

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141322.10 + phase: 0 /pseudo
(409 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
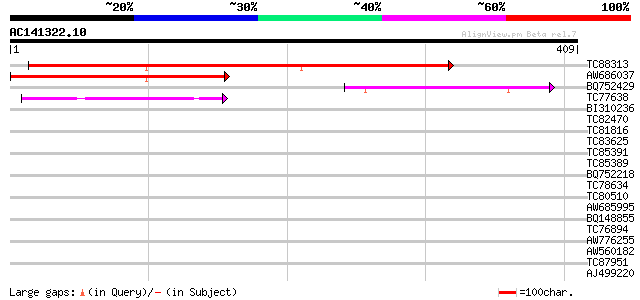
Sequences producing significant alignments: (bits) Value
TC88313 weakly similar to PIR|E84776|E84776 hypothetical protein... 380 e-106
AW686037 similar to PIR|E64203|E642 ATP-dependent nuclease addA ... 256 1e-68
BQ752429 weakly similar to PIR|T39430|T39 mitochondrial import i... 78 7e-15
TC77638 similar to GP|20161455|dbj|BAB90379. ankyrin-like protei... 42 4e-04
BI310236 similar to SP|Q9UZC8|RA50_ DNA double-strand break repa... 38 0.008
TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical prot... 37 0.014
TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana taba... 35 0.053
TC83625 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana taba... 35 0.053
TC85391 homologue to PIR|T06598|T06598 dnaK-type molecular chape... 33 0.15
TC85389 homologue to SP|Q03684|BIP4_TOBAC Luminal binding protei... 33 0.26
BQ752218 similar to GP|19170914|emb hypothetical protein {Enceph... 33 0.26
TC78634 similar to PIR|T48567|T48567 ABA-responsive protein homo... 32 0.45
TC80510 similar to PIR|T05005|T05005 hypothetical protein T19P19... 32 0.45
AW685995 similar to PIR|I51618|I516 nucleolar phosphoprotein - A... 32 0.45
BQ148855 homologue to SP|P12012|GNTP_ Gluconate permease. {Bacil... 31 0.76
TC76894 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protea... 31 1.00
AW776255 weakly similar to PIR|T07672|T07 cyclin a2-type mitosi... 30 1.7
AW560182 similar to PIR|C96543|C96 unknown protein [imported] - ... 30 2.2
TC87951 similar to PIR|C86418|C86418 hypothetical protein AAG126... 30 2.2
AJ499220 similar to SP|Q09863|YAF Hypothetical protein C29E6.10c... 30 2.2
>TC88313 weakly similar to PIR|E84776|E84776 hypothetical protein At2g36070
[imported] - Arabidopsis thaliana, partial (31%)
Length = 1201
Score = 380 bits (976), Expect = e-106
Identities = 213/327 (65%), Positives = 241/327 (73%), Gaps = 20/327 (6%)
Frame = +2
Query: 14 RGYSVFNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFD 73
R Y VFNEFS KVKDETVKNPEFQKSVKELKEKAEELKG+KEGLKEKTKQTTEQLY+QFD
Sbjct: 221 RRYGVFNEFSNKVKDETVKNPEFQKSVKELKEKAEELKGVKEGLKEKTKQTTEQLYKQFD 400
Query: 74 SVWKEAEAAAKKVSHNVKEKISAA----------------TDFSTKQNADAKQGSQKSPE 117
VWKEAEAAAKKVSHNVKEKISAA TD STKQ+ADAKQG Q SPE
Sbjct: 401 GVWKEAEAAAKKVSHNVKEKISAATEEVKAGIGKQDSSGSTDSSTKQDADAKQGRQTSPE 580
Query: 118 EEKNEESPSGNASESLFGKFKSTFSSPMVSTSFQKLKDAKLWT*PRKVMTY*RRS*V-VT 176
EEKN+ES SGNASESLFGKFKSTFSSP VSTSFQKLKDAK+ +K + T
Sbjct: 581 EEKNQESASGNASESLFGKFKSTFSSPKVSTSFQKLKDAKIVDMTKKGYDILKEELSGNT 760
Query: 177 HLRDSVFVSPA-K*VQKLILLLCPLTNLGGVRRL--MSSGIRRHPVSKNFVKYIDPVKTT 233
R V +P+ + K L++ P ++ + ++ HP SK F KY DPVKT
Sbjct: 761 PKRQPVHSTPSGETSTKTDLVVMPSNQSWWSKKFDEIRDKVKSHPASKRFFKYTDPVKTK 940
Query: 234 SRDIVDDVRDIIDRIDNPIIDKIQSNMSRTFQETDAALTYREIRKRDPKFSLPDFVGEVQ 293
S+++VDD+RD I+ DNPII+KIQ FQETDAAL ++EI +RDPKFSLP+FV EVQ
Sbjct: 941 SQEMVDDLRDRIETSDNPIINKIQDINDTIFQETDAALAHKEIHRRDPKFSLPEFVAEVQ 1120
Query: 294 EAIKPVLNAYIKGDFETLKKYCAPQLI 320
EAIKPVLNAYIKGD ETLKKYC+P LI
Sbjct: 1121EAIKPVLNAYIKGDVETLKKYCSP*LI 1201
>AW686037 similar to PIR|E64203|E642 ATP-dependent nuclease addA homolog -
Mycoplasma genitalium, partial (2%)
Length = 682
Score = 256 bits (653), Expect = 1e-68
Identities = 134/174 (77%), Positives = 145/174 (83%), Gaps = 16/174 (9%)
Frame = +1
Query: 1 GVKTRLFVNSVDRRGYSVFNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEK 60
G + RLF N VDRRGYSVFNEFSKKVKDET++NPEFQ+SVKELKEKAEELKG+KEGLKEK
Sbjct: 97 GFRPRLFFNLVDRRGYSVFNEFSKKVKDETLRNPEFQQSVKELKEKAEELKGVKEGLKEK 276
Query: 61 TKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKISAA----------------TDFSTKQ 104
TKQTTEQLYRQFD +WKEAE AAKKVSHNV+EKISAA TD STK
Sbjct: 277 TKQTTEQLYRQFDGMWKEAEDAAKKVSHNVREKISAATEEVKVGIGKQDSSGSTDSSTKL 456
Query: 105 NADAKQGSQKSPEEEKNEESPSGNASESLFGKFKSTFSSPMVSTSFQKLKDAKL 158
+ADAKQGSQ SPEEEKN+E SGN SES+FGKFKSTFSSP VSTSFQKLKDAK+
Sbjct: 457 DADAKQGSQTSPEEEKNQEFASGNTSESMFGKFKSTFSSPNVSTSFQKLKDAKI 618
>BQ752429 weakly similar to PIR|T39430|T39 mitochondrial import inner
membrane translocase subunit precursor - fission yeast,
partial (32%)
Length = 983
Score = 77.8 bits (190), Expect = 7e-15
Identities = 48/159 (30%), Positives = 80/159 (50%), Gaps = 7/159 (4%)
Frame = -2
Query: 242 RDIIDRIDNPIIDK---IQSNMSRTFQETDAALTYREIRKRDPKFSLPDFVGEVQEAIKP 298
R + DNP+I+ + + F E + A ++ R DP F F+ E++E I P
Sbjct: 958 RQVYRESDNPLIETARLVTDKIGSFFAENETAQVIKKFRSMDPNFQTEPFLKELREYILP 779
Query: 299 -VLNAYIKGDFETLKKYCAPQLIERCKAEHGAYKDRGIFYDNKILHISDADVREVKILES 357
VL+AY+KGD TLK + + +A Y G+ D KIL I D+ ++L+
Sbjct: 778 EVLDAYVKGDVPTLKLWLSEAQYSVYEALTKQYLQAGLKSDGKILDIRHVDIVRARMLDP 599
Query: 358 S--PFIIVVFQTQQIHCVRD-RNGEITEGGKDTIHSVYY 393
P ++ +TQ++H R+ ++GE+ G +D + V Y
Sbjct: 598 GEIPVFVITCRTQEVHVYRNAKSGELAAGMEDKVQLVTY 482
>TC77638 similar to GP|20161455|dbj|BAB90379. ankyrin-like protein {Oryza
sativa (japonica cultivar-group)}, partial (62%)
Length = 2345
Score = 42.0 bits (97), Expect = 4e-04
Identities = 35/149 (23%), Positives = 61/149 (40%)
Frame = +1
Query: 9 NSVDRRGYSVFNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQL 68
NS ++ E K ++ N E K +E K E EK ++ E
Sbjct: 616 NSTEKSSEDTKTEDEGKKTEDEGSNTENNKDGEEASTKESE-----SDESEKKDESEENN 780
Query: 69 YRQFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGN 128
D K++ + + NV+EK+ + + + +NA K + ++ NE PSG
Sbjct: 781 KSDSDESEKKSSDSNETTDSNVEEKVEQSQNKESDENASEKNTDDNAKDQSSNEVFPSGA 960
Query: 129 ASESLFGKFKSTFSSPMVSTSFQKLKDAK 157
SE L ++T + ST ++K+ K
Sbjct: 961 QSELL---NETTTQTGSFSTQCSRVKE*K 1038
>BI310236 similar to SP|Q9UZC8|RA50_ DNA double-strand break repair rad50
ATPase. {Pyrococcus abyssi}, partial (2%)
Length = 819
Score = 37.7 bits (86), Expect = 0.008
Identities = 31/111 (27%), Positives = 44/111 (38%)
Frame = +2
Query: 25 KVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAK 84
+VK+ K EF+ VKEL+ + + K E K K K Q + W E E+
Sbjct: 155 QVKERESKEREFEGQVKELESRKKHFKSQVEEFKSKEK--------QLEGRWSELESKEN 310
Query: 85 KVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASESLFG 135
K VKE F D + EE+++ ES + S FG
Sbjct: 311 KFKAKVKELNLKEKQFEGLVK-DPASRKKYIDEEKESVESYMDDQSSRAFG 460
>TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (8%)
Length = 1064
Score = 37.0 bits (84), Expect = 0.014
Identities = 26/89 (29%), Positives = 41/89 (45%), Gaps = 6/89 (6%)
Frame = +2
Query: 21 EFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYR------QFDS 74
EF ++ + K F+ V+E+ K LKG + L K KQ Q+ QF++
Sbjct: 587 EFEGRMNELESKERHFKSEVEEINAKLMPLKGQIKELASKEKQLNGQVKELESKKNQFEN 766
Query: 75 VWKEAEAAAKKVSHNVKEKISAATDFSTK 103
KE E+ K+ VKE S +F ++
Sbjct: 767 RIKELESKEKQHEGRVKEHASKEREFESQ 853
>TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana tabacum},
partial (13%)
Length = 663
Score = 35.0 bits (79), Expect = 0.053
Identities = 26/103 (25%), Positives = 47/103 (45%), Gaps = 2/103 (1%)
Frame = +3
Query: 27 KDETVKNPEFQKSVKELKEKAEELK-GIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKK 85
KD +K+ E KE K+K ++ K + EG +K K+ ++ ++ + +E + KK
Sbjct: 144 KDLEIKSVEKDDEKKEKKDKEKKDKTDVDEGKDKKDKEKKKKEKKEENVKGEEEDGDEKK 323
Query: 86 VSHNVK-EKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSG 127
K EK + K + K K +++KNE+ G
Sbjct: 324 DKEKKKKEKKEKGKEDKDKDGEEKKSKKDKEKKKDKNEDDDEG 452
>TC83625 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana tabacum},
partial (9%)
Length = 908
Score = 35.0 bits (79), Expect = 0.053
Identities = 26/103 (25%), Positives = 47/103 (45%), Gaps = 2/103 (1%)
Frame = +2
Query: 27 KDETVKNPEFQKSVKELKEKAEELK-GIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKK 85
KD +K+ E KE K+K ++ K + EG +K K+ ++ ++ + +E + KK
Sbjct: 536 KDLEIKSVEKDDEKKEKKDKEKKDKTDVDEGKDKKDKEKKKKEKKEENVKGEEEDGDEKK 715
Query: 86 VSHNVK-EKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSG 127
K EK + K + K K +++KNE+ G
Sbjct: 716 DKEKKKKEKKEKGKEDKDKDGEEKKSKKDKEKKKDKNEDDDGG 844
>TC85391 homologue to PIR|T06598|T06598 dnaK-type molecular chaperone BiP-A
- soybean, partial (52%)
Length = 1231
Score = 33.5 bits (75), Expect = 0.15
Identities = 27/89 (30%), Positives = 43/89 (47%)
Frame = +1
Query: 35 EFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKI 94
E ++ V+E +E AEE K +KE + + T +Y + V + + A K+ + KEKI
Sbjct: 676 EIERMVREAEEFAEEDKKVKERIDARNALET-YVYNMKNQV-SDKDKLADKLESDEKEKI 849
Query: 95 SAATDFSTKQNADAKQGSQKSPEEEKNEE 123
A + D Q +K EEK +E
Sbjct: 850 ETAVK-EALEWLDDNQSVEKEEYEEKLKE 933
>TC85389 homologue to SP|Q03684|BIP4_TOBAC Luminal binding protein 4 precursor
(BiP 4) (78 kDa glucose-regulated protein homolog 4) (GRP
78-4)., partial (94%)
Length = 2265
Score = 32.7 bits (73), Expect = 0.26
Identities = 27/89 (30%), Positives = 43/89 (47%)
Frame = +2
Query: 35 EFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKI 94
E + V+E +E AEE K +KE + + T +Y + + + + A K+ + KEKI
Sbjct: 1685 EIDRMVREAEEFAEEDKKVKERIDARNALET-YVYNMKNQI-SDKDKLADKLESDEKEKI 1858
Query: 95 SAATDFSTKQNADAKQGSQKSPEEEKNEE 123
AA + D Q +K EEK +E
Sbjct: 1859 EAAVK-EALEWLDDNQTVEKEEFEEKLKE 1942
>BQ752218 similar to GP|19170914|emb hypothetical protein {Encephalitozoon
cuniculi}, partial (5%)
Length = 637
Score = 32.7 bits (73), Expect = 0.26
Identities = 25/92 (27%), Positives = 43/92 (46%), Gaps = 2/92 (2%)
Frame = -2
Query: 24 KKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAA 83
KK ++E K + + K K E K+ ++K K+ Q ++ KEAE A
Sbjct: 576 KKAREEAAKKAKKAREAAYKKAKEE-----KKEAEKKAKEEKRQAEKKIKEEKKEAEKKA 412
Query: 84 K--KVSHNVKEKISAATDFSTKQNADAKQGSQ 113
K K + K+ SA + T+ + ++K GS+
Sbjct: 411 KKDKKEQDRKDAESALSRQDTEDSKESKDGSK 316
Score = 28.1 bits (61), Expect = 6.5
Identities = 24/79 (30%), Positives = 35/79 (43%)
Frame = -2
Query: 44 KEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKISAATDFSTK 103
+ K +E K KE K+K K E + K EAA KK KE + K
Sbjct: 627 RPKKKEEKAKKEAEKKKKKAREEAAKK----AKKAREAAYKKAKEEKKE--------AEK 484
Query: 104 QNADAKQGSQKSPEEEKNE 122
+ + K+ ++K +EEK E
Sbjct: 483 KAKEEKRQAEKKIKEEKKE 427
>TC78634 similar to PIR|T48567|T48567 ABA-responsive protein homolog
T31B5.20 - Arabidopsis thaliana, partial (67%)
Length = 1208
Score = 32.0 bits (71), Expect = 0.45
Identities = 24/88 (27%), Positives = 41/88 (46%), Gaps = 3/88 (3%)
Frame = +2
Query: 66 EQLYRQFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAK---QGSQKSPEEEKNE 122
E + FDS K+AEA A + HN+K S ++ K N K +G +S ++
Sbjct: 434 ESILHMFDSWSKKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFT 613
Query: 123 ESPSGNASESLFGKFKSTFSSPMVSTSF 150
P+ ++ F + ST + P+ T +
Sbjct: 614 TYPNEKLKKT-FACYLSTTTGPVAGTLY 694
>TC80510 similar to PIR|T05005|T05005 hypothetical protein T19P19.70 -
Arabidopsis thaliana, partial (8%)
Length = 889
Score = 32.0 bits (71), Expect = 0.45
Identities = 21/101 (20%), Positives = 45/101 (43%)
Frame = +3
Query: 40 VKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKISAATD 99
V ELKE+ + K + +GLK E L + D +E AA+ +E A+ +
Sbjct: 492 VTELKEELKRRKLVTKGLK-------EDLINRLDEALREEREAAEAARKKEQEAAEASPE 650
Query: 100 FSTKQNADAKQGSQKSPEEEKNEESPSGNASESLFGKFKST 140
+ + + + + ++ ++S + N +FG +++
Sbjct: 651 QEAAEGSKKDEANGLDTQVDELKDSKTINVDAEVFGTIQAS 773
>AW685995 similar to PIR|I51618|I516 nucleolar phosphoprotein - African
clawed frog, partial (5%)
Length = 516
Score = 32.0 bits (71), Expect = 0.45
Identities = 37/140 (26%), Positives = 62/140 (43%), Gaps = 21/140 (15%)
Frame = +3
Query: 4 TRLFVNSVDRRGYSVFNEFSKKV-KDETVKNPEFQKSVKELK--EKAEELKGIKEGLKEK 60
T F+ + V E KKV K E K + K+ +E K +K +E K ++E
Sbjct: 6 TNPFIMGKSKTSKKVKAEEPKKVEKVEKTKGKKAAKAEEEKKKQQKKKEEKPAPMEVEED 185
Query: 61 TKQTTEQLYRQFDSVWKEA----EAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQK-- 114
+ + E + ++ K+A +AAAKK S + +E S+ ++ ++ K ++K
Sbjct: 186 SSSSEESSSSEDEAPAKKAAPAKKAAAKKESSSEEESSSSESESEEEEKPAKKAAAKKES 365
Query: 115 ------------SPEEEKNE 122
S EEEKNE
Sbjct: 366 SSEEESSSSEEESDEEEKNE 425
>BQ148855 homologue to SP|P12012|GNTP_ Gluconate permease. {Bacillus
subtilis}, partial (2%)
Length = 692
Score = 31.2 bits (69), Expect = 0.76
Identities = 20/69 (28%), Positives = 36/69 (51%)
Frame = +3
Query: 24 KKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAA 83
+K+ ++ VK EF+ +E K + + LKG +++ + +QF+ KE E+
Sbjct: 3 EKLLEDRVK--EFESKEEEFKARMQNLKGFVSQMEDFKSEE-----KQFEGRGKEPESKD 161
Query: 84 KKVSHNVKE 92
KK +VKE
Sbjct: 162KKFKAHVKE 188
>TC76894 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protease
ATP-binding subunit clpA homolog chloroplast precursor.
[Garden pea], partial (71%)
Length = 2413
Score = 30.8 bits (68), Expect = 1.00
Identities = 17/37 (45%), Positives = 24/37 (63%)
Frame = +3
Query: 27 KDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQ 63
KDE V+N EF+K+ EL++K +LK L EK K+
Sbjct: 1761 KDEAVRNQEFEKA-GELRDKEMDLKTQISALIEKNKE 1868
>AW776255 weakly similar to PIR|T07672|T07 cyclin a2-type mitosis-specific -
soybean, partial (22%)
Length = 704
Score = 30.0 bits (66), Expect = 1.7
Identities = 31/156 (19%), Positives = 61/156 (38%), Gaps = 18/156 (11%)
Frame = +1
Query: 23 SKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAA 82
SK K+ V + + LKE + + + GLK+ T + + R S +++ E +
Sbjct: 58 SKMDKENGVNVRVTRARARALKEPSRNEQAVVVGLKDVTNISAKSHKRTHTSNFQKKEGS 237
Query: 83 AKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNAS-ESLFGKFKSTF 141
K+ ++ E + ++ K++A + S +ES S E+ + T
Sbjct: 238 KKRKTNVASEDDVSLQVWTIKEDAKKELAKDSSTSTMTKKESLQVQPSVENSLLSMQDTL 417
Query: 142 SSPMVSTSF-----------------QKLKDAKLWT 160
+SP + KL+D+ +WT
Sbjct: 418 NSPNTEINLICEKLSASVGLGIVDIDSKLRDSPIWT 525
>AW560182 similar to PIR|C96543|C96 unknown protein [imported] - Arabidopsis
thaliana, partial (32%)
Length = 706
Score = 29.6 bits (65), Expect = 2.2
Identities = 25/87 (28%), Positives = 35/87 (39%), Gaps = 11/87 (12%)
Frame = +2
Query: 35 EFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNV--KE 92
E K + E K + EELK L+E+ + L Q VW+E K + V E
Sbjct: 257 ELAKEIGEDKAEVEELKRESMNLREEMDEERRML--QMAEVWREERVHMKLIDAKVALDE 430
Query: 93 KISAATDF---------STKQNADAKQ 110
K S + S N+DAK+
Sbjct: 431 KYSQMNELVAYLETFLKSVNMNSDAKE 511
>TC87951 similar to PIR|C86418|C86418 hypothetical protein AAG12629.1
[imported] - Arabidopsis thaliana, partial (13%)
Length = 922
Score = 29.6 bits (65), Expect = 2.2
Identities = 24/89 (26%), Positives = 43/89 (47%), Gaps = 1/89 (1%)
Frame = +3
Query: 19 FNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKE-KTKQTTEQLYRQFDSVWK 77
FNE S++V++ VK K +EK E +G KEG+ E + +E+ + DS K
Sbjct: 240 FNEGSEEVEEIEVKPSAAAVVAKSSEEKVGEAEGGKEGVVEIEWDMKSEESLEKIDSP-K 416
Query: 78 EAEAAAKKVSHNVKEKISAATDFSTKQNA 106
+ ++K + + I T ++K +
Sbjct: 417 DLNYGSRKNNGGSNDVIETGTAKNSKDES 503
>AJ499220 similar to SP|Q09863|YAF Hypothetical protein C29E6.10c in
chromosome I. [Fission yeast] {Schizosaccharomyces
pombe}, partial (12%)
Length = 518
Score = 29.6 bits (65), Expect = 2.2
Identities = 24/110 (21%), Positives = 53/110 (47%), Gaps = 5/110 (4%)
Frame = -3
Query: 18 VFNEFSKKVKDETVKNPEFQKSVKELKEK-----AEELKGIKEGLKEKTKQTTEQLYRQF 72
V N + +KV E QK ++EL+E+ ELK +KE ++K K + ++
Sbjct: 312 VLNAYREKVAQERQ-----QKLLEELEEENRLKEERELKKLKEKERKKAKNRALKQQKEE 148
Query: 73 DSVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNE 122
+ +EAE A++ + + + + ++ K+ +++ + +KN+
Sbjct: 147 ERARREAERLAEEKAQRAEREKKLEEERKRREEERLKKEAERR-QRKKND 1
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.136 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,948,798
Number of Sequences: 36976
Number of extensions: 110078
Number of successful extensions: 602
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 590
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 597
length of query: 409
length of database: 9,014,727
effective HSP length: 98
effective length of query: 311
effective length of database: 5,391,079
effective search space: 1676625569
effective search space used: 1676625569
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC141322.10


