

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
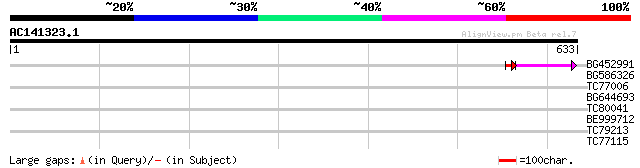
Query= AC141323.1 + phase: 0 /pseudo
(633 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, pa... 48 5e-06
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 38 0.010
TC77006 similar to SP|P21745|EA30_VICFA Embryonic abundant prote... 31 1.7
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 30 2.8
TC80041 similar to GP|8778702|gb|AAF79710.1| T1N15.20 {Arabidops... 30 3.7
BE999712 homologue to PIR|T09361|T0 hypothetical protein F23K16.... 29 6.3
TC79213 similar to GP|15706274|emb|CAC69997. HMG I/Y like protei... 28 8.2
TC77115 similar to GP|9294182|dbj|BAB02084.1 protein kinases-lik... 28 8.2
>BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, partial (33%)
Length = 560
Score = 48.1 bits (113), Expect(2) = 5e-06
Identities = 37/68 (54%), Positives = 40/68 (58%)
Frame = +1
Query: 565 LNSSEI*A*CVKCHLKV*S*EC*RLIMNF*KVLKRHRKLM*SLWTCWLLEIRLKTVILKS 624
LNSSE * VKC +V*+ C*R NF V KR R+ M*S L IRLK VILK
Sbjct: 70 LNSSET*VWFVKCRHRV*NWGC*RSTTNFWIVSKRLRRWM*SW*I*CLGIIRLKMVILKL 249
Query: 625 MIKVC*GF 632
MIK C F
Sbjct: 250 MIKECCSF 273
Score = 20.4 bits (41), Expect(2) = 5e-06
Identities = 8/12 (66%), Positives = 10/12 (82%)
Frame = +3
Query: 554 CLL*W*KNSSCL 565
CLL*W ++ SCL
Sbjct: 36 CLL*WLESWSCL 71
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 38.1 bits (87), Expect = 0.010
Identities = 43/115 (37%), Positives = 54/115 (46%)
Frame = +3
Query: 435 LWYIVMRPSWI*VVCLCKMVK**LMLRDS*EFMRRIIPRTI*SWLLWSLY*RYGDIICTV 494
+W+I R S *V M + + S*E MR P I* WL *R+G C V
Sbjct: 48 MWFIQTRLSLD*VAY*PSMRRSSPTRQGS*ENMRETTPPMI*KWLR*YSP*RFGAHTCMV 227
Query: 495 RDLKCLVIIRV*SICSIRRN*I*GSVDG*NC*RIMILS*AIIRVKLMLLQML*VG 549
+ + I+V*SI +* *G G N * I + IIR KL+ Q L* G
Sbjct: 228 PRFRYIRTIKV*SIFLPSLS*T*GRGGGWNS*LTTI*TSLIIREKLIW*QTL*AG 392
>TC77006 similar to SP|P21745|EA30_VICFA Embryonic abundant protein VF30.1
precursor. [Broad bean] {Vicia faba}, complete
Length = 1136
Score = 30.8 bits (68), Expect = 1.7
Identities = 13/27 (48%), Positives = 17/27 (62%)
Frame = -3
Query: 244 CRRQPSERGMLTMSTRLCLSVLRMHLE 270
C R P G+L S+RLCLS+ H+E
Sbjct: 423 CLRSPVAVGLLQKSSRLCLSLFASHME 343
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 30.0 bits (66), Expect = 2.8
Identities = 49/155 (31%), Positives = 70/155 (44%)
Frame = +3
Query: 157 NN*KISLRRSLLDQVFRLGERRFC**RRKMVA*DCVLTIDS*TR*Q*RIGINFQGLMT*W 216
+N KI +R + VF C *RRKM + + LTI +* *R I L
Sbjct: 126 SNSKIY*KRVSFNLVFIHRVLWSCF*RRKMGSLE*ALTIPN*ITST*R*SILSLLLTNYL 305
Query: 217 IS*WVQKFSAR*I*DQVITILK*KMKICRRQPSERGMLTMSTRLCLSVLRMHLEYLWST* 276
I V +R * T + * +K+ + SE M+TM + CL ++ +LW+ *
Sbjct: 306 IISKVLNGFSRLT*GWDNTNIG**VKMFLKLLSELDMVTMKS**CLLGKQIPRWHLWN** 485
Query: 277 FASFMHFWIGSWLFSLMIF*FTPSLKKNMLSI*SW 311
LF +MIF*+TP +K +M * W
Sbjct: 486 IEFSKITSTH**LFLVMIF*YTPRMKMSMRIT*GW 590
>TC80041 similar to GP|8778702|gb|AAF79710.1| T1N15.20 {Arabidopsis thaliana},
partial (7%)
Length = 1621
Score = 29.6 bits (65), Expect = 3.7
Identities = 15/43 (34%), Positives = 21/43 (47%)
Frame = +3
Query: 255 TMSTRLCLSVLRMHLEYLWST*FASFMHFWIGSWLFSLMIF*F 297
TMSTR V + + *F + H W +W+F L + *F
Sbjct: 1290 TMSTRYTQHVFCKYFHTILPG*FGIWKHIWDCNWIFQLQLS*F 1418
>BE999712 homologue to PIR|T09361|T0 hypothetical protein F23K16.80 -
Arabidopsis thaliana, partial (13%)
Length = 623
Score = 28.9 bits (63), Expect = 6.3
Identities = 10/25 (40%), Positives = 19/25 (76%)
Frame = -3
Query: 103 NYKWYAIFQKFFLMKFLMCLQREKL 127
++ W ++ QKF L+K L+C +R++L
Sbjct: 189 SWHWMSLHQKFCLLKGLLCFKRKQL 115
>TC79213 similar to GP|15706274|emb|CAC69997. HMG I/Y like protein {Glycine
max}, partial (16%)
Length = 811
Score = 28.5 bits (62), Expect = 8.2
Identities = 21/57 (36%), Positives = 27/57 (46%), Gaps = 6/57 (10%)
Frame = -2
Query: 102 INYKWYA------IFQKFFLMKFLMCLQREKLSFRLILFLERSLCL*HLIVCQLLSW 152
I YKW IF + + +L L L + L + L LCL*HL+ QLL W
Sbjct: 591 IFYKWVTKLRPM*IFLAWG*IMWLKQLLHLLLQWHLAILLFIILCL*HLLGMQLLRW 421
>TC77115 similar to GP|9294182|dbj|BAB02084.1 protein kinases-like protein
{Arabidopsis thaliana}, partial (68%)
Length = 1285
Score = 28.5 bits (62), Expect = 8.2
Identities = 13/48 (27%), Positives = 25/48 (52%)
Frame = +3
Query: 79 WNVMTF*CFP*WLPYLLRIKQ*LINYKWYAIFQKFFLMKFLMCLQREK 126
WN F L +RI + ++ W +F K+F +K +M ++R++
Sbjct: 1065 WNWQDHLIFLH*LKLRMRIIRGIVEIMWRYLFMKYFFLKVMMMMKRKR 1208
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.382 0.171 0.711
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,708,131
Number of Sequences: 36976
Number of extensions: 335719
Number of successful extensions: 5643
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1274
Number of HSP's successfully gapped in prelim test: 206
Number of HSP's that attempted gapping in prelim test: 4299
Number of HSP's gapped (non-prelim): 1626
length of query: 633
length of database: 9,014,727
effective HSP length: 102
effective length of query: 531
effective length of database: 5,243,175
effective search space: 2784125925
effective search space used: 2784125925
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC141323.1


