

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
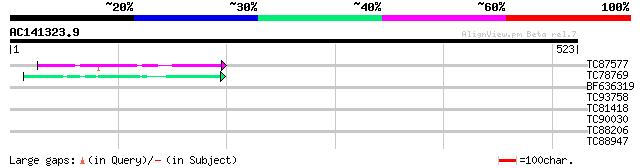
Query= AC141323.9 - phase: 0
(523 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC87577 weakly similar to PIR|H96604|H96604 probable 3'-5' exonu... 85 6e-17
TC78769 similar to PIR|C96586|C96586 hypothetical protein F20D21... 47 2e-05
BF636319 weakly similar to PIR|G71155|G711 hypothetical protein ... 30 2.3
TC93758 weakly similar to GP|22002164|gb|AAM88648.1 putative rec... 29 3.8
TC81418 similar to GP|3047109|gb|AAC13620.1| F6N23.17 gene produ... 29 5.0
TC90030 similar to PIR|C96767|C96767 unknown protein F2P9.17 [im... 29 5.0
TC88206 similar to PIR|A86289|A86289 probable ABC transporter [i... 28 6.6
TC88947 similar to GP|4586586|dbj|BAA76425.1 bZIP DNA binding pr... 28 6.6
>TC87577 weakly similar to PIR|H96604|H96604 probable 3'-5' exonuclease
[imported] - Arabidopsis thaliana, partial (54%)
Length = 1980
Score = 85.1 bits (209), Expect = 6e-17
Identities = 60/178 (33%), Positives = 90/178 (49%), Gaps = 3/178 (1%)
Frame = +1
Query: 26 TRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFLLDLISLPLS---SI 82
T + ++GLD EWKP + VS++QIA ++ F+LDLI L +
Sbjct: 1198 TCQIEDAKVIGLDCEWKPNYVKGSKPNKVSIMQIA----SEKKAFILDLIKLHREVPERL 1365
Query: 83 WEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVEPYLDITSVYNHLQFKK 142
L +L+S ILKLG+ F+ D+ L+ ++ E C F K + LDI V+
Sbjct: 1366 DSCLTRILLSPGILKLGYNFQCDIKQLAHSYEELEC---FKKYKRLLDIQKVFKD----- 1521
Query: 143 NGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRPLTEEQMTYAAMDAHCLLGIFK 200
L+ + ++LG L+K + SDW QRPLT Q+ YAA+DA L+ IF+
Sbjct: 1522 -------PRGGLAKLAEKILGAGLNKTRRNSDWEQRPLTPNQLEYAALDAVVLVHIFR 1674
>TC78769 similar to PIR|C96586|C96586 hypothetical protein F20D21.26
[imported] - Arabidopsis thaliana, partial (46%)
Length = 2124
Score = 46.6 bits (109), Expect = 2e-05
Identities = 45/187 (24%), Positives = 74/187 (39%)
Frame = +2
Query: 13 FVTTTDSPEFTHLTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFLL 72
F + + + L + +D E R+ Q L+QI+ + D F++
Sbjct: 2 FKLVEEVKDLKEMAAKLRSVNEFAVDLEHNQYRSFQG---LTCLMQISTRTED----FVV 160
Query: 73 DLISLPLSSIWEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVEPYLDIT 132
D + L + I LRE+ K+ +D+V+L F CN FD +
Sbjct: 161 DTLKLRIH-IGPHLREVFKDPSKRKVMHGADKDIVWLQRDFGIYVCNM-FDTGQA----- 319
Query: 133 SVYNHLQFKKNGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRPLTEEQMTYAAMDA 192
++ + SL + IT +KE Q +DW RP+ +E + YA D
Sbjct: 320 -----------SKVLKLERNSLEHLLNHFCEITANKEYQNADWRLRPIPDEMLRYAREDT 466
Query: 193 HCLLGIF 199
H LL I+
Sbjct: 467 HYLLYIY 487
>BF636319 weakly similar to PIR|G71155|G711 hypothetical protein PH0446 -
Pyrococcus horikoshii, partial (5%)
Length = 595
Score = 30.0 bits (66), Expect = 2.3
Identities = 18/38 (47%), Positives = 19/38 (49%)
Frame = +3
Query: 473 CLFNKNLEFWQCMDCHQLYWEVLFTCSVCCELLFTIAL 510
C F KN F Q C +L W LF CS C LL I L
Sbjct: 246 CYF*KNGNF-QRDSC*ELIWVFLFCCSHLCTLLSPIKL 356
>TC93758 weakly similar to GP|22002164|gb|AAM88648.1 putative
receptor-like protein kinase {Oryza sativa (japonica
cultivar-group)}, partial (3%)
Length = 814
Score = 29.3 bits (64), Expect = 3.8
Identities = 14/24 (58%), Positives = 18/24 (74%), Gaps = 1/24 (4%)
Frame = -3
Query: 72 LDLISLPLSSIWEP-LREMLVSAD 94
LD +S+PLS++W P LR LVS D
Sbjct: 83 LDNVSVPLSTVWSPTLRRSLVSND 12
>TC81418 similar to GP|3047109|gb|AAC13620.1| F6N23.17 gene product
{Arabidopsis thaliana}, partial (61%)
Length = 1233
Score = 28.9 bits (63), Expect = 5.0
Identities = 11/32 (34%), Positives = 15/32 (46%), Gaps = 1/32 (3%)
Frame = +3
Query: 474 LFNKNLEFWQCMDCH-QLYWEVLFTCSVCCEL 504
+F E W + CH Q +W C +CC L
Sbjct: 198 IFPSRFEIWHWI*CHFQCFWPA*CGCQLCCHL 293
>TC90030 similar to PIR|C96767|C96767 unknown protein F2P9.17 [imported] -
Arabidopsis thaliana, partial (2%)
Length = 768
Score = 28.9 bits (63), Expect = 5.0
Identities = 27/107 (25%), Positives = 50/107 (46%)
Frame = +1
Query: 389 KAQKEKRVLLTRDAKLLRHDYQIYKVKSLLKNEQLLEIIETFQLNINEDQLMSRCTKCNG 448
+ +KEK+ AKLL ++ + + K E+ E+ + QL E + ++ G
Sbjct: 223 RLKKEKKRKEKELAKLLSNEAKRSSIDLSCKKEEP-EVNDAKQLKSVEPSCYNSVSEI-G 396
Query: 449 RFIQKPLSTEEAIEAAKGFQKIPNCLFNKNLEFWQCMDCHQLYWEVL 495
R KP+ E A K KI N + +K+ ++ D +++W V+
Sbjct: 397 RVDPKPVPPEGTSGAPKIRIKIKNRMLSKS*RIYEGNDALRIFWVVV 537
>TC88206 similar to PIR|A86289|A86289 probable ABC transporter [imported] -
Arabidopsis thaliana, partial (21%)
Length = 1483
Score = 28.5 bits (62), Expect = 6.6
Identities = 17/49 (34%), Positives = 24/49 (48%), Gaps = 1/49 (2%)
Frame = -1
Query: 214 NKTNILSIRSANLGLKELFR-KHDTSDKVHSTQFCEALAIVQATSCSDV 261
N TN++SI N+GLK + D ++ Q C L V SC+ V
Sbjct: 457 NMTNVVSISKPNIGLKMIVSIGQRNKDHLNQVQVCLILL*VACFSCTIV 311
>TC88947 similar to GP|4586586|dbj|BAA76425.1 bZIP DNA binding protein
{Cicer arietinum}, partial (15%)
Length = 700
Score = 28.5 bits (62), Expect = 6.6
Identities = 17/60 (28%), Positives = 25/60 (41%)
Frame = +1
Query: 149 KQNKSLSTICGELLGITLSKELQCSDWSQRPLTEEQMTYAAMDAHCLLGIFKVFQATVAK 208
K+ SLS G + LS+ + CS S PL + D + IF Q ++K
Sbjct: 10 KRYSSLSFSSGHCSSLLLSQSMLCSSSSSLPLAPATLRKPPQDCELRMNIFISHQIQISK 189
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.136 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,787,907
Number of Sequences: 36976
Number of extensions: 272263
Number of successful extensions: 1459
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1452
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1457
length of query: 523
length of database: 9,014,727
effective HSP length: 101
effective length of query: 422
effective length of database: 5,280,151
effective search space: 2228223722
effective search space used: 2228223722
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC141323.9


