

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141115.10 + phase: 0 /pseudo
(173 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
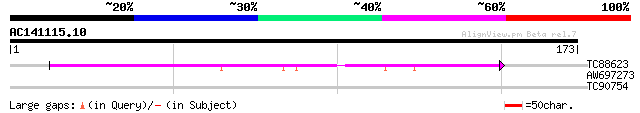
TC88623 similar to PIR|C84609|C84609 hypothetical protein At2g22... 119 8e-28
AW697273 27 4.1
TC90754 similar to GP|1771381|emb|CAA65127.1 phosphoinositide-sp... 27 5.4
>TC88623 similar to PIR|C84609|C84609 hypothetical protein At2g22130
[imported] - Arabidopsis thaliana, partial (24%)
Length = 1960
Score = 119 bits (297), Expect = 8e-28
Identities = 84/182 (46%), Positives = 100/182 (54%), Gaps = 43/182 (23%)
Frame = +1
Query: 13 MDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAEFI-------LV 65
++ALFLL Q WSACP EVSR QS AAA AIP LQ LIQ GP F EKAEF+ LV
Sbjct: 1237 LEALFLLRQAWSACPAEVSRAQSIAAADAIPFLQYLIQSGPPRFQEKAEFLLQCLPGTLV 1416
Query: 66 MIVKRGNNMRQCVGNQG---KITL-------------EANQEWDERFTYMVL*ECSSRTE 109
+IVKRGNNMRQ VG KITL N EW+E FT+ E + +
Sbjct: 1417 VIVKRGNNMRQSVGIPSVYCKITLGNSPPKLTKVVSTGPNPEWEESFTWSF--ESPPKGQ 1590
Query: 110 ASYL-LQKQA*SGK-------------------ADEHTLLPTSKSGQPRNLEVELKWSNK 149
++ + ++ GK A E+TLLP SKSG PRNLE+E +WSNK
Sbjct: 1591 KLHISCKNKSKVGKSKFGKVTIQIDRVVMLGAVAGEYTLLPASKSGPPRNLEIEFQWSNK 1770
Query: 150 PS 151
S
Sbjct: 1771 VS 1776
>AW697273
Length = 937
Score = 26.9 bits (58), Expect = 4.1
Identities = 14/54 (25%), Positives = 27/54 (49%)
Frame = +1
Query: 8 KKLSWMDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAE 61
KK+S + FLL++G CP + ++ P +L F ++FF++ +
Sbjct: 580 KKIS-EENFFLLLRGEGVCPKPPPQKKNLPLGVGAPPRNDLFLFSQIIFFKRGK 738
>TC90754 similar to GP|1771381|emb|CAA65127.1 phosphoinositide-specific
phospholipase C {Nicotiana rustica}, partial (26%)
Length = 968
Score = 26.6 bits (57), Expect = 5.4
Identities = 20/58 (34%), Positives = 29/58 (49%), Gaps = 1/58 (1%)
Frame = +1
Query: 11 SW-MDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAEFILVMI 67
SW M AL LLI+G S CP V D + I L N + F F++ F ++++
Sbjct: 703 SWIMFALTLLIEGVSRCPTCVMSDTNTTPILVITL--NYVIFSK--FYQCRHFRVIVV 864
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.328 0.139 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,549,516
Number of Sequences: 36976
Number of extensions: 67021
Number of successful extensions: 393
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 390
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 391
length of query: 173
length of database: 9,014,727
effective HSP length: 89
effective length of query: 84
effective length of database: 5,723,863
effective search space: 480804492
effective search space used: 480804492
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC141115.10


