

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
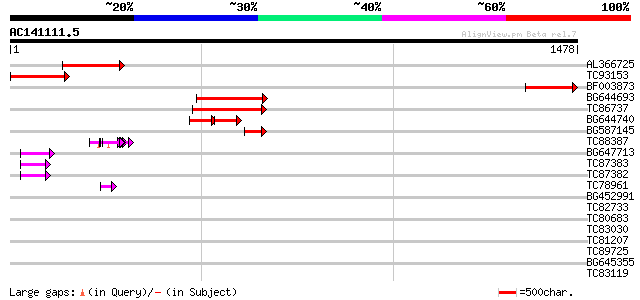
Query= AC141111.5 - phase: 0 /pseudo
(1478 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AL366725 317 3e-86
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 246 5e-65
BF003873 similar to GP|14715222|em putative polyprotein {Cicer a... 222 1e-57
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 195 1e-49
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 182 1e-45
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 88 2e-28
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 56 1e-07
TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 50 5e-06
BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent vir... 49 2e-05
TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA PO... 47 4e-05
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 45 2e-04
TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 44 6e-04
BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, pa... 38 0.004
TC82733 similar to GP|10177404|dbj|BAB10535. gene_id:K24M7.12~pi... 40 0.005
TC80683 homologue to GP|8777424|dbj|BAA97014.1 gb|AAF56406.1~gen... 34 0.48
TC83030 weakly similar to GP|18855061|gb|AAL79753.1 putative RNA... 34 0.48
TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-... 34 0.48
TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.... 32 2.4
BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.... 30 5.3
TC83119 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 30 5.3
>AL366725
Length = 485
Score = 317 bits (811), Expect = 3e-86
Identities = 148/161 (91%), Positives = 153/161 (94%)
Frame = +2
Query: 139 EFYPHYAAETVEFSKCIKFENGLRPDIKRAIEYQQIRVFPDLVNSCRIYEEDTKAHYKVV 198
+FYPHYAAET EFSKCIKFENGLRPDIKRAI YQQ+RVFPDLVN+CRIYEEDTKAH KVV
Sbjct: 2 KFYPHYAAETAEFSKCIKFENGLRPDIKRAIGYQQLRVFPDLVNTCRIYEEDTKAHDKVV 181
Query: 199 NERKTKGQQSRPKPYSAPADKGKQRMVDDRRPKKKDAPAEITCFNCGEKGHKSNVCPEEI 258
NERKTKGQ SRPKPYSAPADKGKQRMVDDRRPKKKDAPAEI CFN GEKGHKSNVCP+EI
Sbjct: 182 NERKTKGQ*SRPKPYSAPADKGKQRMVDDRRPKKKDAPAEIVCFNYGEKGHKSNVCPKEI 361
Query: 259 KKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQCKQPKK 299
KKCVRC KKGHIVADCKRNDIVCFN NEEGHIGSQCKQPK+
Sbjct: 362 KKCVRCDKKGHIVADCKRNDIVCFNCNEEGHIGSQCKQPKR 484
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 246 bits (628), Expect = 5e-65
Identities = 121/157 (77%), Positives = 130/157 (82%), Gaps = 2/157 (1%)
Frame = +2
Query: 1 AVAQAVGQQPAAGNGE--VRMLETFLRNHPPAFKGRYDPDGAQTWLKEVERIFRVMQCSE 58
AVAQAV Q P G RMLETFLRNHPP FKGRY PDGA WLKE+ERIFRVMQC E
Sbjct: 44 AVAQAVQQLPKVDTGSDGTRMLETFLRNHPPTFKGRYAPDGA*KWLKEIERIFRVMQCFE 223
Query: 59 VQKVRFGTHQLAEEADDWWVSLLPNLDQDGVDVTWAVF*REFMRRYFPEDVRRKKEIEFL 118
QKV+FGTH LAEEADDWW+SLLP L+QD VTWA+F +EF+ RYFPEDVR KKEIEFL
Sbjct: 224 TQKVQFGTHMLAEEADDWWISLLPVLEQDDAVVTWAMFRKEFLGRYFPEDVRGKKEIEFL 403
Query: 119 ELKQGNMYVTEYAAKFVELAEFYPHYAAETVEFSKCI 155
ELKQG+M VTEYAAKFVELA FYPHY+AET EFSKCI
Sbjct: 404 ELKQGDMSVTEYAAKFVELATFYPHYSAETAEFSKCI 514
>BF003873 similar to GP|14715222|em putative polyprotein {Cicer arietinum},
partial (82%)
Length = 559
Score = 222 bits (565), Expect = 1e-57
Identities = 112/135 (82%), Positives = 118/135 (86%)
Frame = +1
Query: 1344 WCWMCFEVKEVDSEIFRSVSDIEKIWNGGVSSGITTASFELARCFPCVATSELCSGSISC 1403
W W CFEVKEVD EI SVSDI K WNGGVSSGITTASFE A C CVATSE+CSGSISC
Sbjct: 1 WSWTCFEVKEVDCEIHWSVSDIRKSWNGGVSSGITTASFEFA*CLSCVATSEVCSGSISC 180
Query: 1404 DPE**CAS*RQPYGRDFTGED**SKSEDVERQGDTPC*SR*GWSDW*KLDMGA*E*DAGV 1463
DPE**CAS RQPYGRDFTGED**SKSED+ERQGDT C SR G S+W*KLD+GA*E*D GV
Sbjct: 181 DPE**CASQRQPYGRDFTGED**SKSEDIERQGDTSCESRLGQSEW*KLDVGA*E*DGGV 360
Query: 1464 LSRIVCLR*IFEDEN 1478
LSR+VC+R*IFEDEN
Sbjct: 361 LSRVVCMR*IFEDEN 405
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 195 bits (496), Expect = 1e-49
Identities = 102/186 (54%), Positives = 129/186 (68%)
Frame = +2
Query: 486 DEIPDVPLEREVEFSIDLILGTKPVSMAPYRMSASELAKLKKQLEDLLEKKFVRPSVSPW 545
D + VP E +++F IDL+ P+ + YR++ +L LK QL+DLLEK F++PS+ P
Sbjct: 2 DHLL*VPPEWKIDFGIDLLPNMNPI*IPSYRINPLKLKVLKLQLKDLLEKGFIQPSIYP* 181
Query: 546 GAPVLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYH 605
G VL +KKKDG +R+ IDY QLN V IK +YPLP ID+L D L G+K F KIDLR G H
Sbjct: 182 GVVVLFLKKKDGFLRMSIDYPQLNNVNIKIKYPLPLIDELFDNLQGSKWFFKIDLRLG*H 361
Query: 606 QIKVKDEDMQKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDI 665
Q +V ED+ KTAF RYGHYE VM FG TN P FME MNR+F +LD V+VF +DI
Sbjct: 362 QHRVIGEDVPKTAFRIRYGHYEILVMSFG*TNPPMAFMELMNRVFQDYLDSLVIVFSNDI 541
Query: 666 LIYSKD 671
LIYSK+
Sbjct: 542 LIYSKN 559
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 182 bits (461), Expect = 1e-45
Identities = 94/201 (46%), Positives = 135/201 (66%), Gaps = 7/201 (3%)
Frame = +1
Query: 476 VVCDFPEVF-PDEIPDVPLEREV-EFSIDLIL----GTKPVSMAP-YRMSASELAKLKKQ 528
V+ +FP++F P++ VP R + + +I LI P+ P Y MS EL LKK
Sbjct: 343 VLEEFPDLFNPEKAYQVPASRGLLDHAIPLIPDKDGNDPPLPWGPLYGMSRQELLVLKKT 522
Query: 529 LEDLLEKKFVRPSVSPWGAPVLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQ 588
LEDLL+K F++ S S GAPVL V+K G +R C+DYR LN +T K+RYPLP I + + +
Sbjct: 523 LEDLLDKGFIKASGSAAGAPVLFVRKPGGGIRFCVDYRALNAITKKDRYPLPLISETLRR 702
Query: 589 LVGAKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNR 648
+ GA+ F+K+D+ + +H++++KDED +KTAF TRYG +E+ V PFG+T AP F Y+N+
Sbjct: 703 VAGARWFTKLDVVAAFHKMRIKDEDQEKTAFRTRYGLFEWIVCPFGLTGAPATFQRYINK 882
Query: 649 IFHAFLDRFVVVFIDDILIYS 669
H FLD FV +IDD+LIY+
Sbjct: 883 TLHEFLDDFVTAYIDDVLIYT 945
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 88.2 bits (217), Expect(2) = 2e-28
Identities = 42/69 (60%), Positives = 52/69 (74%)
Frame = -1
Query: 535 KKFVRPSVSPWGAPVLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGAKV 594
K+F +PS+SP GA +L V+KKDG R+CIDYRQ NKVT KN+YPLPRID+L D++
Sbjct: 274 KRFQQPSISP*GAALLFVRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFDKIQEDCY 95
Query: 595 FSKIDLRSG 603
F IDLR G
Sbjct: 94 F*NIDLRLG 68
Score = 57.8 bits (138), Expect(2) = 2e-28
Identities = 35/73 (47%), Positives = 45/73 (60%), Gaps = 3/73 (4%)
Frame = -2
Query: 468 QAVIDKLQVVC---DFPEVFPDEIPDVPLEREVEFSIDLILGTKPVSMAPYRMSASELAK 524
++VI +VV F VFPD P +PLERE+ F IDL+L T+ +S P M +EL K
Sbjct: 483 ESVIPLFEVVLVLKGFS*VFPDNFPVIPLEREIFFCIDLLLDTQLISNPP*LMDRTELKK 304
Query: 525 LKKQLEDLLEKKF 537
LK L+D LEK F
Sbjct: 303 LKI*LKDSLEKGF 265
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 55.8 bits (133), Expect = 1e-07
Identities = 27/58 (46%), Positives = 38/58 (64%)
Frame = +2
Query: 612 EDMQKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYS 669
+D++KTAF T G Y YKVMPFG+ NA + +NR+F L + V+IDD+L+ S
Sbjct: 11 DDLEKTAFITDRGTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVKS 184
>TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (96%)
Length = 1286
Score = 50.4 bits (119), Expect = 5e-06
Identities = 34/115 (29%), Positives = 49/115 (42%), Gaps = 16/115 (13%)
Frame = +1
Query: 207 QSRPKPYS-APADKGKQRMVDDRR-----------PKKKDAPAEIT----CFNCGEKGHK 250
+SR K S +P D+ + R DRR P ++D+ + C NC GH
Sbjct: 274 RSRSKSRSRSPMDRSRSRSPVDRRIRSERFSHREAPYRRDSRRGFSQDNLCKNCKRPGHY 453
Query: 251 SNVCPEEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQCKQPKKSPTTGR 305
CP + C C GHI ++C + C+N E GH+ S C T G+
Sbjct: 454 VRECPN-VAVCHNCSLPGHIASECSTKSL-CWNCKEPGHMASSCPNEGICHTCGK 612
Score = 49.7 bits (117), Expect = 8e-06
Identities = 22/65 (33%), Positives = 33/65 (49%), Gaps = 6/65 (9%)
Frame = +1
Query: 236 PAEITCFNCGEKGHKSNVC------PEEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGH 289
P E C CG+ GH++ C P +++ C C K+GHI +C N+ C N + GH
Sbjct: 580 PNEGICHTCGKAGHRARECTVPQKPPGDLRLCNNCYKQGHIAVEC-TNEKACNNCRKTGH 756
Query: 290 IGSQC 294
+ C
Sbjct: 757 LARDC 771
Score = 48.1 bits (113), Expect = 2e-05
Identities = 28/83 (33%), Positives = 38/83 (45%)
Frame = +1
Query: 241 CFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQCKQPKKS 300
C NC ++GH + C E K C C K GH+ DC ND +C N GH+ QC +
Sbjct: 673 CNNCYKQGHIAVECTNE-KACNNCRKTGHLARDCP-NDPICNLCNISGHVARQCPKSNVI 846
Query: 301 PTTGRVFALDGTQTENEDRLIRC 323
G +L G + R + C
Sbjct: 847 GDRGGGGSLRGGYRDGGFRDVVC 915
Score = 46.2 bits (108), Expect = 9e-05
Identities = 24/64 (37%), Positives = 33/64 (51%), Gaps = 6/64 (9%)
Frame = +1
Query: 241 CFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADC-----KRNDI-VCFNFNEEGHIGSQC 294
C+NC E GH ++ CP E C CGK GH +C D+ +C N ++GHI +C
Sbjct: 538 CWNCKEPGHMASSCPNE-GICHTCGKAGHRARECTVPQKPPGDLRLCNNCYKQGHIAVEC 714
Query: 295 KQPK 298
K
Sbjct: 715 TNEK 726
Score = 45.1 bits (105), Expect = 2e-04
Identities = 20/69 (28%), Positives = 33/69 (46%)
Frame = +1
Query: 233 KDAPAEITCFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIGS 292
++ P C NC GH ++ C + C C + GH+ + C N+ +C + GH
Sbjct: 457 RECPNVAVCHNCSLPGHIASECSTK-SLCWNCKEPGHMASSCP-NEGICHTCGKAGHRAR 630
Query: 293 QCKQPKKSP 301
+C P+K P
Sbjct: 631 ECTVPQKPP 657
Score = 34.3 bits (77), Expect = 0.37
Identities = 13/43 (30%), Positives = 22/43 (50%)
Frame = +1
Query: 238 EITCFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADCKRNDIV 280
E C NC + GH + CP + C C GH+ C +++++
Sbjct: 721 EKACNNCRKTGHLARDCPND-PICNLCNISGHVARQCPKSNVI 846
>BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent virus}, partial
(1%)
Length = 726
Score = 48.5 bits (114), Expect = 2e-05
Identities = 27/95 (28%), Positives = 48/95 (50%), Gaps = 7/95 (7%)
Frame = +2
Query: 29 PAFKGRYDPDGAQTWLKEVERIFRVMQCSEVQKVRFGTHQLAEEADDWWVSLLPNLDQDG 88
P F G+ PD WL+ +ER+F+ + +E QKV+ +L + A WW +L + +G
Sbjct: 347 PDF*GKLQPDEFVDWLQTIERVFKYKEVAEEQKVKIVAAKLKKHASIWWKNLKRKRNCEG 526
Query: 89 VD--VTWAVF*REFMRR-----YFPEDVRRKKEIE 116
TW ++ R+ Y+ ++ +KK I+
Sbjct: 527 KSKIKTWDKMRQKLTRKYLHPHYYQDNFTQKKNIQ 631
>TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA POLYMERASE II
{Encephalitozoon cuniculi}, partial (0%)
Length = 1247
Score = 47.4 bits (111), Expect = 4e-05
Identities = 24/80 (30%), Positives = 40/80 (50%), Gaps = 2/80 (2%)
Frame = -2
Query: 29 PAFKGRYDPDGAQTWLKEVERIFRVMQCSEVQKVRFGTHQLAEEADDWWVSLLPNLDQDG 88
P F+G PD WL+ +ER+F+ + E QKV+ +L + A WW ++ ++G
Sbjct: 724 PDFEGNLQPDDLLDWLQIMERLFKYKEVLEEQKVKIVAAKLKKLASIWWENVKRRRKREG 545
Query: 89 VD--VTWAVF*REFMRRYFP 106
TW ++ R+Y P
Sbjct: 544 KSKIKTWEKMRQKLTRKYLP 485
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 45.1 bits (105), Expect = 2e-04
Identities = 24/80 (30%), Positives = 38/80 (47%), Gaps = 2/80 (2%)
Frame = +2
Query: 29 PAFKGRYDPDGAQTWLKEVERIFRVMQCSEVQKVRFGTHQLAEEADDWWVSLLPNLDQDG 88
P F+G D WL+ +ER+F + E QKV+ +L + A WW +L ++G
Sbjct: 629 PDFEGNLQLDDFLDWLQTIERVFEYKEVPEEQKVKIVAAKLKKHALIWWENLKRRRKREG 808
Query: 89 VD--VTWAVF*REFMRRYFP 106
TW ++ R+Y P
Sbjct: 809 KSKIKTWDKMRQKLTRKYLP 868
>TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (71%)
Length = 974
Score = 43.5 bits (101), Expect = 6e-04
Identities = 19/44 (43%), Positives = 25/44 (56%), Gaps = 2/44 (4%)
Frame = +2
Query: 236 PAEITCFNCGEKGHKSNVCP--EEIKKCVRCGKKGHIVADCKRN 277
P CFNCG GH + C + KC RCG++GHI +CK +
Sbjct: 344 PGSGRCFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNS 475
Score = 38.5 bits (88), Expect = 0.020
Identities = 19/43 (44%), Positives = 21/43 (48%), Gaps = 3/43 (6%)
Frame = +2
Query: 260 KCVRCGKKGHIVADCKRND--IVCFNFNEEGHIGSQCK-QPKK 299
+C CG GH DCK D C+ E GHI CK PKK
Sbjct: 356 RCFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSPKK 484
>BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, partial (33%)
Length = 560
Score = 37.7 bits (86), Expect(2) = 0.004
Identities = 31/71 (43%), Positives = 39/71 (54%), Gaps = 7/71 (9%)
Frame = +2
Query: 971 VRVT*TVQGFEFSL*MVTSECETRNVEE-------Y*RGTES*CQVCRLVGC**SD*RRC 1023
+ + *TVQ EF L* V +ECE +VE+ Y RG+E C+V R *SD*R
Sbjct: 59 LELP*TVQRLEFGL*SVATECEIGDVEDQQRIFG*YQRGSEGGCEVGRFNVWE*SD*RW* 238
Query: 1024 L*SRRSRCVEI 1034
*S SR V +
Sbjct: 239 F*S**SRSVAV 271
Score = 21.9 bits (45), Expect(2) = 0.004
Identities = 7/10 (70%), Positives = 9/10 (90%)
Frame = +3
Query: 952 CCRCFEQEDI 961
CCRCF+ E+I
Sbjct: 3 CCRCFK*ENI 32
>TC82733 similar to GP|10177404|dbj|BAB10535.
gene_id:K24M7.12~pir||S42136~similar to unknown protein
{Arabidopsis thaliana}, partial (57%)
Length = 710
Score = 40.4 bits (93), Expect = 0.005
Identities = 20/66 (30%), Positives = 28/66 (42%), Gaps = 12/66 (18%)
Frame = +3
Query: 241 CFNCGEKGHKSNVCP-----EEIKKCVRCGKKGHIVADCKRN-------DIVCFNFNEEG 288
C C +GH++ CP E+ K CG GH +A+C CF E+G
Sbjct: 357 CLRCRRRGHRAQNCPDGGSKEDFKY*YNCGDNGHSLANCPHPLQEGGTMFAQCFVCKEQG 536
Query: 289 HIGSQC 294
H+ C
Sbjct: 537 HLSKNC 554
Score = 36.6 bits (83), Expect = 0.074
Identities = 26/108 (24%), Positives = 44/108 (40%), Gaps = 10/108 (9%)
Frame = +3
Query: 200 ERKTKGQQSRPKPYSAPADKGKQRMVDDRRPKKKDAPAEITCFNCGEKGHKSNVCP---- 255
++K K ++ +P S P + V +P +CF C H + C
Sbjct: 180 KKKNKFKRKKPDSNSKPRTGKRPLRVPGMKPGD-------SCFICKGLDHIAKFCTQKAE 338
Query: 256 -EEIKKCVRCGKKGHIVADC-----KRNDIVCFNFNEEGHIGSQCKQP 297
E+ K C+RC ++GH +C K + +N + GH + C P
Sbjct: 339 WEKNKICLRCRRRGHRAQNCPDGGSKEDFKY*YNCGDNGHSLANCPHP 482
Score = 34.7 bits (78), Expect = 0.28
Identities = 13/43 (30%), Positives = 23/43 (53%), Gaps = 7/43 (16%)
Frame = +3
Query: 242 FNCGEKGHKSNVCPEEIK-------KCVRCGKKGHIVADCKRN 277
+NCG+ GH CP ++ +C C ++GH+ +C +N
Sbjct: 435 YNCGDNGHSLANCPHPLQEGGTMFAQCFVCKEQGHLSKNCPKN 563
>TC80683 homologue to GP|8777424|dbj|BAA97014.1
gb|AAF56406.1~gene_id:K9P8.7~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (16%)
Length = 1360
Score = 33.9 bits (76), Expect = 0.48
Identities = 25/88 (28%), Positives = 41/88 (46%), Gaps = 8/88 (9%)
Frame = +1
Query: 201 RKTKGQQSRPKPYSAPADKGKQR-----MVDDRRPKKKDAPAEITCFNCGEKGHKSNVCP 255
R KG+ + K A D+ ++ + +P KK+ + +KG KS+ P
Sbjct: 886 RGQKGKLKKMKEKYADQDEEERSIRMSLLASSGKPIKKEETLPV--IETSDKGKKSDSGP 1059
Query: 256 EEIKK-CVRCGKKGHIVADCKR--NDIV 280
+ K C +C K GH+ DCK ND++
Sbjct: 1060IDAPKICYKCKKVGHLSRDCKEQPNDLL 1143
>TC83030 weakly similar to GP|18855061|gb|AAL79753.1 putative RNA helicase
{Oryza sativa}, partial (7%)
Length = 624
Score = 33.9 bits (76), Expect = 0.48
Identities = 20/61 (32%), Positives = 27/61 (43%), Gaps = 4/61 (6%)
Frame = +1
Query: 201 RKTKGQQSRPKPYSAPADK----GKQRMVDDRRPKKKDAPAEITCFNCGEKGHKSNVCPE 256
R K S KP + D G+Q P + A TCF CGE GH+++ CP
Sbjct: 43 RSYKSGNSWSKPERSSRDDWLIGGRQSSRSSSSPNRSFAG---TCFTCGESGHRASDCPN 213
Query: 257 E 257
+
Sbjct: 214 K 216
>TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-1 {Triticum
aestivum}, partial (39%)
Length = 630
Score = 33.9 bits (76), Expect = 0.48
Identities = 23/68 (33%), Positives = 29/68 (41%), Gaps = 4/68 (5%)
Frame = +3
Query: 215 APADKGKQRMVDDRRPKKKDAPAEITCFNCGEKGHKSNVCPEEIKK----CVRCGKKGHI 270
A D GK + VD PK + P ++ N G G E + C CG GHI
Sbjct: 207 ADGDNGKTKAVDVTGPKGE--PLQVRQDNHGGGGGGRGFRGGERRNGGGGCYTCGDTGHI 380
Query: 271 VADCKRND 278
DC R+D
Sbjct: 381 ARDCDRSD 404
Score = 30.8 bits (68), Expect = 4.1
Identities = 19/88 (21%), Positives = 25/88 (27%), Gaps = 31/88 (35%)
Frame = +3
Query: 241 CFNCGEKGHKSNVCP-----------------EEIKKCVRCGKKGHIVADCKR------- 276
C+ CG+ GH + C + + C CG H DC R
Sbjct: 351 CYTCGDTGHIARDCDRSDRNDRNDRSGGGGGGDRDRACYTCGSFEHFARDCMRGGGNNNN 530
Query: 277 -------NDIVCFNFNEEGHIGSQCKQP 297
C+ GHI C P
Sbjct: 531 GGGGYGGGGTSCYRCGGVGHIARDCATP 614
>TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.150 -
Arabidopsis thaliana, partial (17%)
Length = 378
Score = 31.6 bits (70), Expect = 2.4
Identities = 15/54 (27%), Positives = 18/54 (32%), Gaps = 20/54 (37%)
Frame = +1
Query: 241 CFNCGEKGHKSNVCPE--------------------EIKKCVRCGKKGHIVADC 274
C+NCGE GH + C C CG+ GH DC
Sbjct: 16 CYNCGESGHMARECTSGGGGGGGRYGGGGGGGGGGGGGGSCYSCGESGHFARDC 177
>BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.5 [imported]
- Arabidopsis thaliana, partial (5%)
Length = 627
Score = 30.4 bits (67), Expect = 5.3
Identities = 21/96 (21%), Positives = 30/96 (30%), Gaps = 35/96 (36%)
Frame = -1
Query: 241 CFNCGEKGHKSNVC------PEEIK----------------KCVRCGKKGHIVADCKRND 278
C NCG H S C PE+ KC +C + GH ++C
Sbjct: 627 CSNCGGSDHSSAQCLHLRNPPEQTAGGAYVNTVSGSGGASGKCYKCQQPGHWASNCPSMS 448
Query: 279 IV-------------CFNFNEEGHIGSQCKQPKKSP 301
C+ N+ GH + C +P
Sbjct: 447 AANRVSGGSGGASGNCYKCNQPGHWANNCPNMSAAP 340
>TC83119 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (16%)
Length = 421
Score = 30.4 bits (67), Expect = 5.3
Identities = 15/64 (23%), Positives = 26/64 (40%)
Frame = +3
Query: 202 KTKGQQSRPKPYSAPADKGKQRMVDDRRPKKKDAPAEITCFNCGEKGHKSNVCPEEIKKC 261
+++ + P+ +D+ R RR + + C NC GH + CP + C
Sbjct: 231 RSRSRSRSPRIRKIRSDRHSYRDAPYRRDSSRGFSRDNLCKNCKRPGHYARECP-NVAVC 407
Query: 262 VRCG 265
CG
Sbjct: 408 HNCG 419
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.349 0.155 0.537
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 45,147,411
Number of Sequences: 36976
Number of extensions: 642061
Number of successful extensions: 5717
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 2262
Number of HSP's successfully gapped in prelim test: 264
Number of HSP's that attempted gapping in prelim test: 3275
Number of HSP's gapped (non-prelim): 2802
length of query: 1478
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1370
effective length of database: 5,021,319
effective search space: 6879207030
effective search space used: 6879207030
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC141111.5


