

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
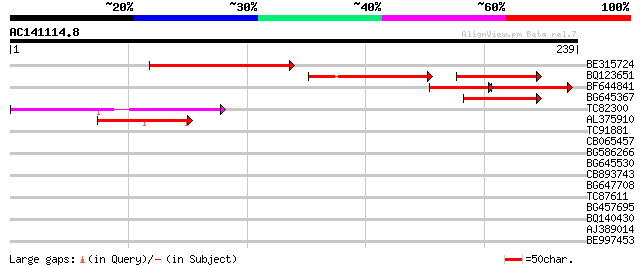
Query= AC141114.8 - phase: 0
(239 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BE315724 116 9e-27
BQ123651 58 2e-17
BF644841 similar to GP|18175662|gb| putative receptor protein ki... 47 2e-11
BG645367 52 2e-07
TC82300 51 3e-07
AL375910 47 9e-06
TC91881 39 0.001
CB065457 30 0.002
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 35 0.033
BG645530 similar to SP|P42347|P3K1_ Phosphatidylinositol 3-kinas... 33 0.097
CB893743 29 1.4
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 29 1.8
TC87611 similar to GP|4406770|gb|AAD20081.1| unknown protein {Ar... 28 2.4
BG457695 28 2.4
BQ140430 similar to GP|21592347|gb| unknown {Arabidopsis thalian... 27 7.0
AJ389014 PIR|A25642|HS histone H4 - maize, partial (98%) 27 7.0
BE997453 27 9.1
>BE315724
Length = 392
Score = 116 bits (290), Expect = 9e-27
Identities = 58/61 (95%), Positives = 59/61 (96%)
Frame = -2
Query: 60 DVVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVKVGDHII 119
+VVLVGESREEVNGRLETWRQALEAY FRLSRSKTEYME NFSGRRSRSTLEVKVGDHII
Sbjct: 184 NVVLVGESREEVNGRLETWRQALEAYEFRLSRSKTEYMECNFSGRRSRSTLEVKVGDHII 5
Query: 120 P 120
P
Sbjct: 4 P 2
>BQ123651
Length = 654
Score = 57.8 bits (138), Expect(2) = 2e-17
Identities = 30/52 (57%), Positives = 38/52 (72%)
Frame = -3
Query: 127 YLGSFVQNDGEIEADVSHRIQAGWLKWRRASGVLCDKKVPLKLKGKFYRTAI 178
YLGS +Q + + DV++RIQA +K R GVLCDKKVPLK+KGKFY T +
Sbjct: 421 YLGSVIQK*QK-QRDVNNRIQARLMK*RSPLGVLCDKKVPLKIKGKFYLTTV 269
Score = 47.8 bits (112), Expect(2) = 2e-17
Identities = 20/36 (55%), Positives = 26/36 (71%)
Frame = -2
Query: 189 WAVKSQHENQVSVTEMRMLRWMSGKTRQDRIRNDTI 224
WA HE ++S + RMLRWM G TR+D+IRND+I
Sbjct: 248 WAE*CHHERRISEAKKRMLRWMCGNTRRDKIRNDSI 141
>BF644841 similar to GP|18175662|gb| putative receptor protein kinase
{Arabidopsis thaliana}, partial (1%)
Length = 676
Score = 47.4 bits (111), Expect(2) = 2e-11
Identities = 19/27 (70%), Positives = 23/27 (84%)
Frame = +2
Query: 178 IRPALLYGTECWAVKSQHENQVSVTEM 204
+RPA+LY ECWAVK+QHEN+VS EM
Sbjct: 158 LRPAMLYAMECWAVKNQHENKVSYVEM 238
Score = 37.7 bits (86), Expect(2) = 2e-11
Identities = 17/34 (50%), Positives = 24/34 (70%)
Frame = +3
Query: 204 MRMLRWMSGKTRQDRIRNDTIREGRGGIHSRKVG 237
M+ML W+ GKTR+ +N+ IRE G ++SRK G
Sbjct: 243 MKMLCWICGKTRRYPFQNNNIRESLGIVNSRKYG 344
>BG645367
Length = 739
Score = 52.0 bits (123), Expect = 2e-07
Identities = 21/33 (63%), Positives = 28/33 (84%)
Frame = +1
Query: 192 KSQHENQVSVTEMRMLRWMSGKTRQDRIRNDTI 224
K+QH+N++S +EMRML WM GKTR RIRN+T+
Sbjct: 640 KNQHKNKISTSEMRMLHWMCGKTRHGRIRNETL 738
>TC82300
Length = 589
Score = 51.2 bits (121), Expect = 3e-07
Identities = 39/99 (39%), Positives = 51/99 (51%), Gaps = 8/99 (8%)
Frame = +2
Query: 1 MYEGVSTSVRTQDGTTEVFPITIGLHQGSTLSPYLF--------TLVLDVLTEHIQELAP 52
MYE + SV T G E FPIT G++ GS LSPY T +V+T
Sbjct: 299 MYERGTASVTTLGGNVEDFPITKGVN*GSVLSPYFSIGSGFVY*T*AREVVT------GS 460
Query: 53 RCMLFADDVVLVGESREEVNGRLETWRQALEAYGFRLSR 91
+ ML +++VLV ES E +N L+ Q LE Y RL+R
Sbjct: 461 QGMLVGNEIVLVEES*EAINKILKIR*QVLETYDNRLTR 577
>AL375910
Length = 244
Score = 46.6 bits (109), Expect = 9e-06
Identities = 29/48 (60%), Positives = 32/48 (66%), Gaps = 8/48 (16%)
Frame = -1
Query: 38 LVLDVLTEHIQELAPRCM------LFADDVVLVGESREEVNG--RLET 77
+ LDVLTE+IQELA RCM F DD VL GE REE+ G RLET
Sbjct: 181 IFLDVLTENIQELALRCMFFFFLFFFTDDAVLHGELREELMGG*RLET 38
>TC91881
Length = 860
Score = 39.3 bits (90), Expect = 0.001
Identities = 18/49 (36%), Positives = 33/49 (66%)
Frame = +3
Query: 173 FYRTAIRPALLYGTECWAVKSQHENQVSVTEMRMLRWMSGKTRQDRIRN 221
+Y T+++ +++ +CWAVKSQ E S+ +MRML + G +++I+N
Sbjct: 315 YYCTSLK-LVIFRIDCWAVKSQ*ERMTSIAKMRMLW*ICGHIGRNKIKN 458
Score = 27.3 bits (59), Expect = 5.3
Identities = 15/35 (42%), Positives = 20/35 (56%)
Frame = +1
Query: 80 QALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVKV 114
Q L F L+RSK EY+ FS + +L+VKV
Sbjct: 16 QTLNFNDFCLTRSKLEYIGCRFSRGHTNVSLDVKV 120
>CB065457
Length = 831
Score = 30.4 bits (67), Expect(2) = 0.002
Identities = 12/16 (75%), Positives = 14/16 (87%)
Frame = +3
Query: 142 VSHRIQAGWLKWRRAS 157
V+H IQ GWLKW+RAS
Sbjct: 48 VNH*IQDGWLKWQRAS 95
Score = 27.7 bits (60), Expect(2) = 0.002
Identities = 12/14 (85%), Positives = 12/14 (85%)
Frame = +1
Query: 125 FKYLGSFVQNDGEI 138
FKYLGS V NDGEI
Sbjct: 1 FKYLGSKVHNDGEI 42
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 34.7 bits (78), Expect = 0.033
Identities = 23/60 (38%), Positives = 30/60 (49%), Gaps = 16/60 (26%)
Frame = -3
Query: 24 GLHQGSTLSPYLFTLVLDVLTEHIQE-----------LAPRC-----MLFADDVVLVGES 67
GL QG LSPYLF L +VL+ Q+ +A C +LFADD + G+S
Sbjct: 385 GLRQGDPLSPYLFILCTEVLSGLCQQALRKGTLPGVKVARNCPPINHLLFADDTMFFGKS 206
>BG645530 similar to SP|P42347|P3K1_ Phosphatidylinositol 3-kinase root
isoform (EC 2.7.1.137) (PI3-kinase) (PtdIns-3-kinase)
(PI3K), partial (7%)
Length = 821
Score = 33.1 bits (74), Expect = 0.097
Identities = 12/19 (63%), Positives = 14/19 (73%)
Frame = +1
Query: 174 YRTAIRPALLYGTECWAVK 192
YRT IRP + Y T+CW VK
Sbjct: 757 YRTVIRPMIFYETKCWGVK 813
>CB893743
Length = 832
Score = 29.3 bits (64), Expect = 1.4
Identities = 15/49 (30%), Positives = 27/49 (54%), Gaps = 4/49 (8%)
Frame = -3
Query: 88 RLSRSKTEYMEWNFSGRRSRSTL----EVKVGDHIIPQVTRFKYLGSFV 132
RL ++ TE +W + R + +TL VK ++IP + R+ G+F+
Sbjct: 260 RLCKATTEPSDWGKTSRYASTTLCKLYMVKTWPNVIPVMLRYSRFGAFI 114
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 28.9 bits (63), Expect = 1.8
Identities = 13/21 (61%), Positives = 15/21 (70%)
Frame = +1
Query: 24 GLHQGSTLSPYLFTLVLDVLT 44
GL QG LSPYLF L +VL+
Sbjct: 13 GLRQGDPLSPYLFILCANVLS 75
>TC87611 similar to GP|4406770|gb|AAD20081.1| unknown protein {Arabidopsis
thaliana}, partial (84%)
Length = 1338
Score = 28.5 bits (62), Expect = 2.4
Identities = 16/60 (26%), Positives = 28/60 (46%)
Frame = +1
Query: 136 GEIEADVSHRIQAGWLKWRRASGVLCDKKVPLKLKGKFYRTAIRPALLYGTECWAVKSQH 195
GEI A + + GW K +G L ++ + + FY +I ++ TEC + +H
Sbjct: 841 GEIIAHLDQKSFKGWQK-NAITGFLTEQNIKIGKPNNFY*VSI*EDVIVNTECADCRQRH 1017
>BG457695
Length = 628
Score = 28.5 bits (62), Expect = 2.4
Identities = 18/58 (31%), Positives = 27/58 (46%), Gaps = 4/58 (6%)
Frame = +3
Query: 94 TEYMEWNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGS----FVQNDGEIEADVSHRIQ 147
TEY +WN + + S GD ++ +T YLG+ VQND S R++
Sbjct: 252 TEYYQWNPTNNQDSSQRSTNSGDILLGSIT---YLGNQEYRLVQNDTTAGVTSSMRVK 416
>BQ140430 similar to GP|21592347|gb| unknown {Arabidopsis thaliana}, partial
(31%)
Length = 409
Score = 26.9 bits (58), Expect = 7.0
Identities = 12/40 (30%), Positives = 22/40 (55%)
Frame = +3
Query: 64 VGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSG 103
+ ESR E+ GR+++ +Q L+ + +L Y + F G
Sbjct: 168 LNESRSELLGRIQSLKQDLQGWRSKLDTQVKVYRDTKFHG 287
>AJ389014 PIR|A25642|HS histone H4 - maize, partial (98%)
Length = 568
Score = 26.9 bits (58), Expect = 7.0
Identities = 17/67 (25%), Positives = 33/67 (48%), Gaps = 4/67 (5%)
Frame = -2
Query: 13 DGTTEVFPITIGLHQGSTLSPYLFT----LVLDVLTEHIQELAPRCMLFADDVVLVGESR 68
+G++ F +H+G+ L P + + + D+L E +Q+ + + D + SR
Sbjct: 357 EGSSLPFQSIDNIHRGNCLPPSMLSVCDRITDDILQEDLQDTSGFFIDQTTDPLHTTSSR 178
Query: 69 EEVNGRL 75
+ NGRL
Sbjct: 177 QSTNGRL 157
>BE997453
Length = 795
Score = 26.6 bits (57), Expect = 9.1
Identities = 16/33 (48%), Positives = 18/33 (54%)
Frame = +1
Query: 205 RMLRWMSGKTRQDRIRNDTIREGRGGIHSRKVG 237
RML MSG TR DRIRN G + + VG
Sbjct: 1 RML*RMSGHTR*DRIRNKVFERNLG**YCK*VG 99
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.136 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,154,946
Number of Sequences: 36976
Number of extensions: 93828
Number of successful extensions: 431
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 431
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 431
length of query: 239
length of database: 9,014,727
effective HSP length: 93
effective length of query: 146
effective length of database: 5,575,959
effective search space: 814090014
effective search space used: 814090014
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC141114.8


