

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
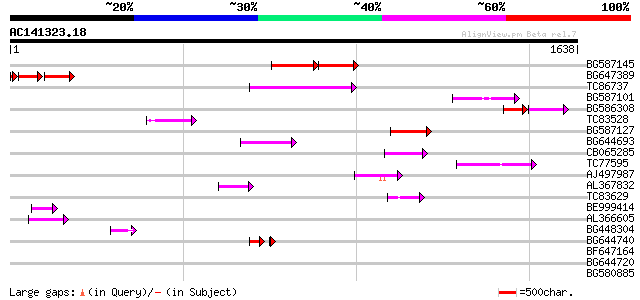
Query= AC141323.18 - phase: 0
(1638 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 145 6e-63
BG647389 116 3e-46
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 138 2e-32
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 117 3e-26
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 74 5e-25
TC83528 102 1e-21
BG587127 weakly similar to PIR|H84506|H84 probable retroelement ... 101 3e-21
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 82 2e-15
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 81 3e-15
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 75 2e-13
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 73 1e-12
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 65 2e-10
TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNa... 56 1e-07
BE999414 55 2e-07
AL366605 52 1e-06
BG448304 50 7e-06
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 45 3e-05
BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis t... 41 0.004
BG644720 40 0.006
BG580885 39 0.017
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 145 bits (367), Expect(2) = 6e-63
Identities = 67/136 (49%), Positives = 103/136 (75%)
Frame = +2
Query: 756 MDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGATFQRSMDTIFSNQIGRNLEVYIDDLVVK 815
M P D KTAF+T++ Y Y+VM FGL+NAG+T+QR ++ +F++++G +EVYIDD++VK
Sbjct: 2 MHPDDLEKTAFITDRGTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVK 181
Query: 816 TSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFGVQAGKFLGFLLTKKGIEANPDKCQAII 875
+ H LKE F+ + ++ M+LNP KCTFGV +G+FLG+++T++GIE NP + AI+
Sbjct: 182 SLRATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQITAIL 361
Query: 876 NMRNPCNIREVQQLTG 891
++ +P N REVQ+LTG
Sbjct: 362 DLPSPKNSREVQRLTG 409
Score = 115 bits (288), Expect(2) = 6e-63
Identities = 53/118 (44%), Positives = 82/118 (68%)
Frame = +3
Query: 891 GRLAALSRFLSCAGDKAFAFFATIKKKEEFEWNQECDEAFGKIKQFLTTPPILHRPTKGA 950
GR+AAL+RF+S + DK F+ + + F W+++C+EAF ++KQ+LTTPP+L +P G
Sbjct: 408 GRIAALNRFISRSTDKCLPFYKLLCGNKRFVWDEKCEEAFEQLKQYLTTPPVLSKPEAGD 587
Query: 951 GLFLYLSVSENALSSVLVEESDEREKPIYFVSRVLKGAELRYQKIEKLALAVIITARK 1008
L LY+++S A+SSVL+ E +KPI++ S+ + E RY +EK+A AVI +ARK
Sbjct: 588 TLSLYIAISSTAVSSVLIREDRGEQKPIFYTSKRMTDPETRYPTLEKMAFAVITSARK 761
>BG647389
Length = 718
Score = 116 bits (291), Expect(3) = 3e-46
Identities = 55/85 (64%), Positives = 67/85 (78%)
Frame = +1
Query: 102 LADAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQGPNESLREYLA 161
L +A +WYM+LPRFSI GYQ++T K QFS ++H KV +T+LFNVRQ PNESL EYL
Sbjct: 448 LKEATQQWYMNLPRFSITGYQNMTHKLVHQFSASKHCKVSTTNLFNVRQDPNESLPEYLV 627
Query: 162 RFNDSTIKVSNPNQEVFVGAFQNGL 186
FN++TIKV NP QE+F GAFQNGL
Sbjct: 628 *FNNATIKVVNPKQELFAGAFQNGL 702
Score = 82.4 bits (202), Expect(3) = 3e-46
Identities = 37/70 (52%), Positives = 48/70 (67%)
Frame = +2
Query: 26 QQQEKTDSHHGDLDEPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMS 85
QQ E+ + H D+DE +PQP S +IWNAPV +N K L +FD +SDP EH T FNT+M+
Sbjct: 224 QQHEQDELHREDVDELDPQPLSMDIWNAPVLENLKSLSLLSFDDKSDPVEHATTFNTQMA 403
Query: 86 VYGVADSLKC 95
V G SL+C
Sbjct: 404 VIGALKSLRC 433
Score = 27.3 bits (59), Expect(3) = 3e-46
Identities = 13/23 (56%), Positives = 18/23 (77%)
Frame = +3
Query: 1 MRSQIAQLLEQNEALLASVETIQ 23
M+ QI QLL++NEA+ AS+ IQ
Sbjct: 153 MKLQIKQLLQKNEAM*ASMTAIQ 221
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 138 bits (347), Expect = 2e-32
Identities = 92/312 (29%), Positives = 144/312 (45%), Gaps = 3/312 (0%)
Frame = +1
Query: 694 ANTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPSIDHLIDNASGYKTLSFMDAYSGYNQ 753
A + V+K G R CVDY LN KD YPLP I + +G + + +D + +++
Sbjct: 577 APVLFVRKPGGGIRFCVDYRALNAITKKDRYPLPLISETLRRVAGARWFTKLDVVAAFHK 756
Query: 754 IKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAGATFQRSMDTIFSNQIGRNLEVYIDD-L 812
+++ D KTAF T + + V FGL A ATFQR ++ + + YIDD L
Sbjct: 757 MRIKDEDQEKTAFRTRYGLFEWIVCPFGLTGAPATFQRYINKTLHEFLDDFVTAYIDDVL 936
Query: 813 VVKTSEKQSHSVDLKEIFQQIRKFSMRLNPTKCTFGVQAGKFLGFLLTK-KGIEANPDKC 871
+ T K+ H ++ + +++ + L+P KC F V K++GF+LT KG+ +P K
Sbjct: 937 IYTTGSKKDHEAQVRRVLRRLADAGLSLDPKKCEFSVTTVKYVGFILTAGKGVSCDPLKL 1116
Query: 872 QAIINMRNPCNIREVQQLTGRLAALSRFLSCAGDKAFAFFATIKKKEEFEWNQECDEAFG 931
AI + P +++ + G F+ + +K F W E + AF
Sbjct: 1117 AAIRDWLPPGSVKGARSFLGFCNYYKDFIPGYSEITEPLTRLTRKDFPFRWGAEQEAAFT 1296
Query: 932 KIKQFLTTPPILHRPTKGAGLFLYLSVSENALSSVLVEESDE-REKPIYFVSRVLKGAEL 990
K+K+ P+L A + S AL VL +E P+ F S+ L AE
Sbjct: 1297 KLKRLFAEEPVLRMFDPEAVTTVETDCSGFALGGVLTQEDGTGAAHPVAFHSQRLSPAEY 1476
Query: 991 RYQKIEKLALAV 1002
Y +K LAV
Sbjct: 1477 NYPIHDKELLAV 1512
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 117 bits (294), Expect = 3e-26
Identities = 65/197 (32%), Positives = 99/197 (49%), Gaps = 3/197 (1%)
Frame = +2
Query: 1279 LYKRGRASPLLRCLGEGETELVLLEVHEGVCGSHIGGRSLAAKLLRAGYYWPRMAHDCCE 1338
LYK+ + +RC+ E E +L H H +K+ +AG++WP M D
Sbjct: 41 LYKQCADNIYIRCVAEEEIPGILFHCHGSNYAGHFAVSKTVSKIQQAGFWWPTMFKDAHS 220
Query: 1339 FVKKCDKCQR---FSDKKNAPANELTSVFSPWPFHKWGVDIVGPFPQAPGQLKFLIVDVD 1395
F+ KCD CQR S + P N + V F WG+D +GPFP + K+++V VD
Sbjct: 221 FISKCDPCQRQGNIS*RNEMPQNFILEVEV---FDVWGIDFMGPFPSSYNN-KYILVAVD 388
Query: 1396 YFTKWVEAEAVSKITAERVVKFYWKKIICHFGLPKYIVTDNGTQFASSKVVNFCKQLGIE 1455
Y +KWVEA A A VVK + I FG+P+ +++D G+ F + K+ G+
Sbjct: 389 YVSKWVEAIASPTNDATVVVKMFKSVIFPRFGVPRVVISDGGSHFINKVFEKLLKKNGVR 568
Query: 1456 TKFVSVIHPQANGQAES 1472
K + HPQ +++S
Sbjct: 569 HKVATAYHPQKAERSKS 619
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 686
Score = 73.6 bits (179), Expect(2) = 5e-25
Identities = 34/70 (48%), Positives = 50/70 (70%)
Frame = -2
Query: 1427 GLPKYIVTDNGTQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLE 1486
GLP IVTDNG+ F S+K FC++ I S +PQ+NGQAE++NK+I++ +KK+L+
Sbjct: 682 GLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKIIIDGLKKRLD 503
Query: 1487 DAKGLWAEQL 1496
KG WA++L
Sbjct: 502 LKKGCWADEL 473
Score = 60.8 bits (146), Expect(2) = 5e-25
Identities = 38/119 (31%), Positives = 61/119 (50%), Gaps = 5/119 (4%)
Frame = -3
Query: 1499 VLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSE--ESNEVGIRCTMDMI 1556
VLWS+ T P T TPF+M + +AM P E++ + R + E N + ++ I
Sbjct: 465 VLWSHRTNPRGATKSTPFSMAHRVEAMAPAEVNVTSL*RSRMPQNIELNNDRLFNALETI 286
Query: 1557 DEVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTR---IGKLLPSWKG 1612
+E R+ A +R + + YN V + +K GD+VL +V T+ GKL +W+G
Sbjct: 285 EERRDQALLRIQNYQHQIESYYNKTVKSQPLKLGDIVLCKVFENTKELNAGKLGTNWEG 109
>TC83528
Length = 555
Score = 102 bits (254), Expect = 1e-21
Identities = 60/148 (40%), Positives = 84/148 (56%), Gaps = 4/148 (2%)
Frame = +3
Query: 395 PKKYPHVTFSSEDFGRVLPHDDDPLVISVQLFN-WEIKRVLIDTGSSADVLYFDAFSKMG 453
PKKY TF+ D P+VI +Q+ N + + RV ++ S D+LY+ AF KM
Sbjct: 141 PKKY---TFTGPDVAL-------PMVIKLQINNNFSVLRVFVNPMSKVDILYWSAFLKMK 290
Query: 454 LSEEQLQPFNGTLSGFTGEKVHVRGYVTLKTIFGTGDQQTSIKVRYLVINSPSS---YNV 510
L E L+P G L G G+ + V+GY+ L T FG G+ +IKVRY V+ SP S YNV
Sbjct: 291 LQESMLKPCQGFLKGTFGKGLPVKGYIDLDTTFGKGENTKTIKVRYFVVESPPSVSIYNV 470
Query: 511 IIGRPSINLLDAFVSTKHLLMKYPLDSG 538
++G PS+ L A +S +KYP+ G
Sbjct: 471 VLGWPSLKDLKAVLSVAEFTIKYPVGDG 554
>BG587127 weakly similar to PIR|H84506|H84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(13%)
Length = 415
Score = 101 bits (251), Expect = 3e-21
Identities = 51/119 (42%), Positives = 74/119 (61%)
Frame = +3
Query: 1099 GIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQEMGAKNLRARSDSQLMTNQI 1158
GI L P + +++QS + +F ASNN+ YEALIAG+ LA + +N+ A DSQL+ +Q
Sbjct: 3 GIRLTSPTNEILKQSFRLEFHASNNETNYEALIAGVRLAHGLKIRNIHAYCDSQLVASQF 182
Query: 1159 SGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSRADLLAKLASTKKPGNNRTV 1217
SGEY+ +D+ + YL V+KLA +F +PR N +AD LA LAS+ P R +
Sbjct: 183 SGEYEARDELMDTYLKLVQKLAQKLDYFALTRIPRSENVQADALAALASSSDPELKRVI 359
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 82.0 bits (201), Expect = 2e-15
Identities = 53/161 (32%), Positives = 78/161 (47%)
Frame = +2
Query: 667 KRKAVDEEVKKLQEAHFICEIKYPTWLANTVLVKKASGKWRMCVDYTDLNMACPKDPYPL 726
K K + ++K L E FI YP + + +KK G RM +DY LN K YPL
Sbjct: 107 KLKVLKLQLKDLLEKGFIQPSIYP*GVV-VLFLKKKDGFLRMSIDYPQLNNVNIKIKYPL 283
Query: 727 PSIDHLIDNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTNQKNYHYRVMSFGLRNAG 786
P ID L DN G K +D G +Q ++ D PKTAF +Y VMSFG N
Sbjct: 284 PLIDELFDNLQGSKWFFKIDLRLG*HQHRVIGEDVPKTAFRIRYGHYEILVMSFG*TNPP 463
Query: 787 ATFQRSMDTIFSNQIGRNLEVYIDDLVVKTSEKQSHSVDLK 827
F M+ +F + + + V+ +D+++ + + H L+
Sbjct: 464 MAFMELMNRVFQDYLDSLVIVFSNDILIYSKNENEHENHLR 586
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 81.3 bits (199), Expect = 3e-15
Identities = 48/125 (38%), Positives = 66/125 (52%)
Frame = -2
Query: 1083 WLLSVDGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQEMGA 1142
W L DG+ N G G G ++ P I + + F+ +NN AEYEA I G+ A +M
Sbjct: 534 WGLVFDGAVNAYGKGIGAVIVSPQGHYIPFTARILFECTNNMAEYEACIFGIEEAIDMRI 355
Query: 1143 KNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSRADLL 1202
K+L DS L+ NQI GE++T L Y R+L F E ++PR+ N AD L
Sbjct: 354 KHLDIYGDSALVINQIKGEWETHHANLIPYRDYARRLLTYFTKVELHHIPRDENQMADAL 175
Query: 1203 AKLAS 1207
A L+S
Sbjct: 174 ATLSS 160
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial (14%)
Length = 1708
Score = 75.5 bits (184), Expect = 2e-13
Identities = 60/235 (25%), Positives = 105/235 (44%), Gaps = 5/235 (2%)
Frame = +2
Query: 1292 LGEGETELVLLEVHEGVCGSHIGGRSLAAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSD 1351
L E T+LV E H+ H GR+ +++ ++WP + FV+ CD C
Sbjct: 158 LNELRTKLVQ-ESHDSTAAGH-PGRNGTLEIVSRKFFWPGQSQTVRRFVRNCDVCGGIHI 331
Query: 1352 KKNAPANELTSVFSPWPFHK-WGVDIVGPFPQAPGQ-LKFLIVDVDYFTKWVEAEAVSKI 1409
+ A L + P H +D + P G+ ++L V VD +K V E + +
Sbjct: 332 WRQAKRGFLKPLPVPNRLHSDLSMDFITSLPPTRGRGSQYLWVIVDRLSKSVTLEEMDTM 511
Query: 1410 TAERVVKFYWKKIICHF---GLPKYIVTDNGTQFASSKVVNFCKQLGIETKFVSVIHPQA 1466
AE + + + CH+ G+P+ IV+D G+ + FC+ G+ + HPQ
Sbjct: 512 EAEACAQRF---LSCHYRFHGMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQT 682
Query: 1467 NGQAESANKMIVNDIKKKLEDAKGLWAEQLHEVLWSYHTTPHSTTGETPFTMVYG 1521
+G E N+ I ++ + ++ W + L V + +S+ G TPF + +G
Sbjct: 683 DGGTERWNQEIQAVLRAYVCWSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHG 847
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 72.8 bits (177), Expect = 1e-12
Identities = 51/167 (30%), Positives = 77/167 (45%), Gaps = 29/167 (17%)
Frame = -2
Query: 997 KLALAVIITARKLRPYFQSHKV-VIRTNYPVKQILGKLDLAGRMLSWSVELSEYDIQFAP 1055
K A+ A++LR Y +H ++ P+K I K L GR+ W + LSEYDI++
Sbjct: 635 KTCCALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRS 456
Query: 1056 RNNIKSQVLADFV----------VEFTSPIQEAV------------------PHVWLLSV 1087
+ IK +LAD + ++F P +E + VW L
Sbjct: 455 QKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIF 276
Query: 1088 DGSSNIKGSGAGIILEGPGDLLIEQSLKFDFKASNNQAEYEALIAGM 1134
DG+ N+ G+G G +L P I + + F +NN AEYEA I G+
Sbjct: 275 DGAVNVYGNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGI 135
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 65.5 bits (158), Expect = 2e-10
Identities = 34/101 (33%), Positives = 55/101 (53%)
Frame = -1
Query: 604 KIGTSLSGLEEEILVQLLRENVDLFAWAPSDMPGIDIGVACHHLAVRTSVKPVVQRKRKM 663
K+G +L + + QLLRE +D+FA + DMPG+D + H + + PV + R+
Sbjct: 303 KVGAALEEGVKRKIFQLLREYLDIFACSYEDMPGLDPKIVEHRIPTKPECPPVR*KLRRT 124
Query: 664 GEEKRKAVDEEVKKLQEAHFICEIKYPTWLANTVLVKKASG 704
+ + EV+K +A F+ ++YP W+AN V V K G
Sbjct: 123 HPDMALKIKSEVQKQIDAGFLMTVEYPEWVANIVPVPKKDG 1
>TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNase H
{Arabidopsis thaliana}, partial (21%)
Length = 1071
Score = 55.8 bits (133), Expect = 1e-07
Identities = 40/111 (36%), Positives = 56/111 (50%), Gaps = 5/111 (4%)
Frame = +2
Query: 1091 SNIKGSGAG---IILEGPGDLL--IEQSLKFDFKASNNQAEYEALIAGMLLAQEMGAKNL 1145
SNI AG ++L G LL Q L K S AEY AL+ G+ A G K +
Sbjct: 578 SNINRGPAGAGALLLAEDGSLLYGFRQGLGHQTKES---AEYRALLLGLKHASMKGFKYV 748
Query: 1146 RARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESN 1196
A+ DS+L+ NQI ++ KD+ L K A +L+ +F F ++ RE N
Sbjct: 749 TAKGDSELVINQILDPWKIKDEHLKKLCAEALELSDNFHSFRIQHISRERN 901
>BE999414
Length = 613
Score = 55.5 bits (132), Expect = 2e-07
Identities = 27/75 (36%), Positives = 42/75 (56%)
Frame = +3
Query: 62 PQLPTFDGRSDPSEHVTVFNTRMSVYGVADSLKCKLLAGTLADAALRWYMSLPRFSIVGY 121
PQ+ ++DG DP EH+ ++ V ++KCKL TL A+ W+ +L R SI +
Sbjct: 3 PQMDSYDGT*DPDEHMENIEVVLTYRSVRGAVKCKLFVTTLRRGAMTWFKNLRRNSIGSW 182
Query: 122 QDLTKKFTQQFSGTR 136
DL +FT F+ +R
Sbjct: 183 GDLCHEFTTHFTVSR 227
>AL366605
Length = 422
Score = 52.4 bits (124), Expect = 1e-06
Identities = 31/115 (26%), Positives = 48/115 (40%)
Frame = -3
Query: 55 VPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVADSLKCKLLAGTLADAALRWYMSLP 114
VP FK P ++G + P H+ + +M Y DSL +L + A WY SL
Sbjct: 345 VPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEWYTSLS 166
Query: 115 RFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQGPNESLREYLARFNDSTIK 169
+ I + +L F + K L ++ Q ES REY R+ + +
Sbjct: 165 KNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAAAR 1
>BG448304
Length = 637
Score = 50.1 bits (118), Expect = 7e-06
Identities = 28/77 (36%), Positives = 37/77 (47%), Gaps = 1/77 (1%)
Frame = +2
Query: 290 PPPFNRSKTMGQDKNAWCKYHLIEGHNTDDCVHLKREIEKLLLNGKLRGYAKEKHHSERR 349
PP + G DK +C++H H TDDC+HLK IE L+ G+L K
Sbjct: 62 PPKTPTQENKGTDKTKYCRFHKCHRHLTDDCIHLKDTIEILIQRGRLNQLTK-------- 217
Query: 350 EDKPNPEP-KHTLHTIS 365
NPEP K T+ I+
Sbjct: 218 ----NPEPEKQTVKLIT 256
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 44.7 bits (104), Expect(2) = 3e-05
Identities = 19/41 (46%), Positives = 25/41 (60%)
Frame = -1
Query: 694 ANTVLVKKASGKWRMCVDYTDLNMACPKDPYPLPSIDHLID 734
A + V+K G +RMC+DY N K+ YPLP ID+L D
Sbjct: 238 AALLFVRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFD 116
Score = 22.7 bits (47), Expect(2) = 3e-05
Identities = 7/19 (36%), Positives = 14/19 (72%)
Frame = -3
Query: 750 GYNQIKMDPLDAPKTAFMT 768
GY+Q +++ ++ PKT +T
Sbjct: 71 GYHQFRVNEVNIPKTTLLT 15
>BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana},
partial (6%)
Length = 469
Score = 40.8 bits (94), Expect = 0.004
Identities = 25/86 (29%), Positives = 47/86 (54%), Gaps = 4/86 (4%)
Frame = +1
Query: 1553 MDMIDEVREAAHIREFAAKQRAARRYNSKVIPRSMKEGDLVLKQVVAPTRI----GKLLP 1608
++ ++E+R A+ K+R + ++ ++I R +EG+LVL + +R+ GKL
Sbjct: 106 LNELEELRLDAYENAKIYKERTKKWHDRRIIRREFREGELVL---LFNSRLKLFPGKLRS 276
Query: 1609 SWKGPYRVKEKLQHGAYKLEELSGNP 1634
W GP++VK + GA ++ S P
Sbjct: 277 HWSGPFQVKNVMPSGAVEVWSESTGP 354
>BG644720
Length = 678
Score = 40.4 bits (93), Expect = 0.006
Identities = 37/144 (25%), Positives = 53/144 (36%)
Frame = -3
Query: 1242 WMTPLLKFLTGSFVAKNDEYAQLVRRRATKFVVIAGKLYKRGRASPLLRCLGEGETELVL 1301
W P++ ++ +N + +RR LY LL CLGE ET
Sbjct: 463 WRQPIIIYMCYGIFLENPRRSTYIRRHTPHLH*YNETLYI*LFEGVLL*CLGEEETIQAF 284
Query: 1302 LEVHEGVCGSHIGGRSLAAKLLRAGYYWPRMAHDCCEFVKKCDKCQRFSDKKNAPANELT 1361
H VCGSH L + R GYYW + K F + P L
Sbjct: 283 Q*AHSRVCGSH--KSKLYFHIKRMGYYWN------T*IMLKGALLANFIQQ---PPEVLH 137
Query: 1362 SVFSPWPFHKWGVDIVGPFPQAPG 1385
+ F W +++ G FP++ G
Sbjct: 136 PTITS*LFESWELNVFGLFPKSSG 65
>BG580885
Length = 609
Score = 38.9 bits (89), Expect = 0.017
Identities = 36/143 (25%), Positives = 59/143 (41%), Gaps = 7/143 (4%)
Frame = +3
Query: 64 LPTFDG--RSDPSEHVTVFNTRMSVYGVAD-SLKCKLLAGTLADAALRWY-MSL-PRFSI 118
LP F G P H++ FN + ++ K+ TL + + WY +++ P +
Sbjct: 45 LPIFRGTPNESPITHLSRFNKVCRANNASSVEMQKKIFPVTLEEESALWYDLNIEPYYIS 224
Query: 119 VGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQGPNESLREYLARFNDSTIKVSNPNQE-- 176
+ + ++ F Q + + L + L + QG E +R Y R + E
Sbjct: 225 LSWDEIKLSFLQAYYEIEPVEELRSELMGIHQGEKERVRSYFLRLQWILKRWPEHGLEDD 404
Query: 177 VFVGAFQNGLRAGQFNESLAQKP 199
V G F NGLR + L QKP
Sbjct: 405 VIKGVFVNGLREEFHDWVLMQKP 473
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.135 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,897,396
Number of Sequences: 36976
Number of extensions: 664576
Number of successful extensions: 3384
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 3296
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3374
length of query: 1638
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1529
effective length of database: 4,984,343
effective search space: 7621060447
effective search space used: 7621060447
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC141323.18


