSLI +
Sbjct: 625 KDALHIVQFFPKDYPLDRKELVLLWMAEGFLDCSQRGKKMEELGDDCFAELLSRSLIQQL 804
Query: 487 SVND-CVHYKMHDLINDLATMVSSSYCIRYGDRKLQESVERVRHLS*NKGKYNSFNKFDS 545
S +D + MHDL+NDLAT VS C R + + E VRH S N+ Y+ F KF+
Sbjct: 805 SDDDRGEKFVMHDLVNDLATFVSGKSCCRL---ECGDIPENVRHFSYNQENYDIFMKFEK 975
Query: 546 LYESKRLRTFISLPVRLEWLPDQHYAKYFLSNKVLHDLLSEIRPLRVLSLS 596
L+ K LR+F+ + + + W + +LS KV++DLL + LRVLSLS
Sbjct: 976 LHNFKCLRSFLFICL-MTWRDN------YLSFKVVNDLLPSHKRLRVLSLS 1107
>TC78373 weakly similar to GP|14348622|gb|AAK61318.1 NBS-LRR resistance-like
protein J78 {Phaseolus vulgaris}, partial (7%)
Length = 1463
Score = 154 bits (389), Expect(3) = 2e-56
Identities = 131/423 (30%), Positives = 188/423 (43%), Gaps = 9/423 (2%)
Frame = +1
Query: 690 FIVSKSQDGLKVGELKNFTNLQGKLSISKLQNVTDPFEAFRANLKSKEKVDELSLEWDYG 749
F+V + Q G + EL N +LQG+L IS +++V DP +A ANLK K+ V+EL Y
Sbjct: 460 FVVGE-QSGSDIKELGNLNHLQGELCISGMEHVIDPVDAAGANLKDKKHVEELIWSGSYK 636
Query: 750 ATLDTQIERLVLEQLQPPSSLKKLTIKSYGGTSFPNWFGDSSFAHMVYLCISDCDHCWSL 809
+ + E V E LQP SSLK+L I Y G F NW ++V + ++ C C L
Sbjct: 637 FNTNGR-ESDVFEALQPNSSLKRLIISHYKGNRFANWMRGCDLPNLVSIRLNLCALCSEL 813
Query: 810 PPLGQLLGLRELYISGMKSVKIVGAEFYGSSSSSSLFQPFPSLQVLRFRDMPEWEDWNLI 869
PPLGQL L+E+ ISG ++I+G EFYG++S++ PF SL++L F M EWE+W+ +
Sbjct: 814 PPLGQLPCLKEISISGCDKIRIIGKEFYGNNSTN---VPFRSLEILHFDSMSEWEEWSHL 984
Query: 870 GDTTTDFPNLLHLSLKDCPKLKGTLPINQIS-STFELSGCPLLFPNSMLYFTENIPTNFH 928
FP L LS+K CPKLK LP + S E+ C L
Sbjct: 985 ----EGFPLLKELSIKKCPKLKRALPQHLPSLQKLEIKDCKKL----------------- 1101
Query: 929 SSLVLNCTNLILDLTLSRIPSSASFPRDGLPTTLRIVVIH*HLLL*ALFLCSKVLELCVV 988
+L+ C N+I +L + R C ++L
Sbjct: 1102 EALIPKCDNMI-ELDIKR--------------------------------CDRIL----- 1167
Query: 989 NI*NSFQLRKIQHSLPINLRSLNVCSRGSSWTRAISEWILQRLTFLTTLRIGGDDLLNAL 1048
+ LP NL+SL +C + +T + L + FL L+ +N
Sbjct: 1168 -----------VNELPTNLKSLFLCD--NQYTEFSVDQNLINILFLEVLKFDFRGCVNC- 1305
Query: 1049 MEMNVPLLPNSLVSLYIYNLLDVKCLDGKW--------LQHLTSLENLEIAYCRKLESLP 1100
+ L YN L + G W L TSL L + +C +LES P
Sbjct: 1306 ----------PSLDLRCYNSLRDLSIKG-WHSSSLPLELHLFTSLSALRLYHCPELESFP 1452
Query: 1101 EEG 1103
G
Sbjct: 1453 MGG 1461
Score = 58.9 bits (141), Expect(3) = 2e-56
Identities = 37/94 (39%), Positives = 58/94 (61%), Gaps = 1/94 (1%)
Frame = +2
Query: 550 KRLRTFISLPVRLEWLPDQHYAKYFL-SNKVLHDLLSEIRPLRVLSLSYYLNITDLPQYL 608
K LR+F+++P + YF+ SN + DL S+++ LR+LS + + +L +
Sbjct: 59 KGLRSFLAVP-------GGYRKDYFMTSNNLQRDLFSKLKYLRMLSFCGF-KLKELSGEI 214
Query: 609 GNLIHLRYLDLSNTKIQRLPYETCKLYNLQTLLL 642
GNL LRYL+L+ + I+RLP CKLY L+TL+L
Sbjct: 215 GNLKLLRYLNLTASLIERLPDSICKLYKLETLIL 316
Score = 46.6 bits (109), Expect(3) = 2e-56
Identities = 21/52 (40%), Positives = 33/52 (63%)
Frame = +3
Query: 645 CWLLIELPEDMGNLINLRHLDICGTNLKYMPSQIAKLQNLQTLSAFIVSKSQ 696
C+ L +LP ++LRHL + G N+K MP +I +L +LQTLS F +++
Sbjct: 324 CFELTKLPSKFYKFVSLRHLHLEGCNIKKMPKKIGRLNHLQTLSDFCCGRTE 479
Score = 30.8 bits (68), Expect = 3.1
Identities = 17/37 (45%), Positives = 24/37 (63%)
Frame = +1
Query: 1080 QHLTSLENLEIAYCRKLESLPEEGLPSSLSVLTIKKC 1116
QHL SL+ LEI C+KLE+L + ++ L IK+C
Sbjct: 1051 QHLPSLQKLEIKDCKKLEALIPK--CDNMIELDIKRC 1155
>AW774198
Length = 554
Score = 213 bits (541), Expect = 4e-55
Identities = 110/184 (59%), Positives = 135/184 (72%), Gaps = 1/184 (0%)
Frame = +2
Query: 675 PSQIAKLQNLQTLSAFIVSKSQDGLKVGELKNFTNLQGKLSISKLQNVTDPFEAFRANLK 734
P QIAKL+NL TLS F++SK GLKV EL F +L GKL IS+LQNV DP EAF+ANL
Sbjct: 5 PVQIAKLENLHTLSDFVISKHNGGLKVAELGKFPHLHGKLYISQLQNVNDPSEAFQANLN 184
Query: 735 SKEKVDELSLEWDYGAT-LDTQIERLVLEQLQPPSSLKKLTIKSYGGTSFPNWFGDSSFA 793
+KE++ EL+LEWD G+T LD+Q VL+ LQP ++LK LTIK YGGTSFPNW GD+ F
Sbjct: 185 TKERIHELALEWDCGSTFLDSQAHSAVLKHLQPSTNLKSLTIKGYGGTSFPNWLGDNVFG 364
Query: 794 HMVYLCISDCDHCWSLPPLGQLLGLRELYISGMKSVKIVGAEFYGSSSSSSLFQPFPSLQ 853
+M+YL IS+C +C LP LGQL L+EL I + S+K VG EFYG+ S FQPFPSL+
Sbjct: 365 NMLYLRISNCVNCLWLPSLGQLGNLKELVIDSLLSIKSVGTEFYGNDGHPS-FQPFPSLE 541
Query: 854 VLRF 857
L F
Sbjct: 542 TLHF 553
>TC87193 similar to GP|21616922|gb|AAM66423.1 NBS-LRR protein {Oryza sativa
(japonica cultivar-group)}, partial (26%)
Length = 3119
Score = 197 bits (502), Expect = 1e-50
Identities = 153/489 (31%), Positives = 243/489 (49%), Gaps = 19/489 (3%)
Frame = +3
Query: 41 LKKLKITLLSLQAVLNDAEEKQITNPAVKEWLDELTHVVFDADDLLDEINTEALRWKIEG 100
L++LK T+ S++AVL DAE+KQ + AV+ W+ L ++ ADDL+DE E + K +
Sbjct: 180 LERLKKTVESIKAVLLDAEDKQEQSHAVQLWVRRLNDLLLPADDLIDEFFIEDMIHKRDK 359
Query: 101 CPQSQTIIDQVIYLYSSPFKRFPEAIYSRIHELFQRLEHFALQKDILQLKQGV-----SN 155
+++ + QV + +S F + I ++ + + L+L V +N
Sbjct: 360 AHKNK--VKQVFHSFSPSRTAFRRKMAHEIEKIQRSFKDVEKDMSYLKLNSNVVVVAKTN 533
Query: 156 SIWYGNPTSSVVVDESSICGRDDEKKKLKEFLLLEDGSVSGSKIGVISIVGMGGLGKTTL 215
++ T S V+ ES I GR+D+K K+ L S + +++IVG+GGLGKT L
Sbjct: 534 NV--RRETCSYVL-ESEIIGREDDKNKIISLLRQ---SHEHQNVSLVAIVGIGGLGKTAL 695
Query: 216 AKLLFNDHEVEDNFDLKAWAYISKDFDVCRVTKVILESITFKPVDTNNLNILQVELQQSL 275
A+L++ D EV++ F+ W +S +FD + K ++ S+T + V L LQ LQ +L
Sbjct: 696 AQLVYKDGEVKNLFEKHMWVCVSDNFDFKTILKNMVASLTKEDVVNKTLQELQSMLQVNL 875
Query: 276 RNRRFLLVLDDIWDGSYVDWNNLMDIFSAGEKGSRIIVTTRDESVARSMQTSFPIYHLLP 335
+R+LLVLDD+W+ S+ W+ L G +GS++++TT + VA M S + L
Sbjct: 876 TGKRYLLVLDDVWNESFEKWDQLRPYLMCGAQGSKVVMTTCSKIVADRMGVS-DQHVLRG 1052
Query: 336 LASEDCWSLLAKHAFGPYNCRNRSNLEFIGKEIVKKCDGLPIAAVALGGLLRSELSENRW 395
L E W L FG LE IGK+I +KC G+P+A +LGG+LRSE E+ W
Sbjct: 1053LTPEKSWVLFKNIVFGDVTVGVNQPLESIGKKIAEKCKGVPLAIRSLGGILRSESKESEW 1232
Query: 396 NKVLKSNIWDLPNVKVLPALLLSYHHLPSPLKQCFTYCSIFPKNFILEKQMVVQLWIAEG 455
VL+ L N + +++ + P T F L + L I EG
Sbjct: 1233INVLQGEC--LENCVIRENSIIAGPKIELPELITSTKAM-----FCLLLFISSGLGI*EG 1391
Query: 456 FVHQSKSG---------KTMEE-----VADEYFDELVSRSLIHRWSVNDCVHYKMHDLIN 501
V+ + G K M E + + +E + S +KMHDL++
Sbjct: 1392*VNSNVDGTRLSWLFS*KAMHERCW*SICKNFLEEFILPRCKFE*SWVMVTGFKMHDLMH 1571
Query: 502 DLATMVSSS 510
DLAT V+ +
Sbjct: 1572DLATQVAGN 1598
Score = 55.8 bits (133), Expect = 9e-08
Identities = 37/124 (29%), Positives = 60/124 (47%)
Frame = +2
Query: 767 PSSLKKLTIKSYGGTSFPNWFGDSSFAHMVYLCISDCDHCWSLPPLGQLLGLRELYISGM 826
PS +KL+I+ G S NW S +++ LC+ +C LPPL +L L+ L + +
Sbjct: 2042 PSGFRKLSIQQPKGLSLSNWL--SPVTNIIELCLYNCQDLRYLPPLERLPFLKSLDLRWL 2215
Query: 827 KSVKIVGAEFYGSSSSSSLFQPFPSLQVLRFRDMPEWEDWNLIGDTTTDFPNLLHLSLKD 886
++ + +Y F FPSL++L F + + + W +GD D + HL L
Sbjct: 2216 NQLEYI---YYEEPILHESF--FPSLEILNFFECWKLKGWRRMGDDLNDINSSHHLFLPH 2380
Query: 887 CPKL 890
P L
Sbjct: 2381 FPCL 2392
Score = 40.0 bits (92), Expect = 0.005
Identities = 38/143 (26%), Positives = 64/143 (44%), Gaps = 8/143 (5%)
Frame = +1
Query: 1007 LRSLNVCSRGSSWTRAISEWILQRLTFLTTLRIGGDDLLNALMEMNVPLLPNSLVSLYIY 1066
L+S ++ +W L+ LT L L + D L + E+ + L+ L
Sbjct: 2554 LKSFHIIETIMGMENVPKDW-LKNLTSLENLDL--DSLSSQQFEVIEMWFKDDLICLPSL 2724
Query: 1067 NLLDVKCLDGK----WLQHLTSLENLEIAYCRKLESLPEEGLP--SSLSVLTIKKCPL-- 1118
+ ++ K W+ +++SL++L + C L LPE G+P ++L L I CP+
Sbjct: 2725 QKIQIRTCRLKALPDWICNISSLQHLAVYGCESLVDLPE-GMPRLTNLHTLEIIGCPISY 2901
Query: 1119 LEASCKSNGGKEWPKISHIPCLI 1141
G W KI+HIP +I
Sbjct: 2902 YYDEFLRKTGATWSKIAHIPKII 2970
Score = 35.4 bits (80), Expect = 0.13
Identities = 29/77 (37%), Positives = 39/77 (49%), Gaps = 2/77 (2%)
Frame = +2
Query: 607 YLGNLIHLRYLDLSNTKI-QRLPYETCKLYNLQTLLLSRCWLLIELPEDMGNLINLRHLD 665
++ L HLR+LDL N + L L LQT+ L+ +++ PE + LINLRHL
Sbjct: 1838 FIEKLKHLRHLDLRNYDSGESLSKSIGNLVCLQTIKLNDS--VVDSPEVVSKLINLRHLT 2011
Query: 666 ICGTNLK-YMPSQIAKL 681
I K PS KL
Sbjct: 2012 INHVTFKDKTPSGFRKL 2062
>BG585038 weakly similar to GP|11994216|db disease resistance comples protein
{Arabidopsis thaliana}, partial (5%)
Length = 528
Score = 190 bits (483), Expect = 2e-48
Identities = 99/176 (56%), Positives = 124/176 (70%)
Frame = +3
Query: 611 LIHLRYLDLSNTKIQRLPYETCKLYNLQTLLLSRCWLLIELPEDMGNLINLRHLDICGTN 670
L+ LRYLDLS T I+ LP TC LYNLQTL L+RC L ELP + G LINLRHLDI TN
Sbjct: 3 LVELRYLDLSFTGIKSLPNATCNLYNLQTLNLTRCENLTELPPNFGKLINLRHLDISETN 182
Query: 671 LKYMPSQIAKLQNLQTLSAFIVSKSQDGLKVGELKNFTNLQGKLSISKLQNVTDPFEAFR 730
+K MP QI L NLQTL+ F V K GL + E+ F NL+GKL I LQNV D EA+
Sbjct: 183 IKEMPMQIVGLNNLQTLTVFSVGKQDTGLSLKEVCKFPNLRGKLCIKNLQNVIDAIEAYD 362
Query: 731 ANLKSKEKVDELSLEWDYGATLDTQIERLVLEQLQPPSSLKKLTIKSYGGTSFPNW 786
N+++KE ++EL L+W T D++IE+ VL+ LQP +L+KL+I+ YGGTSFP+W
Sbjct: 363 VNMRNKEDIEELELQWS-KQTEDSRIEKDVLDMLQPSFNLRKLSIRLYGGTSFPSW 527
>BG453139 similar to GP|5817349|gb| putative NBS-LRR type disease resistance
protein {Pisum sativum}, partial (65%)
Length = 638
Score = 190 bits (483), Expect = 2e-48
Identities = 102/212 (48%), Positives = 148/212 (69%)
Frame = -2
Query: 105 QTIIDQVIYLYSSPFKRFPEAIYSRIHELFQRLEHFALQKDILQLKQGVSNSIWYGNPTS 164
+ I QV SSPF+ I S++ + +RL+ FA QKDIL L Q VS ++ P+S
Sbjct: 637 ENITGQVWNFLSSPFQNLYGEINSQMKSMIERLQLFAQQKDILGL-QTVSGRVFLRTPSS 461
Query: 165 SVVVDESSICGRDDEKKKLKEFLLLEDGSVSGSKIGVISIVGMGGLGKTTLAKLLFNDHE 224
S+V D+S + GR +K+KL +LL D S I V+SI GMGG+GKT LA+LL+ND +
Sbjct: 460 SLV-DQSFMVGRKSDKEKLTN-MLLSDVSTVNDNINVVSIFGMGGVGKTALAQLLYNDKD 287
Query: 225 VEDNFDLKAWAYISKDFDVCRVTKVILESITFKPVDTNNLNILQVELQQSLRNRRFLLVL 284
V++ FD+KAW +S++FDV RVT+ +LESIT +++NL+ L+V+L+Q+L+ +RFL VL
Sbjct: 286 VQERFDVKAWVCLSEEFDVLRVTRTLLESITSNVQESDNLDFLRVKLKQNLKGKRFLFVL 107
Query: 285 DDIWDGSYVDWNNLMDIFSAGEKGSRIIVTTR 316
DD+W+ SY DW++L+ F GE GSR+I+TTR
Sbjct: 106 DDMWNDSYNDWHHLVTPFIDGESGSRVIITTR 11
>TC93462
Length = 814
Score = 121 bits (304), Expect(2) = 2e-48
Identities = 60/105 (57%), Positives = 79/105 (75%), Gaps = 1/105 (0%)
Frame = +1
Query: 733 LKSKEKVDELSLEWDYGATL-DTQIERLVLEQLQPPSSLKKLTIKSYGGTSFPNWFGDSS 791
+K KE++DEL+LEW+ +T ++Q + +VLE L+P ++LK LTIK YGG SF NW GDS
Sbjct: 1 VKMKEQLDELALEWNCCSTSSNSQSQSVVLEHLRPSTNLKNLTIKGYGGISFSNWLGDSL 180
Query: 792 FAHMVYLCISDCDHCWSLPPLGQLLGLRELYISGMKSVKIVGAEF 836
F +MVYL IS CDHC LPPLGQL L++L I GM+SV+ +G EF
Sbjct: 181 FRNMVYLRISSCDHCLWLPPLGQLGNLKKLIIEGMQSVETIGVEF 315
Score = 90.5 bits (223), Expect(2) = 2e-48
Identities = 57/126 (45%), Positives = 69/126 (54%), Gaps = 2/126 (1%)
Frame = +2
Query: 844 SLFQPFPSLQVLRFRDMPEWEDWNLIGDTTTDFPNLLHLSLKDCPKLK-GTLPINQISST 902
S FQPFPSL+ L F DM EWE+WNLI TTT+FP+L LSL CPKL+ G + S T
Sbjct: 332 SSFQPFPSLETLHFEDMQEWEEWNLIEGTTTEFPSLKTLSLSKCPKLRVGNIADKFPSLT 511
Query: 903 -FELSGCPLLFPNSMLYFTENIPTNFHSSLVLNCTNLILDLTLSRIPSSASFPRDGLPTT 961
EL CPLL + + L LNC + LT+ P FP DGLP T
Sbjct: 512 ELELRECPLLVQS----VRSSGRVLRQLMLPLNC---LQQLTIDGFPFPVCFPTDGLPKT 670
Query: 962 LRIVVI 967
L+ + I
Sbjct: 671 LKFLKI 688
>TC83046 weakly similar to GP|10280097|emb|CAC09980. nucleotide sequence of
the I-2 resistance gene {Fusarium oxysporum}, partial
(8%)
Length = 884
Score = 189 bits (481), Expect = 4e-48
Identities = 96/188 (51%), Positives = 130/188 (69%), Gaps = 3/188 (1%)
Frame = +1
Query: 360 NLEFIGKEIVKKCDGLPIAAVALGGLLRSELSENRWNKVLKSNIWDLPNVKVLPALLLSY 419
NLE IG++I KKC GLPIAA LGG+LRS++ W+ +L S+IW+LPN +LPAL LSY
Sbjct: 13 NLEEIGRKIAKKCGGLPIAAKTLGGILRSKVDAKEWSTILNSDIWNLPNDNILPALRLSY 192
Query: 420 HHLPSPLKQCFTYCSIFPKNFILEKQMVVQLWIAEGFVHQSKSGKTMEEVADEYFDELVS 479
+LPS LK+CF YCSIFPK+F L+K+ ++ LW+AEGF+ S+ KT EEV +YF EL+S
Sbjct: 193 QYLPSHLKRCFAYCSIFPKDFSLDKKELILLWMAEGFLEHSQCNKTAEEVGHDYFIELLS 372
Query: 480 RSLIHRWSVNDCVHYKMHDLINDLATMVSSSYCIRY---GDRKLQESVERVRHLS*NKGK 536
RSLI + + + + MHDL+NDLA +VS + C R G+ + VRH S N+G
Sbjct: 373 RSLIQQSNDDGKEKFVMHDLVNDLALVVSGTSCFRLECGGNMS-----KNVRHFSYNQGV 537
Query: 537 YNSFNKFD 544
Y+ KF+
Sbjct: 538 YDFLKKFE 561
>BG587633 similar to GP|20385442|gb| resistance gene analog {Vitis vinifera},
partial (22%)
Length = 677
Score = 167 bits (422), Expect(2) = 4e-46
Identities = 83/156 (53%), Positives = 111/156 (70%), Gaps = 2/156 (1%)
Frame = +3
Query: 359 SNLEFIGKEIVKKCDGLPIAAVALGGLLRSELSENRWNKVLKSNIWDLPNVKVLPALLLS 418
+ LE IG++ +KC GLPIAA LGGLLRS++ W +L S+IW+L N +LPAL LS
Sbjct: 21 TTLEAIGRKTARKCGGLPIAAKTLGGLLRSKVEITEWTSILNSDIWNLSNDNILPALHLS 200
Query: 419 YHHLPSPLKQCFTYCSIFPKNFILEKQMVVQLWIAEGFVHQSKSGKTMEEVADEYFDELV 478
Y +LP LK+CF YCSIFPK++ L+++ +V LW+AEGF+ S GK MEE+ D+ F EL+
Sbjct: 201 YQYLPCHLKRCFAYCSIFPKDYPLDRKQLVLLWMAEGFLDCSHGGKAMEELGDDCFAELL 380
Query: 479 SRSLIHRWSVNDC--VHYKMHDLINDLATMVSSSYC 512
SRSLI + S ND + MHDL+NDLAT++S C
Sbjct: 381 SRSLIQQLS-NDARGEKFVMHDLVNDLATVISGQSC 485
Score = 37.7 bits (86), Expect(2) = 4e-46
Identities = 25/55 (45%), Positives = 32/55 (57%)
Frame = +2
Query: 525 ERVRHLS*NKGKYNSFNKFDSLYESKRLRTFISLPVRLEWLPDQHYAKYFLSNKV 579
ERVRH+S N+ Y+ F KF L+ K LR+F+S+ P Y KY LS KV
Sbjct: 512 ERVRHVSYNQELYDIFMKFAKLFNFKVLRSFLSI------YPTTSYDKY-LSLKV 655
>TC93402 weakly similar to GP|16588610|gb|AAL26857.1 NBS-type putative
resistance protein {Glycine max}, partial (61%)
Length = 676
Score = 183 bits (464), Expect = 4e-46
Identities = 93/227 (40%), Positives = 146/227 (63%), Gaps = 2/227 (0%)
Frame = +2
Query: 198 KIGVISIVGMGGLGKTTLAKLLFNDHEVEDNFDLKAWAYISKDFDVCRVTKVILESITFK 257
++ ISIVG+GG+GKTTLA+L++ND +++ F+LKAW Y+S+ FDV +T IL
Sbjct: 2 QVSTISIVGLGGMGKTTLAQLVYNDRRIQETFELKAWVYVSEYFDVIGLTNAILRKFGTA 181
Query: 258 PVDTNNLNILQVELQQSLRNRRFLLVLDDIWDGSYVDWNNLMDIFSAGEKGSRIIVTTRD 317
++ +L++LQ +LQ+ L + +LLV+DD+W + W L+ F+ G GS+II+TTRD
Sbjct: 182 E-NSEDLDLLQWQLQKKLTGKNYLLVMDDVWKLNEESWEKLLLPFNYGSSGSKIILTTRD 358
Query: 318 ESVARSMQTSFPIYHLLPLASEDCWSLLAKHAFGPYNCRNRSNLEFIGKEIVKKCDGLPI 377
+ VA ++++ + L L ++DCWSL + AF N LE IGK IV KC GLP+
Sbjct: 359 KKVALIVKST-ELVDLEQLKNKDCWSLFKRLAFHGSNVSEYPKLESIGKNIVDKCGGLPL 535
Query: 378 AAVALGGLLRSELSENRWNKVLKSNIWDL--PNVKVLPALLLSYHHL 422
+G LLR + +++ W K+L++++W L + + AL LSYH+L
Sbjct: 536 TGKTMGNLLRKKFTQSEWEKILEADMWRLTDDDSNINSALRLSYHNL 676
>BG649127 weakly similar to GP|11994217|db disease resistance comples protein
{Arabidopsis thaliana}, partial (6%)
Length = 688
Score = 166 bits (419), Expect = 6e-41
Identities = 105/236 (44%), Positives = 146/236 (61%)
Frame = +2
Query: 434 SIFPKNFILEKQMVVQLWIAEGFVHQSKSGKTMEEVADEYFDELVSRSLIHRWSVNDCVH 493
SIFPK+ +K ++ LW+AEG + Q ++ K ME+V +E F+ L+SRS ++ S H
Sbjct: 8 SIFPKD*N-KKWNLIYLWMAEGILPQQRTDKRMEDVREECFEVLLSRSFFYQ-STYHASH 181
Query: 494 YKMHDLINDLATMVSSSYCIRYGDRKLQESVERVRHLS*NKGKYNSFNKFDSLYESKRLR 553
Y MHDLI+D+A V+ +C D ++ VRHLS +G Y+ KF+ E K+LR
Sbjct: 182 YMMHDLIHDVAQFVAGEFCYNLDDNNPRKITTIVRHLSYLQGIYDDPEKFEIFSEFKQLR 361
Query: 554 TFISLPVRLEWLPDQHYAKYFLSNKVLHDLLSEIRPLRVLSLSYYLNITDLPQYLGNLIH 613
TFI P + + Y+ S ++ LL +++ LRVLSLS+Y IT+L +G L+H
Sbjct: 362 TFI--PFKFSYFV---YSSSITS--MVSILLPKLKRLRVLSLSHY-PITNLSDSIGVLMH 517
Query: 614 LRYLDLSNTKIQRLPYETCKLYNLQTLLLSRCWLLIELPEDMGNLINLRHLDICGT 669
+RYLDLS T I+ LP LYNL+TLLLS C L LPE+M NLINLR LDI G+
Sbjct: 518 MRYLDLSYTGIECLPDSVSTLYNLETLLLSGCRCLTILPENMSNLINLRQLDISGS 685
>AW686540 weakly similar to GP|15289776|db Lycopersicon esculentum resistance
complex protein I2C-2 like protein, partial (2%)
Length = 566
Score = 165 bits (418), Expect = 8e-41
Identities = 87/176 (49%), Positives = 117/176 (66%), Gaps = 3/176 (1%)
Frame = +1
Query: 720 QNVTDPFEAFRANLKSKEKVDELSLEWDYGATLDTQIE-RLVLEQLQPPSSLKKLTIKSY 778
QNV D EA+ A+LKSKE ++EL+L+W G D ++ + VL+ L+PP +L +L I Y
Sbjct: 1 QNVIDVAEAYDADLKSKEHIEELTLQW--GVETDDPLKGKDVLDMLKPPVNLNRLNIDLY 174
Query: 779 GGTSFPNWFGDSSFAHMVYLCISDCDHCWSLPPLGQLLGLRELYISGMKSVKIVGAEFYG 838
GGTSFP+W GDSSF++MV L I C +C +LPPLGQL L++L I GM ++ +G EFYG
Sbjct: 175 GGTSFPSWLGDSSFSNMVSLSIQHCGYCVTLPPLGQLSSLKDLSIRGMYILETIGPEFYG 354
Query: 839 --SSSSSSLFQPFPSLQVLRFRDMPEWEDWNLIGDTTTDFPNLLHLSLKDCPKLKG 892
S+S FQPFPSL+ L+F MP W+ W D FP L L L +CP+L+G
Sbjct: 355 IVGGGSNSSFQPFPSLEKLQFVKMPNWKKWLPFQDGIFPFPCLKSLILYNCPELRG 522
>AW694719 similar to GP|11994217|db disease resistance comples protein
{Arabidopsis thaliana}, partial (6%)
Length = 511
Score = 164 bits (415), Expect = 2e-40
Identities = 86/158 (54%), Positives = 112/158 (70%), Gaps = 4/158 (2%)
Frame = +1
Query: 361 LEFIGKEIVKKCDGLPIAAVALGGLLRSELSENRWNKV---LKSNIWDLPNVKVLPALLL 417
LE IG++I ++C GLPIAA LGGLLRS++ W + L S+IW+L N +LPAL L
Sbjct: 13 LEEIGRKIARRCGGLPIAAKTLGGLLRSKVDITEWTSIFSILNSSIWNLRNDNILPALHL 192
Query: 418 SYHHLPSPLKQCFTYCSIFPKNFILEKQMVVQLWIAEGFVHQSKSGKTMEEVADEYFDEL 477
SY +LPS LK+CF YCSIFPK+ L+K+ +V LW+AEGF+ S+ GK +EE+ D+ F EL
Sbjct: 193 SYQYLPSHLKRCFAYCSIFPKDCPLDKKQLVLLWMAEGFLDCSQGGKKLEELGDDCFAEL 372
Query: 478 VSRSLIHRWSVND-CVHYKMHDLINDLATMVSSSYCIR 514
+SRSLI + S +D + MHDLINDLAT VS C R
Sbjct: 373 LSRSLIQQLSDDDRGEKFVMHDLINDLATFVSGKSCCR 486
>CA920463 weakly similar to GP|2852684|gb|A resistance protein candidate
{Lactuca sativa}, partial (6%)
Length = 810
Score = 161 bits (407), Expect = 2e-39
Identities = 95/235 (40%), Positives = 136/235 (57%), Gaps = 4/235 (1%)
Frame = -1
Query: 278 RRFLLVLDDIWDGSYVDWNNLMDIFSAGEKGSRIIVTTRDESVARSMQTSFPIYHLLPLA 337
++FLL LDD+W V W + ++ G++GS+++VTTR S+A+ M T+ Y L L+
Sbjct: 756 KKFLLRLDDVWSEDRVKWIEVKNLLQVGDEGSKVLVTTRSHSIAKMMCTNTS-YTLQGLS 580
Query: 338 SEDCWSLLAKHAFGPYNCRNRSNLEFIGKEIVKKCDGLPIAAVALGGLLRSELSENRWNK 397
ED S+ K AF + L IGKEIV+KC GLP+A LG L + W
Sbjct: 579 REDSLSVFVKWAFKEGEEKKYPKLIEIGKEIVQKCGGLPLALRTLGSSLFLKDDIEEWKF 400
Query: 398 VLKSNIWDLPNVK--VLPALLLSYHHLPSPLKQCFTYCSIFPKNFILEKQMVVQLWIAEG 455
V + IW+LP + +LPAL LS+ LPS LK+CF S+F K+F V LW A
Sbjct: 399 VRDNEIWNLPQKEDDILPALKLSFDQLPSYLKRCFACFSLFVKDFHFSNYSVTVLWEALD 220
Query: 456 FVHQSKSGKTMEEVADEYFDELVSRSLIHRWSV--NDCVHYKMHDLINDLATMVS 508
F+ GKT+E+V +++ EL SRS + + V N CV +K+HDL++DLA V+
Sbjct: 219 FLPSPNKGKTLEDVGNQFLHELQSRSFLQDFYVSGNVCV-FKLHDLVHDLALYVA 58
>BG451218 similar to GP|11994217|db disease resistance comples protein
{Arabidopsis thaliana}, partial (5%)
Length = 666
Score = 155 bits (392), Expect = 8e-38
Identities = 85/183 (46%), Positives = 116/183 (62%), Gaps = 1/183 (0%)
Frame = +1
Query: 379 AVALGGLLRSELSENRWNKVLKSNIWDLPNVKVLPALLLSYHHLPSPLKQCFTYCSIFPK 438
A +GGLL S++ W +L SN+W+LPN K+LP L LSY LPS LK CF YCSIFPK
Sbjct: 85 AKTIGGLLGSKVDIIEWTTILNSNVWNLPNDKILPTLHLSYQCLPSHLKICFAYCSIFPK 264
Query: 439 NFILEKQMVVQLWIAEGFVHQSKSGKTMEEVADEYFDELVSRSLIHRWSVND-CVHYKMH 497
+++ +V LW+AEGF+ S KTMEE+ D+ F EL+SRSLI + + N + MH
Sbjct: 265 GHTHDRKKLVLLWMAEGFLDYSHGEKTMEELGDDCFAELLSRSLIQQSNDNGRGEKFFMH 444
Query: 498 DLINDLATMVSSSYCIRYGDRKLQESVERVRHLS*NKGKYNSFNKFDSLYESKRLRTFIS 557
DL+NDLAT+VS C R+ + E+ VRH+S + +Y+ KF + K LRTF+
Sbjct: 445 DLVNDLATVVSGKSCCRFECGNISEN---VRHVSYIQEEYDIVTKFKPFHNLKCLRTFLP 615
Query: 558 LPV 560
+ V
Sbjct: 616 IHV 624
Score = 30.0 bits (66), Expect = 5.3
Identities = 17/40 (42%), Positives = 23/40 (57%)
Frame = +3
Query: 359 SNLEFIGKEIVKKCDGLPIAAVALGGLLRSELSENRWNKV 398
S E IG++I +KC GLPIA R+ E R+N+V
Sbjct: 24 STFEEIGRKIARKCGGLPIAC---*NNRRTSRFEGRYNRV 134
>BF634887 weakly similar to GP|10280097|em nucleotide sequence of the I-2
resistance gene {Fusarium oxysporum}, partial (3%)
Length = 694
Score = 152 bits (384), Expect = 7e-37
Identities = 94/238 (39%), Positives = 135/238 (56%), Gaps = 1/238 (0%)
Frame = +1
Query: 672 KYMPSQIAKLQNLQTLSAFIVSKSQDGLKVGELKNFTNLQGKLSISKLQNVTDPFEAFRA 731
K MP QI L +LQTLS F+V + ++G + +L N LQGKL IS L++V +P +A A
Sbjct: 1 KKMPKQIGSLNHLQTLSHFVVGE-ENGSNIQQLGNLNCLQGKLCISGLEHVINPKDAAGA 177
Query: 732 NLKSKEKVDELSLEWDYGATLDTQ-IERLVLEQLQPPSSLKKLTIKSYGGTSFPNWFGDS 790
NLK K+ V+EL++E+ + E V E L P S+LK+L I Y G +FPNW
Sbjct: 178 NLKDKKHVEELNMEYSDNFKFNNNGRESNVFEALHPNSNLKRLYIGRYQGNNFPNWITGC 357
Query: 791 SFAHMVYLCISDCDHCWSLPPLGQLLGLRELYISGMKSVKIVGAEFYGSSSSSSLFQPFP 850
++V L + C C +PP+ QL L+E IS +KI+ EFYG+SS++ F+
Sbjct: 358 HLPNLVSLQLKGCG-CSHMPPIEQLPSLKEFSISNCNGIKIITEEFYGNSSTNVSFR--- 525
Query: 851 SLQVLRFRDMPEWEDWNLIGDTTTDFPNLLHLSLKDCPKLKGTLPINQISSTFELSGC 908
SL+VL+F +M WE+W FP L + + +CPKLK L + S +L C
Sbjct: 526 SLEVLKFEEMNNWEEW----FCPEGFPLLKEIYIWNCPKLKXALLPQHLPSIQKLKIC 687
>BG586564 weakly similar to GP|14348622|gb NBS-LRR resistance-like protein
J78 {Phaseolus vulgaris}, partial (6%)
Length = 733
Score = 151 bits (381), Expect = 2e-36
Identities = 82/177 (46%), Positives = 108/177 (60%), Gaps = 5/177 (2%)
Frame = +2
Query: 760 VLEQLQPPSSLKKLTIKSYGGTSFPNWFGDSSFAHMVYLCISDCDHCWSLPPLGQLLGLR 819
VL+ L PP +L +L I YGGTSFP+W GDSSF++MV LCI +C +C +LPPLGQL L+
Sbjct: 35 VLDMLIPPVNLNRLNIYFYGGTSFPSWLGDSSFSNMVSLCIENCRYCVTLPPLGQLSSLK 214
Query: 820 ELYISGMKSVKIVGAEFYG--SSSSSSLFQPFPSLQVLRFRDMPEWEDWNLIGDTTTDFP 877
+L I GM ++ +G EFYG S+S FQPF SL+ L F +MP W+ W L D FP
Sbjct: 215 DLTIRGMSILETIGPEFYGIVGGGSNSSFQPFSSLEKLEFTNMPNWKKWLLFQDGILPFP 394
Query: 878 NLLHLSLKDCPKLKGTLPINQIS-STFELSGCPLLF--PNSMLYFTENIPTNFHSSL 931
L L L DC +L+G LP + S F GCP L P ++ + + +F SL
Sbjct: 395 CLKSLKLYDCTELRGNLPSHLSSIEEFVNKGCPHLLESPPTLEWLSSIKEIDFSGSL 565
>BF643456 weakly similar to GP|14348622|gb NBS-LRR resistance-like protein
J78 {Phaseolus vulgaris}, partial (6%)
Length = 521
Score = 148 bits (373), Expect = 1e-35
Identities = 75/170 (44%), Positives = 113/170 (66%), Gaps = 1/170 (0%)
Frame = +2
Query: 221 NDHEVEDNFDLKAWAYISKDFDVCRVTKVILESITFKPVDTNNLNILQVELQQSLRNRRF 280
ND+ ++ NFD++AWA +S FD +VTK I+E++T + NN+ +L ++L++ L ++F
Sbjct: 2 NDN-IKQNFDVQAWACVSDHFDEFKVTKAIMEAVTRSACNINNIELLHLDLKEKLAGKKF 178
Query: 281 LLVLDDIWDGSYVDWNNLMDIFSAGEKGSRIIVTTRDESVARSMQTSFPIYHLLPLASED 340
L+VLDD+W Y W +L+ +KGS+I+VTTR + VA +QT F Y L L+ ED
Sbjct: 179 LIVLDDVWTEDYDAWKSLLRPLQYSDKGSKILVTTRIKKVASMLQT-FQGYSLEQLSDED 355
Query: 341 CWSLLAKHA-FGPYNCRNRSNLEFIGKEIVKKCDGLPIAAVALGGLLRSE 389
CWS+ HA + + L+ IGKEI++KC GLP+AA +LGGLLRS+
Sbjct: 356 CWSVFENHACLSLKHSIEKMELQKIGKEIIRKCQGLPLAAQSLGGLLRSK 505
>BI265031 weakly similar to GP|13872974|db putative NBS-LRR type resistance
protein {Oryza sativa (japonica cultivar-group)},
partial (3%)
Length = 662
Score = 127 bits (320), Expect(2) = 1e-32
Identities = 70/170 (41%), Positives = 105/170 (61%)
Frame = +1
Query: 181 KKLKEFLLLEDGSVSGSKIGVISIVGMGGLGKTTLAKLLFNDHEVEDNFDLKAWAYISKD 240
+K EFLL + S S + + +ISIVG+ G+GKTT+A+L++NDH++ + F+LKAW Y+S+
Sbjct: 79 RK*PEFLLSD--SYSETFVPIISIVGVIGMGKTTIARLVYNDHKIHEQFELKAWVYVSES 252
Query: 241 FDVCRVTKVILESITFKPVDTNNLNILQVELQQSLRNRRFLLVLDDIWDGSYVDWNNLMD 300
FD+ +T+ IL + ++ ILQ +LQQ L +++LLVLD+IW+ + L+
Sbjct: 253 FDLVHLTQAILREFHSSETYSEDMEILQRQLQQRLAGKKYLLVLDNIWNENVECRKKLLL 432
Query: 301 IFSAGEKGSRIIVTTRDESVARSMQTSFPIYHLLPLASEDCWSLLAKHAF 350
FS G GS++IV T VA S+ S + L L D WSL HAF
Sbjct: 433 PFSNGSSGSKLIVRTPHNEVA-SIMASTRLLRLNQLNESDSWSLFVHHAF 579
Score = 32.0 bits (71), Expect(2) = 1e-32
Identities = 16/34 (47%), Positives = 19/34 (55%)
Frame = +3
Query: 344 LLAKHAFGPYNCRNRSNLEFIGKEIVKKCDGLPI 377
L+ F N NLE IGK+IV KC GLP+
Sbjct: 558 LICASCFFGQNIFEYPNLESIGKKIVXKCGGLPL 659
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 40,510,163
Number of Sequences: 36976
Number of extensions: 653419
Number of successful extensions: 4350
Number of sequences better than 10.0: 336
Number of HSP's better than 10.0 without gapping: 3869
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4131
length of query: 1147
length of database: 9,014,727
effective HSP length: 107
effective length of query: 1040
effective length of database: 5,058,295
effective search space: 5260626800
effective search space used: 5260626800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 64 (29.3 bits)
Medicago: description of AC141414.10


